![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | lane2_Undetermined_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 3750802 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 46 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
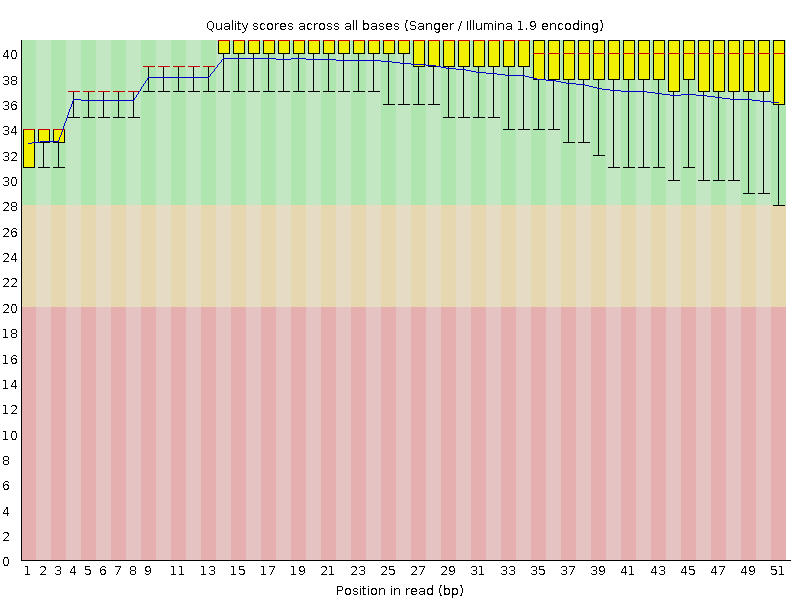
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
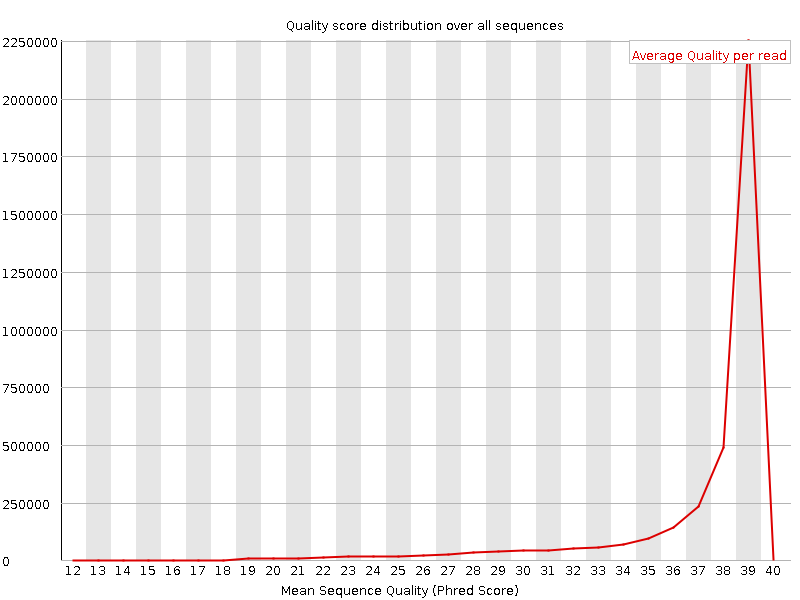
![[OK]](Icons/tick.png) Per base sequence content
Per base sequence content
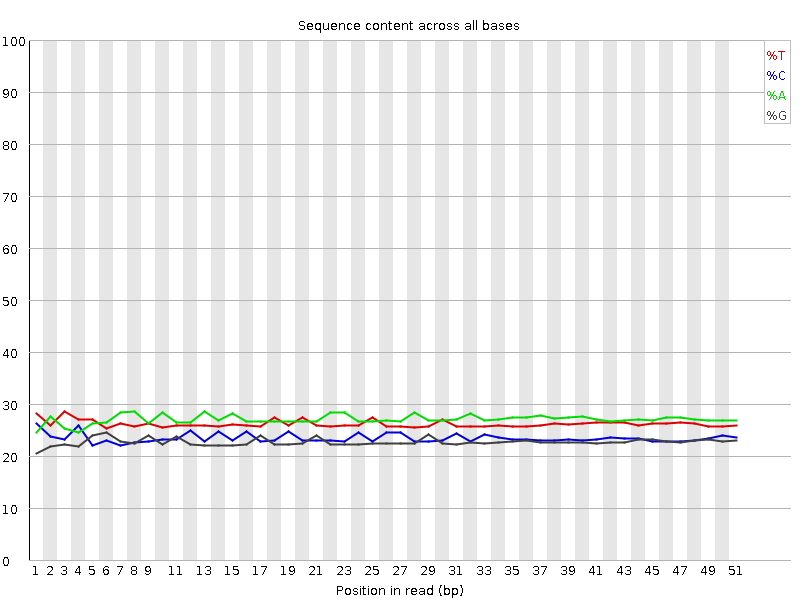
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
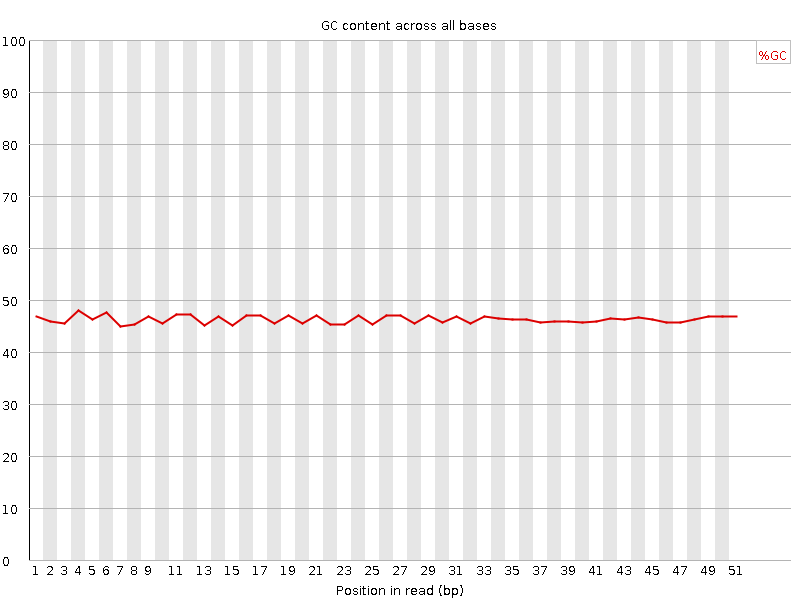
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
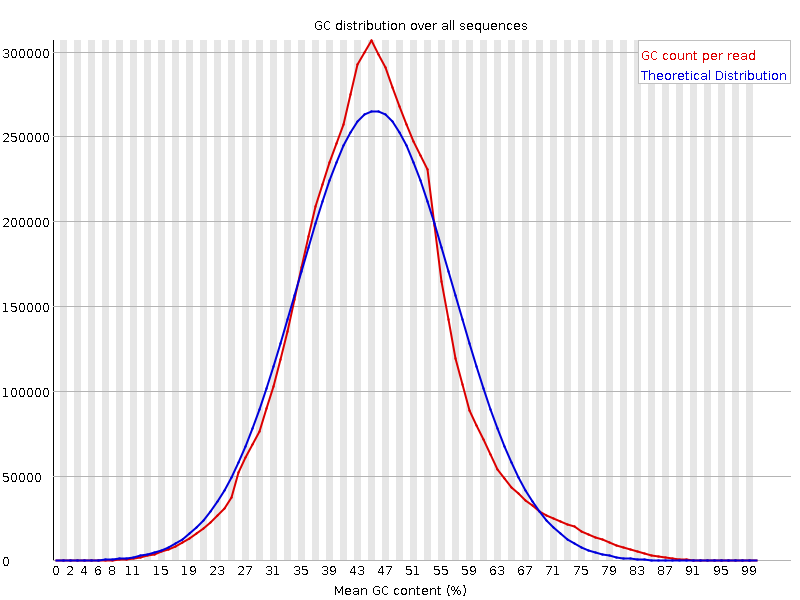
![[WARN]](Icons/warning.png) Per base N content
Per base N content
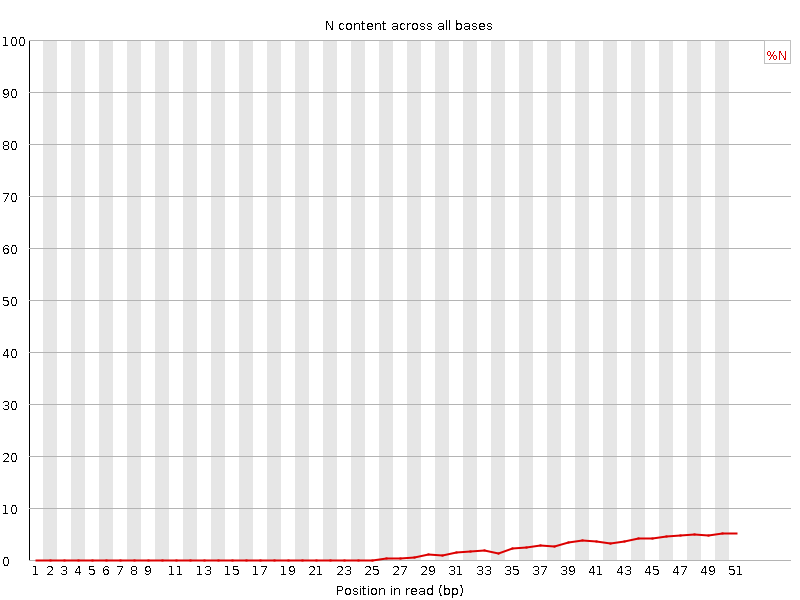
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
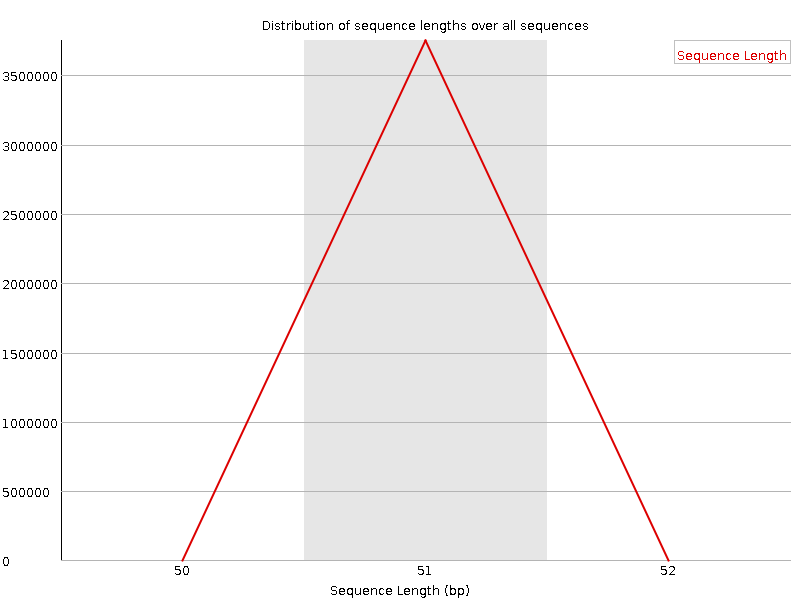
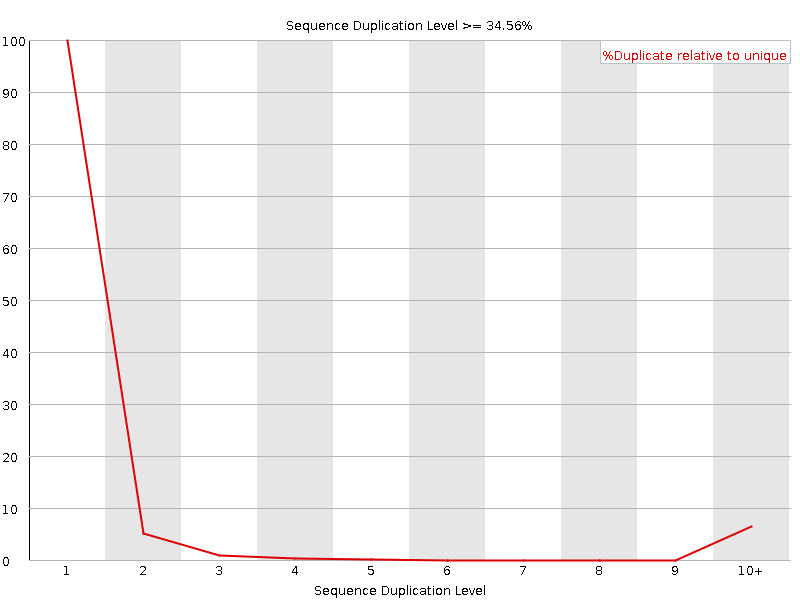
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCAGATCATCTCGTATGCC | 7543 | 0.2011036572978259 | TruSeq Adapter, Index 7 (100% over 51bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
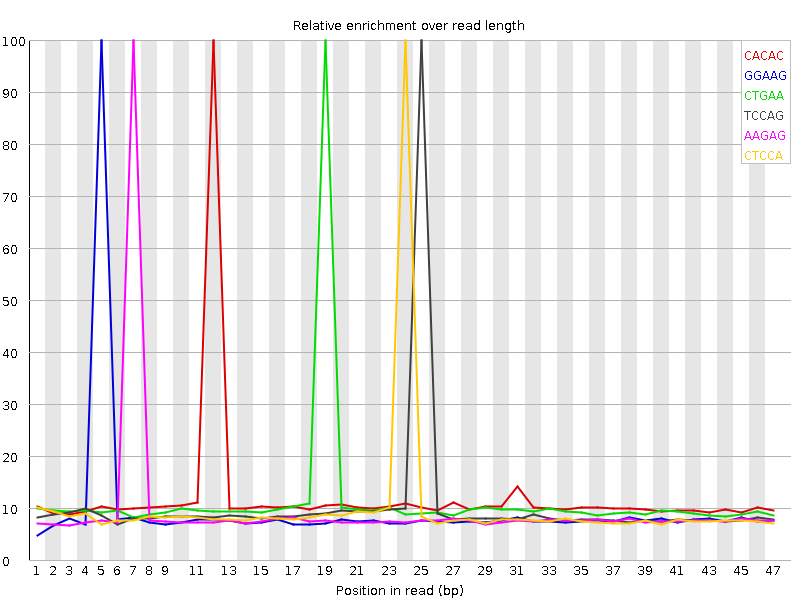
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CACAC | 426160 | 2.5233467 | 20.718317 | 12 |
| GGAAG | 348940 | 2.2874384 | 23.842667 | 5 |
| CTGAA | 375895 | 2.0647104 | 17.980726 | 19 |
| TCCAG | 305430 | 1.9345102 | 18.578587 | 25 |
| AAGAG | 344930 | 1.8955334 | 19.714172 | 7 |
| CTCCA | 305840 | 1.8725015 | 18.516493 | 24 |
| AGAGC | 289570 | 1.8349353 | 22.255947 | 8 |
| GAAGA | 323885 | 1.7798824 | 19.683752 | 6 |
| TCTGA | 308335 | 1.7512133 | 18.314362 | 18 |
| CACCA | 294170 | 1.7418175 | 6.2068 | 31 |
| ACCAG | 278385 | 1.7052246 | 5.404847 | 32 |
| GTCTG | 246455 | 1.6697524 | 21.530214 | 17 |
| AGCAC | 272325 | 1.6681043 | 20.947151 | 10 |
| GAGCA | 262855 | 1.6656487 | 21.833567 | 9 |
| GCACA | 268450 | 1.6443685 | 20.667423 | 11 |
| CAGTC | 253300 | 1.604333 | 17.254549 | 27 |
| CCAGT | 244210 | 1.5467595 | 17.866793 | 26 |
| ATGCC | 233480 | 1.4787985 | 8.631361 | 47 |
| AGTCA | 266025 | 1.4612179 | 14.284152 | 28 |
| ATCTC | 249400 | 1.3692449 | 7.6405954 | 40 |
| TCACC | 220215 | 1.3482634 | 7.877709 | 30 |
| TGAAC | 234500 | 1.2880579 | 17.02452 | 20 |
| ACTCC | 207910 | 1.2729262 | 18.129366 | 23 |
| AACTC | 238870 | 1.2683021 | 16.196177 | 22 |
| GAACT | 229220 | 1.259056 | 16.841673 | 21 |
| GTCAC | 198320 | 1.2561047 | 14.974693 | 29 |
| GTATG | 212965 | 1.251284 | 7.7485104 | 45 |
| CACGT | 183230 | 1.1605288 | 20.250515 | 14 |
| TATGC | 199090 | 1.1307474 | 7.4485235 | 46 |
| TGCCG | 144665 | 1.0930021 | 8.464666 | 47 |
| CGGAA | 167685 | 1.0625794 | 22.019733 | 4 |
| TCTCG | 146170 | 0.9572849 | 8.476314 | 41 |
| ACGTC | 150710 | 0.954556 | 19.791862 | 15 |
| CGTCT | 143940 | 0.9426803 | 20.22429 | 16 |
| CTCGT | 135710 | 0.88878113 | 8.295182 | 42 |
| ACACG | 133690 | 0.8189071 | 19.383215 | 13 |
| ATCGG | 120600 | 0.79020196 | 22.51775 | 2 |
| TCGGA | 118990 | 0.7796529 | 22.504026 | 3 |
| CGTAT | 135815 | 0.7713721 | 7.0522876 | 44 |
| GATCG | 94620 | 0.6199744 | 22.298536 | 1 |
| TCGTA | 107275 | 0.60927695 | 6.963906 | 43 |