![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | lane1_Undetermined_L001_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 4707422 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 46 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
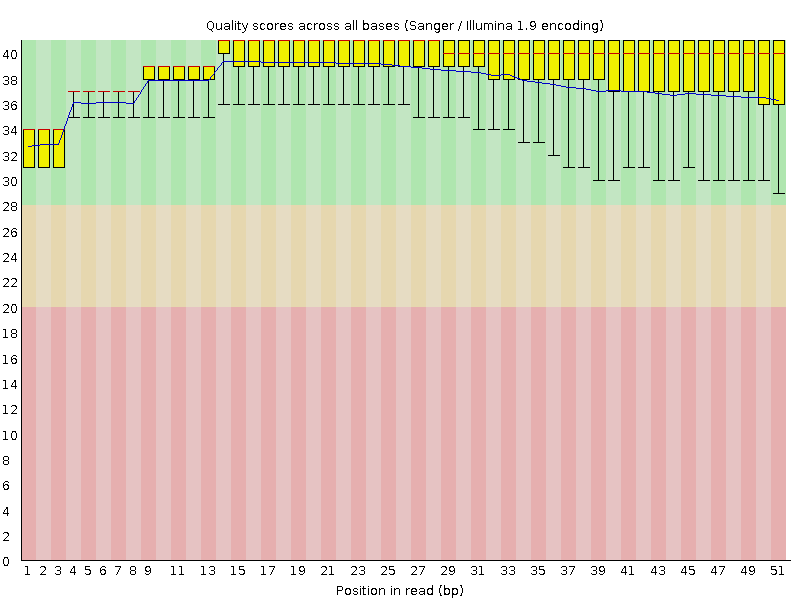
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
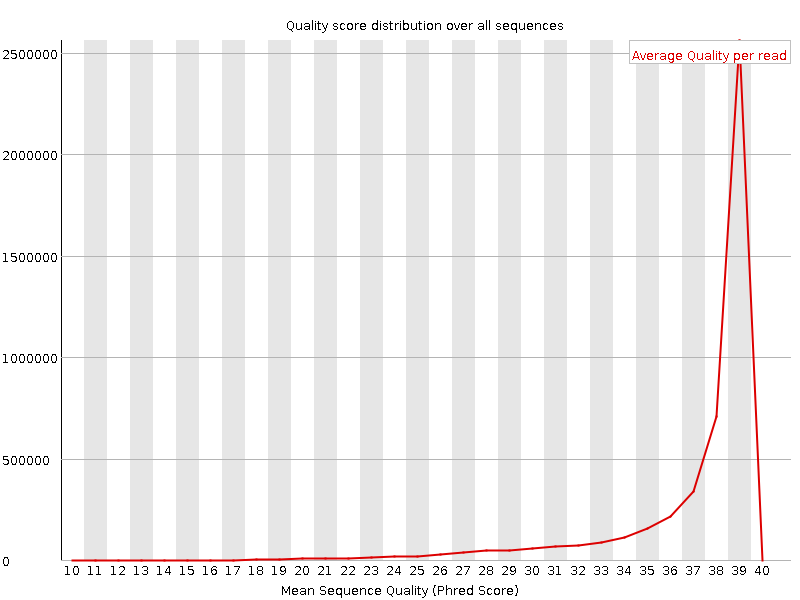
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
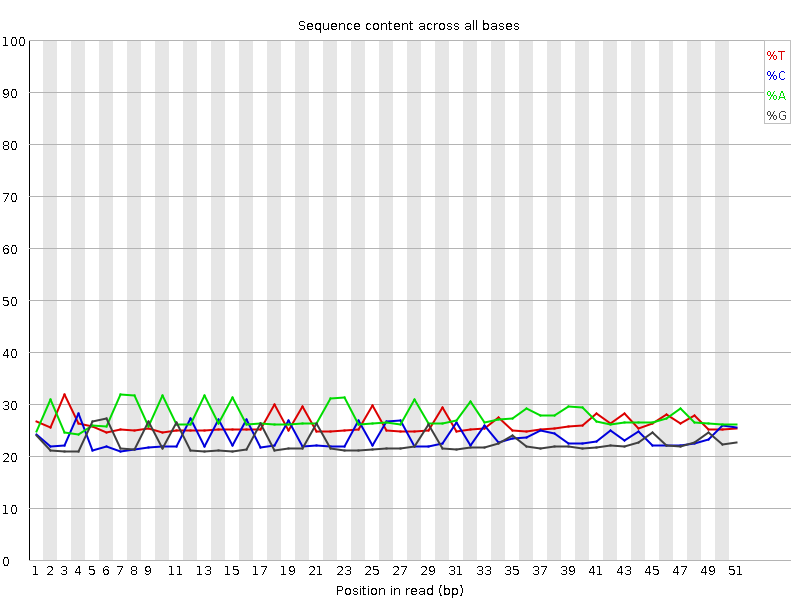
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
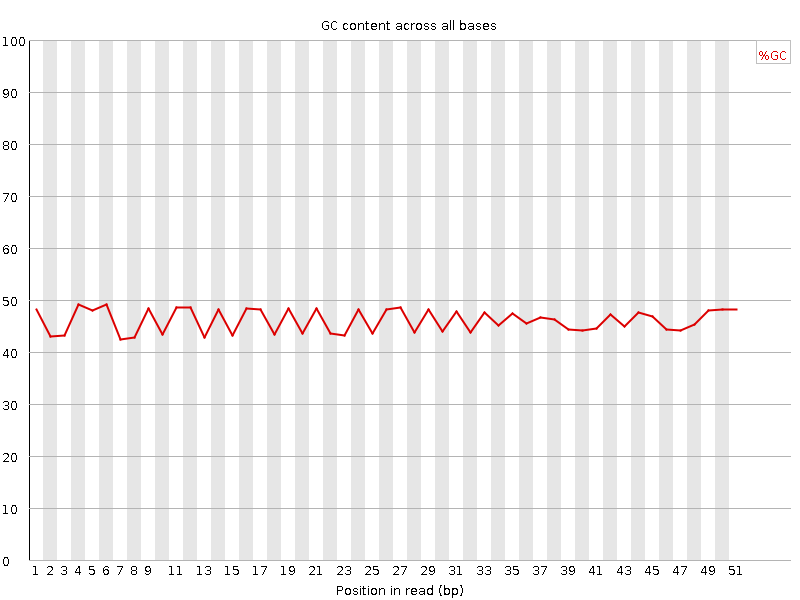
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
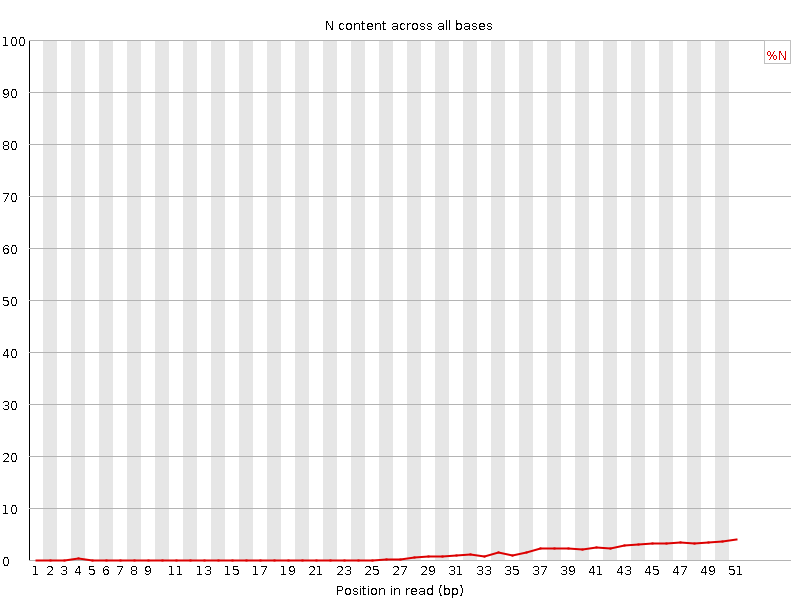
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
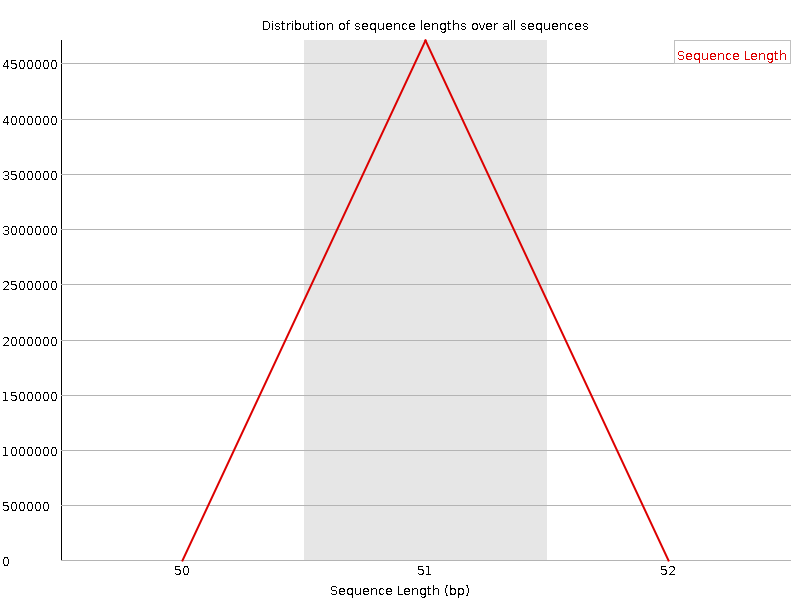
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACTGACCAATCTCGTATGCC | 66317 | 1.4087753339301214 | TruSeq Adapter, Index 4 (100% over 51bp) |
| TATTCGTTGGAATTCCTCGGGGAATTCGGTATTCCCAGGCGGTCTCCCATC | 6461 | 0.1372513447912679 | No Hit |
| ACTTCCAGGGATTTATAAGCCGATGACGTCATAACATCCCTGACCCTTTAA | 5834 | 0.12393195256342007 | No Hit |
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACTGCCCAATCTCGTATGCC | 5651 | 0.12004447444907212 | TruSeq Adapter, Index 4 (98% over 51bp) |
| GATCGGAAGAGCACACGTCTGAACTCCATCACTGACCAATCTCGTATGCCG | 5035 | 0.10695875576908125 | Illumina Multiplexing PCR Primer 2.01 (96% over 28bp) |
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACTACCAATCTCGTATGCCG | 4871 | 0.10347489560103174 | TruSeq Adapter, Index 10 (97% over 37bp) |
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGCAATATCTCGTATGCCG | 4718 | 0.1002247089808392 | TruSeq Adapter, Index 6 (97% over 37bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
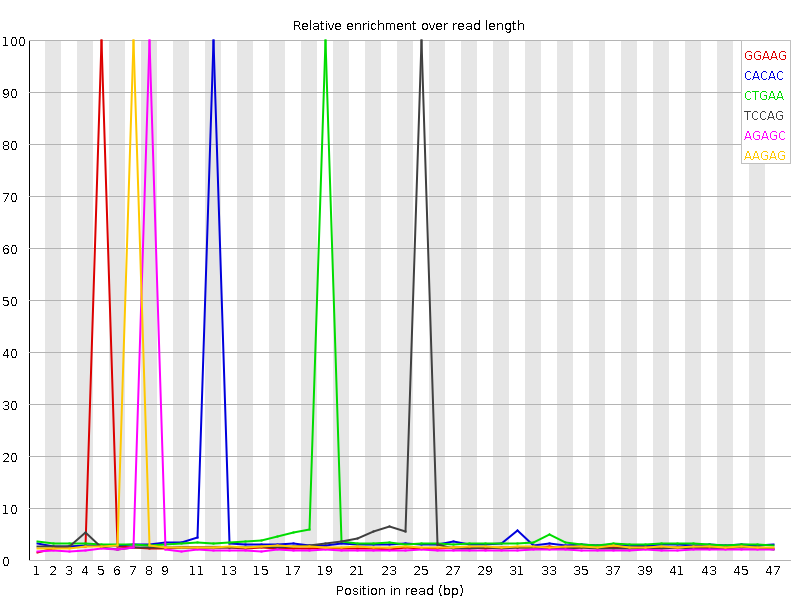
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGAAG | 592655 | 3.0697176 | 66.31335 | 5 |
| CACAC | 629670 | 2.8766398 | 53.89318 | 12 |
| CTGAA | 624180 | 2.6841857 | 47.87755 | 19 |
| TCCAG | 511900 | 2.5776725 | 51.34249 | 25 |
| AGAGC | 518355 | 2.5748334 | 61.138947 | 8 |
| AAGAG | 597750 | 2.5357208 | 53.063503 | 7 |
| CTCCA | 517700 | 2.500035 | 51.010277 | 24 |
| GTCTG | 445170 | 2.470806 | 61.35477 | 17 |
| TCTGA | 534150 | 2.4280713 | 50.521523 | 18 |
| GAAGA | 567315 | 2.4066126 | 53.7219 | 6 |
| GAGCA | 477755 | 2.3731604 | 59.975883 | 9 |
| AGCAC | 497825 | 2.371504 | 57.089436 | 10 |
| CAGTC | 468150 | 2.3573692 | 48.967026 | 27 |
| GCACA | 469765 | 2.2378337 | 55.954624 | 11 |
| ATGCC | 444245 | 2.236996 | 33.93915 | 47 |
| CCAGT | 443330 | 2.232388 | 50.179913 | 26 |
| CACTG | 425195 | 2.1410694 | 26.445932 | 31 |
| ACCAA | 532275 | 2.0766757 | 17.290148 | 36 |
| ATCTC | 474810 | 2.0698712 | 29.520054 | 40 |
| GTATG | 433915 | 2.056731 | 31.817629 | 45 |
| AGTCA | 477755 | 2.0545087 | 40.20707 | 28 |
| ACTCC | 416645 | 2.0120282 | 51.272095 | 23 |
| CACGT | 393050 | 1.9792032 | 56.77769 | 14 |
| CGGAA | 397820 | 1.976098 | 63.059574 | 4 |
| TGAAC | 458265 | 1.970695 | 46.809116 | 20 |
| CTGAC | 386230 | 1.9448613 | 21.693855 | 33 |
| GTCAC | 385910 | 1.9432497 | 44.10557 | 29 |
| TATGC | 416355 | 1.8926136 | 30.428024 | 46 |
| GAACT | 439435 | 1.8897196 | 46.552788 | 21 |
| ACTGA | 437905 | 1.8831398 | 19.691042 | 32 |
| AACTC | 452545 | 1.8663357 | 44.498676 | 22 |
| GACCA | 391735 | 1.8661199 | 19.98162 | 35 |
| TCTCG | 350390 | 1.8650469 | 35.36317 | 41 |
| CGTCT | 350340 | 1.8647808 | 58.721725 | 16 |
| CTCGT | 343335 | 1.8274947 | 35.255688 | 42 |
| ACGTC | 357590 | 1.8006444 | 55.869785 | 15 |
| ATCGG | 335540 | 1.7618202 | 66.58716 | 2 |
| TCGGA | 329670 | 1.7309986 | 66.58047 | 3 |
| TGACC | 337450 | 1.6992296 | 21.02003 | 34 |
| AATCT | 439300 | 1.6354783 | 19.702158 | 39 |
| TCACT | 371530 | 1.6196359 | 26.409721 | 30 |
| CGTAT | 350740 | 1.5943491 | 30.126942 | 44 |
| ACACG | 328835 | 1.5664812 | 53.786217 | 13 |
| CCAAT | 375545 | 1.5487812 | 19.391151 | 37 |
| GATCG | 293575 | 1.5414746 | 66.38361 | 1 |
| CAATC | 373510 | 1.5403883 | 20.013666 | 38 |
| TGCCG | 248840 | 1.5299505 | 16.200943 | 47 |
| TCGTA | 304995 | 1.3864075 | 29.887953 | 43 |
| GCCGT | 178015 | 1.094495 | 6.889492 | 47 |
| TCACG | 202860 | 1.0215015 | 8.090554 | 30 |
| CACGC | 149085 | 0.83161247 | 6.604468 | 31 |