![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | SV39_GCCAAT_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 26997017 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 45 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
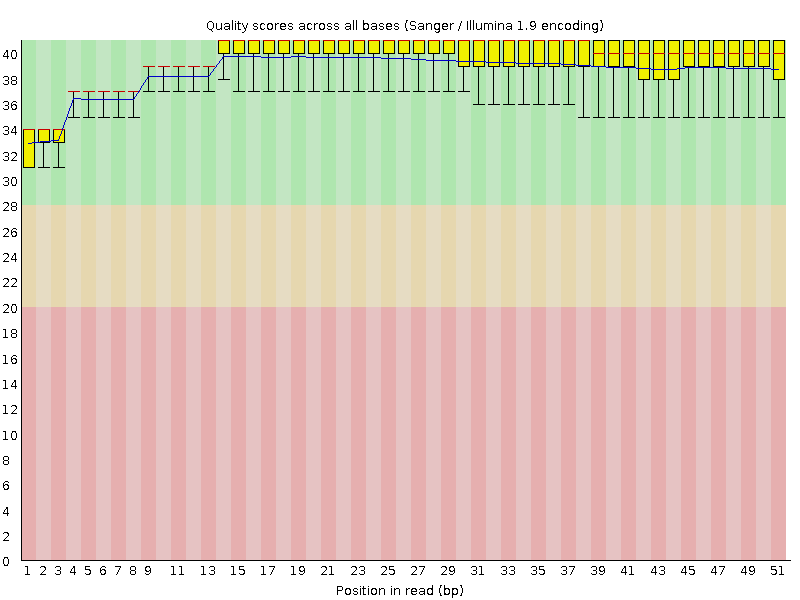
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
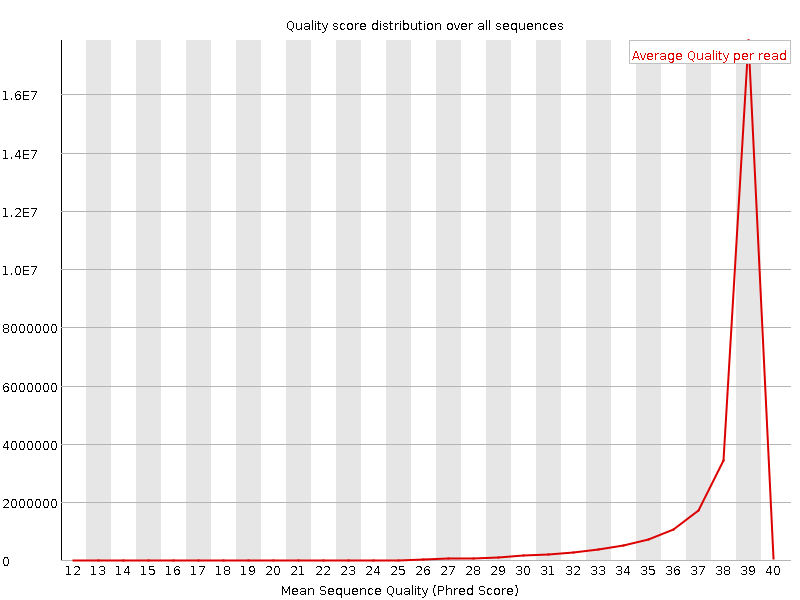
![[OK]](Icons/tick.png) Per base sequence content
Per base sequence content
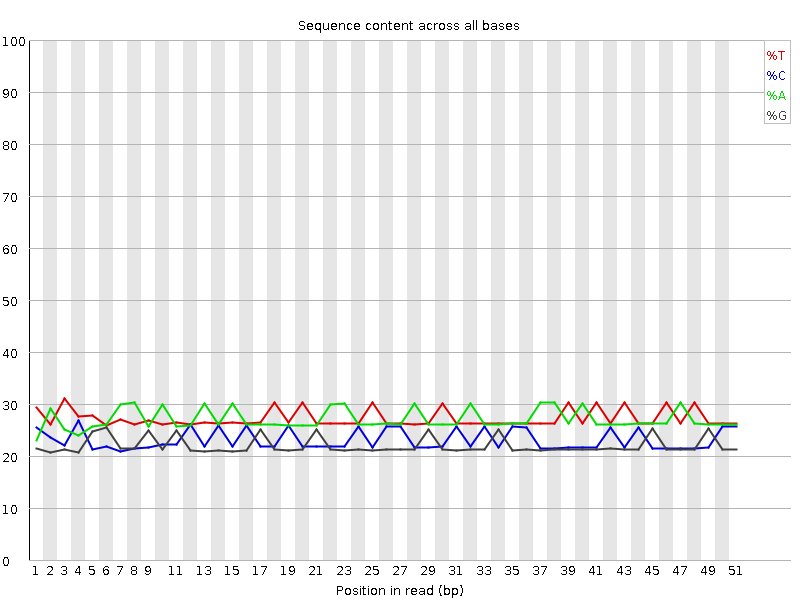
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
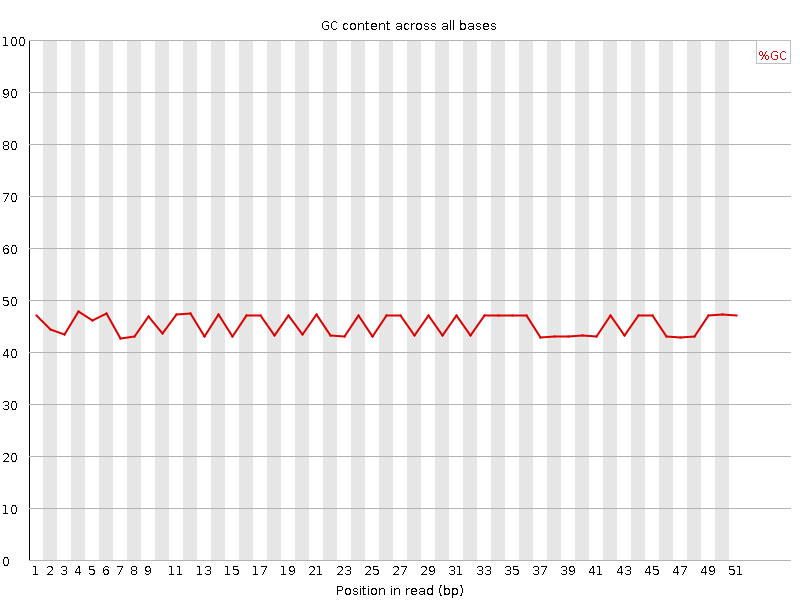
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
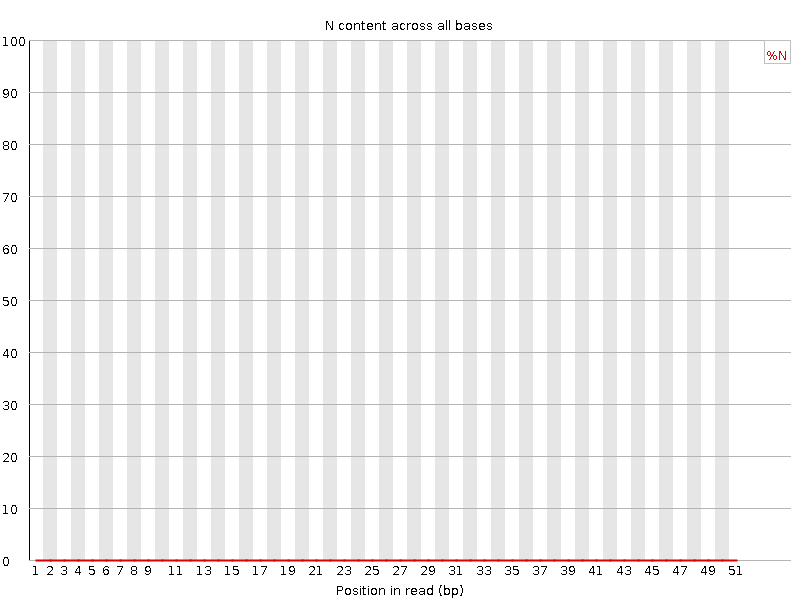
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
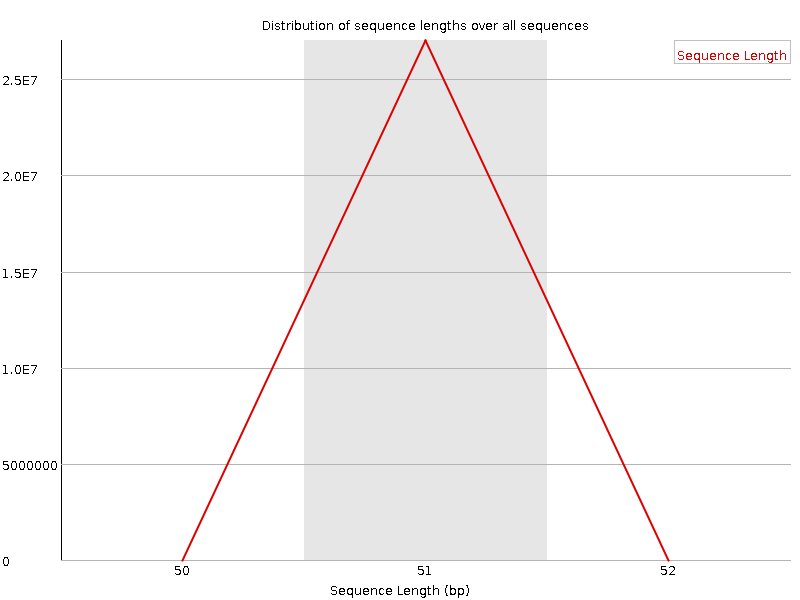
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
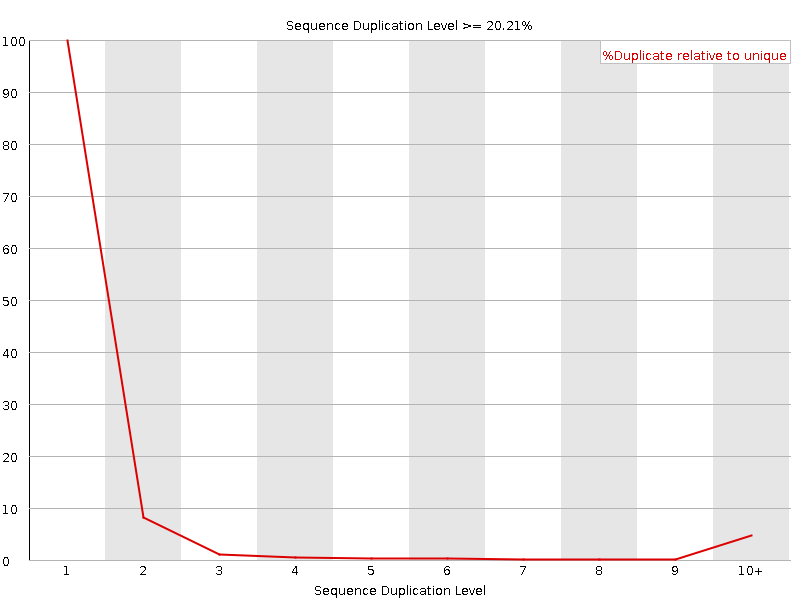
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGCCAATATCTCGTATGCC | 1048897 | 3.8852329499959195 | TruSeq Adapter, Index 6 (100% over 51bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
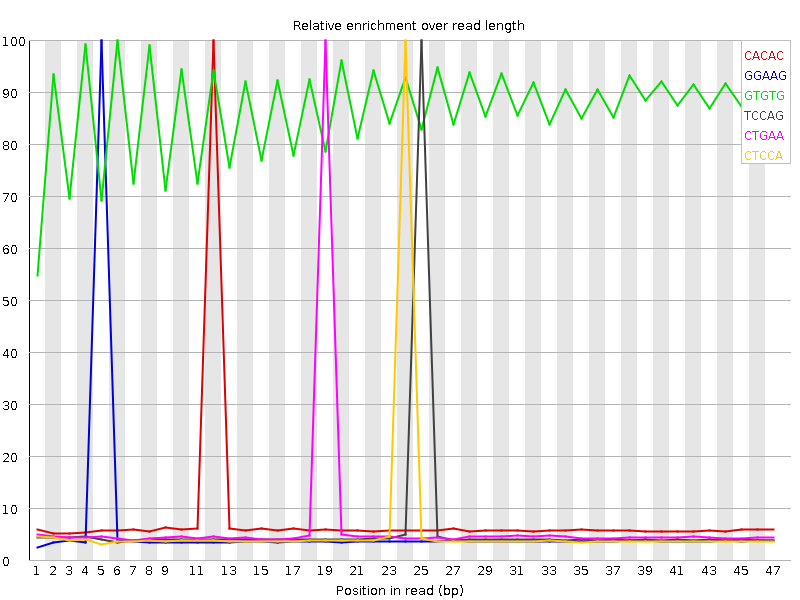
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CACAC | 4343685 | 3.6737866 | 46.53308 | 12 |
| GGAAG | 3127985 | 3.0635104 | 53.221306 | 5 |
| GTGTG | 3099865 | 3.0003328 | 3.4651349 | 6 |
| TCCAG | 3304670 | 2.9177816 | 47.14272 | 25 |
| CTGAA | 3550065 | 2.6783442 | 40.711517 | 19 |
| CTCCA | 3142935 | 2.6425734 | 44.74196 | 24 |
| GAAGA | 3169665 | 2.5260417 | 43.331524 | 6 |
| AGAGC | 2652495 | 2.4738665 | 50.134827 | 8 |
| GCACA | 2726415 | 2.4214785 | 47.58799 | 11 |
| GAGCA | 2590055 | 2.4156318 | 49.982944 | 9 |
| AAGAG | 3021200 | 2.4077234 | 43.116344 | 7 |
| GTCTG | 2531210 | 2.3330383 | 48.898716 | 17 |
| TCTGA | 2987980 | 2.24101 | 40.0829 | 18 |
| CCAGT | 2516415 | 2.2218099 | 46.302834 | 26 |
| AGCAC | 2457945 | 2.1830359 | 47.452335 | 10 |
| CAGTC | 2368805 | 2.0914812 | 46.1964 | 27 |
| ATGCC | 2201870 | 1.9440899 | 45.987152 | 47 |
| ACTCC | 2266145 | 1.9053699 | 44.099228 | 23 |
| GCCAA | 2119465 | 1.882413 | 46.039165 | 34 |
| AGTCA | 2481645 | 1.8722755 | 39.557903 | 28 |
| GTCAC | 2109360 | 1.8624102 | 45.961323 | 29 |
| CGCCA | 1770740 | 1.8405093 | 53.50976 | 33 |
| GAACT | 2409790 | 1.8180646 | 39.744232 | 21 |
| AACTC | 2495310 | 1.7927575 | 37.896965 | 22 |
| ATCTC | 2448140 | 1.7485148 | 37.295082 | 40 |
| CACGT | 1960610 | 1.7310749 | 46.48293 | 14 |
| TGAAC | 2260920 | 1.7057496 | 39.67899 | 20 |
| TCACG | 1881540 | 1.6612619 | 45.67998 | 30 |
| GTATG | 2037600 | 1.6047894 | 40.84993 | 45 |
| AATAT | 3066280 | 1.5990268 | 27.510208 | 37 |
| CACGC | 1480540 | 1.538875 | 53.24597 | 31 |
| CCAAT | 2105905 | 1.5129892 | 37.25778 | 35 |
| TATCT | 2436630 | 1.4783051 | 31.618982 | 39 |
| TATGC | 1956830 | 1.4676387 | 38.861073 | 46 |
| ACGTC | 1650285 | 1.4570806 | 46.18457 | 15 |
| CGGAA | 1539800 | 1.4361045 | 49.314487 | 4 |
| ACGCC | 1364965 | 1.4187461 | 53.109184 | 32 |
| TCTCG | 1578245 | 1.385272 | 45.068596 | 41 |
| ACACG | 1478160 | 1.3128347 | 46.36934 | 13 |
| CAATA | 2100130 | 1.2892842 | 31.847248 | 36 |
| ATATC | 2068870 | 1.2626172 | 31.584208 | 38 |
| TCGGA | 1340110 | 1.2425051 | 48.875843 | 3 |
| CGTCT | 1396900 | 1.2261003 | 45.607243 | 16 |
| CTCGT | 1377870 | 1.2093971 | 44.86997 | 42 |
| ATCGG | 1281910 | 1.1885438 | 48.79541 | 2 |
| GATCG | 1275750 | 1.1828326 | 48.728577 | 1 |
| CGTAT | 1283285 | 0.9624745 | 38.355118 | 44 |
| TCGTA | 1243910 | 0.9329429 | 38.308197 | 43 |