![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | SV40_CAGATC_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 24496430 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 43 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
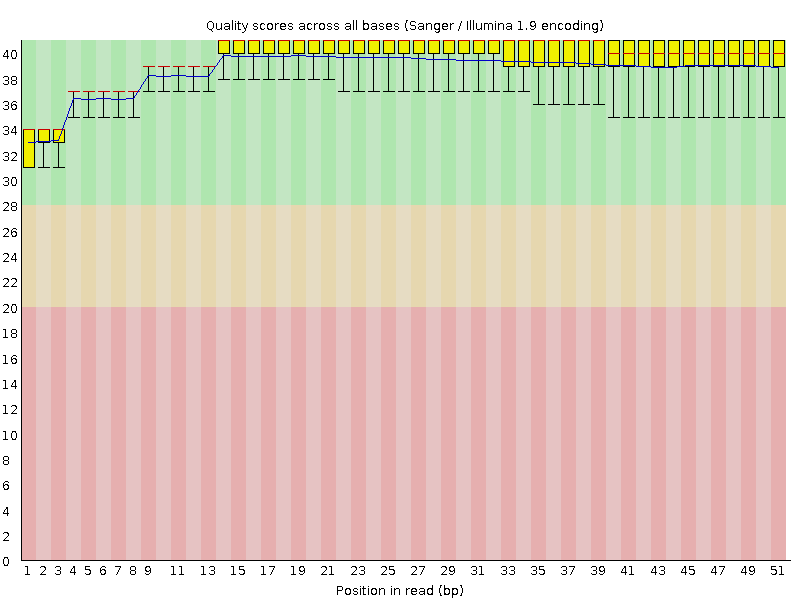
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
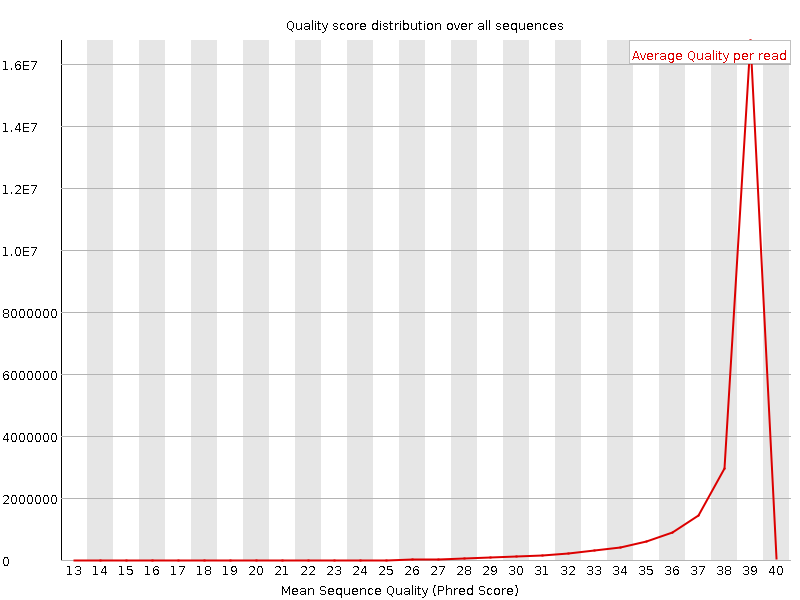
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
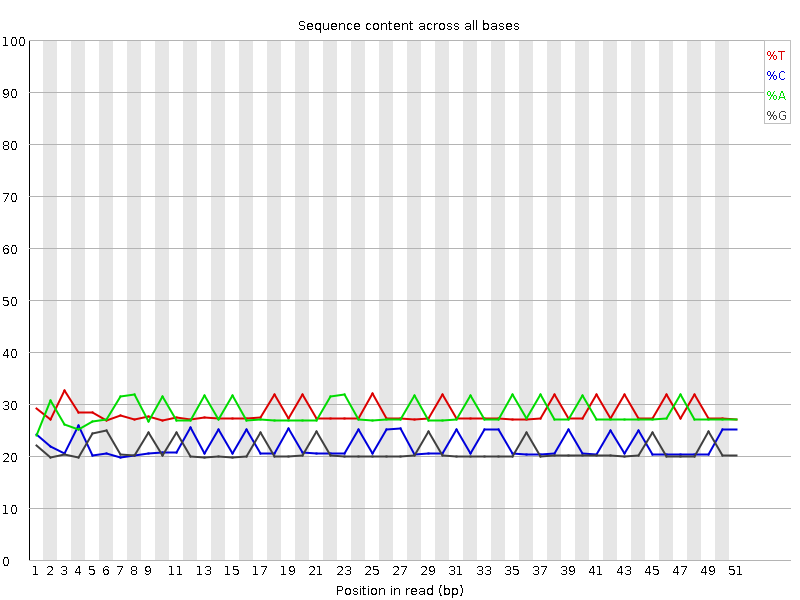
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
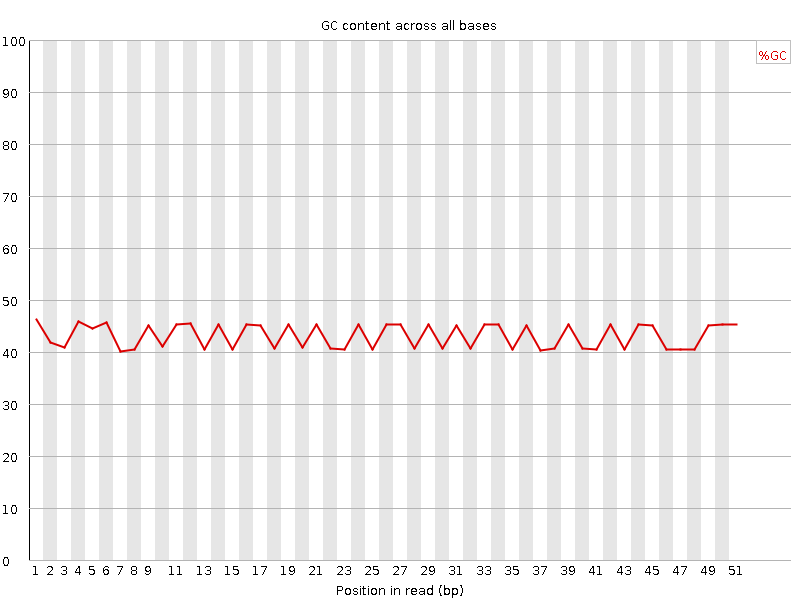
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
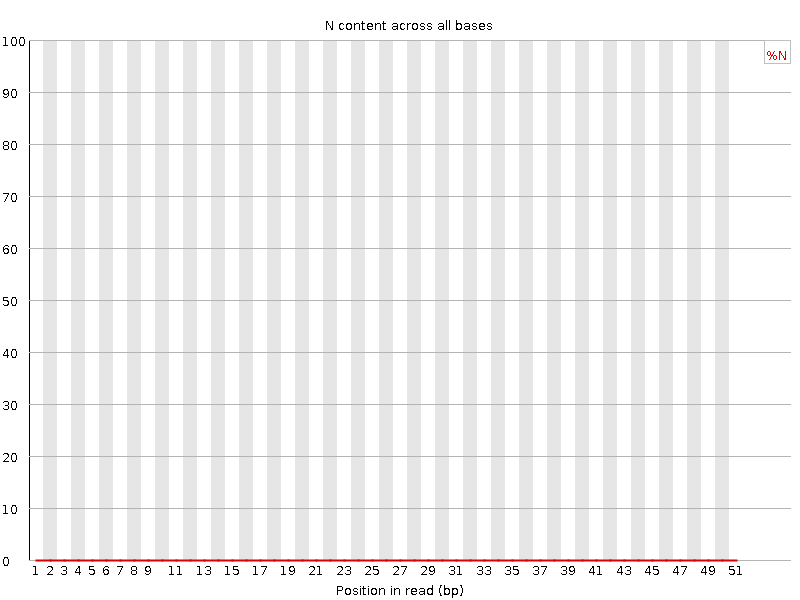
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
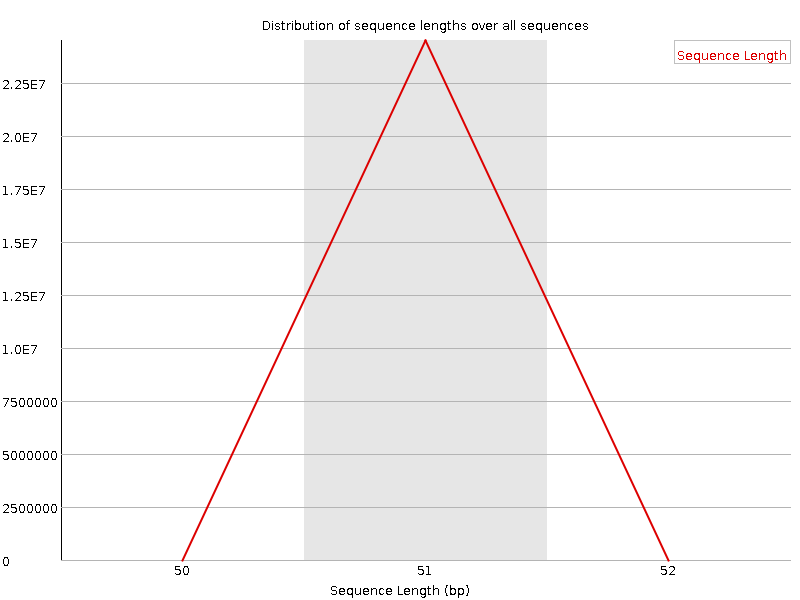
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
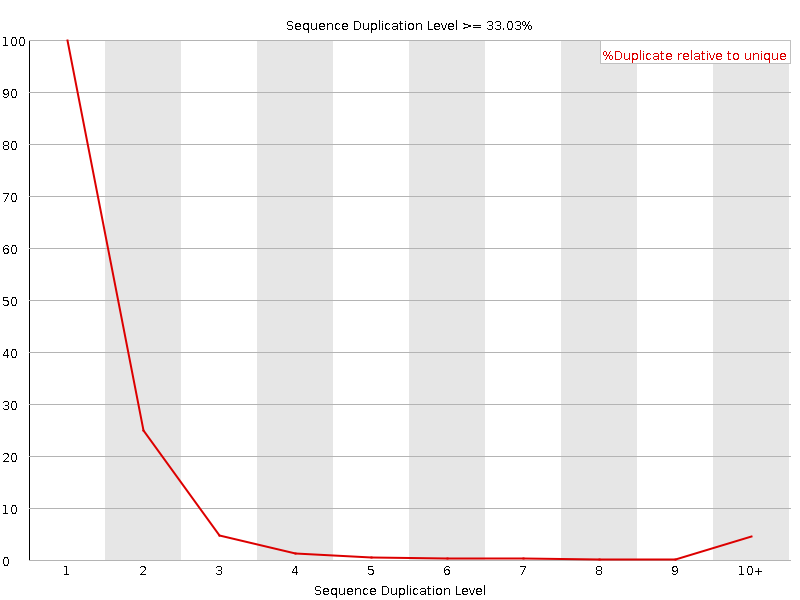
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCAGATCATCTCGTATGCC | 1113539 | 4.54571951912993 | TruSeq Adapter, Index 7 (100% over 51bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
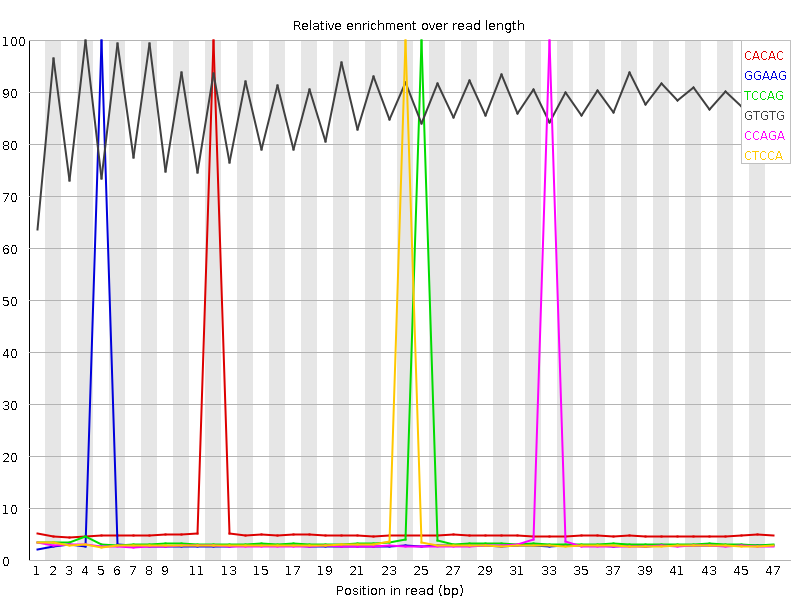
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CACAC | 3982965 | 3.950926 | 57.10507 | 12 |
| GGAAG | 2818030 | 3.2506382 | 65.58499 | 5 |
| TCCAG | 3011405 | 3.1316164 | 58.14411 | 25 |
| GTGTG | 2656470 | 3.0454733 | 3.4918354 | 4 |
| CCAGA | 2778030 | 2.89783 | 57.954174 | 33 |
| CTCCA | 2825305 | 2.7939663 | 55.127922 | 24 |
| CTGAA | 3399435 | 2.7705107 | 46.223118 | 19 |
| AGAGC | 2451290 | 2.688897 | 61.83807 | 8 |
| GCACA | 2507485 | 2.615618 | 58.611843 | 11 |
| GAGCA | 2377110 | 2.6075268 | 61.660732 | 9 |
| GTCTG | 2376835 | 2.5912263 | 60.792023 | 17 |
| CACCA | 2594135 | 2.5732677 | 54.936073 | 31 |
| GAAGA | 2978975 | 2.5609436 | 49.00372 | 6 |
| AAGAG | 2879595 | 2.4755096 | 48.851204 | 7 |
| ACCAG | 2367810 | 2.4699194 | 57.510498 | 32 |
| CCAGT | 2356570 | 2.4506412 | 57.35064 | 26 |
| AGCAC | 2298430 | 2.3975477 | 58.496635 | 10 |
| TCTGA | 2876895 | 2.3374403 | 45.692547 | 18 |
| CAGTC | 2235285 | 2.3245146 | 57.228455 | 27 |
| ATGCC | 2109585 | 2.1937969 | 56.992252 | 47 |
| CATCT | 2794170 | 2.1588666 | 42.755646 | 39 |
| TCACC | 2156755 | 2.132832 | 54.355015 | 30 |
| ACTCC | 2134440 | 2.1107645 | 54.582985 | 23 |
| GTCAC | 2020265 | 2.1009114 | 57.014877 | 29 |
| CAGAT | 2546820 | 2.0756366 | 45.085327 | 34 |
| CACGT | 1905425 | 1.981487 | 57.647797 | 14 |
| AGTCA | 2424060 | 1.9755883 | 45.006996 | 28 |
| GAACT | 2349925 | 1.9151689 | 45.225132 | 21 |
| AACTC | 2459645 | 1.9062594 | 43.109074 | 22 |
| ATCTC | 2382640 | 1.8409052 | 42.473866 | 40 |
| TGAAC | 2222510 | 1.8113269 | 45.159256 | 20 |
| TCATC | 2314175 | 1.7880069 | 42.409046 | 38 |
| GTATG | 2059995 | 1.760055 | 46.751083 | 45 |
| AGATC | 2132345 | 1.737843 | 44.680656 | 35 |
| ACGTC | 1639520 | 1.7049675 | 57.315453 | 15 |
| CGGAA | 1529050 | 1.677263 | 61.035007 | 4 |
| TCTCG | 1567605 | 1.6251725 | 56.15797 | 41 |
| GATCA | 1964015 | 1.6006556 | 44.548126 | 36 |
| TATGC | 1967520 | 1.5985848 | 44.38593 | 46 |
| ATCAT | 2520185 | 1.5260162 | 33.477165 | 37 |
| ACACG | 1461220 | 1.5242337 | 57.400684 | 13 |
| TCGGA | 1345330 | 1.4712001 | 60.708294 | 3 |
| CGTCT | 1403560 | 1.4551032 | 56.845524 | 16 |
| CTCGT | 1385260 | 1.4361311 | 55.924572 | 42 |
| ATCGG | 1306985 | 1.4292675 | 60.69754 | 2 |
| GATCG | 1299875 | 1.4214922 | 60.62434 | 1 |
| CGTAT | 1335435 | 1.0850238 | 43.88064 | 44 |
| TCGTA | 1296120 | 1.0530809 | 43.826187 | 43 |