![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | SV38_ACAGTG_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 22649290 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 46 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
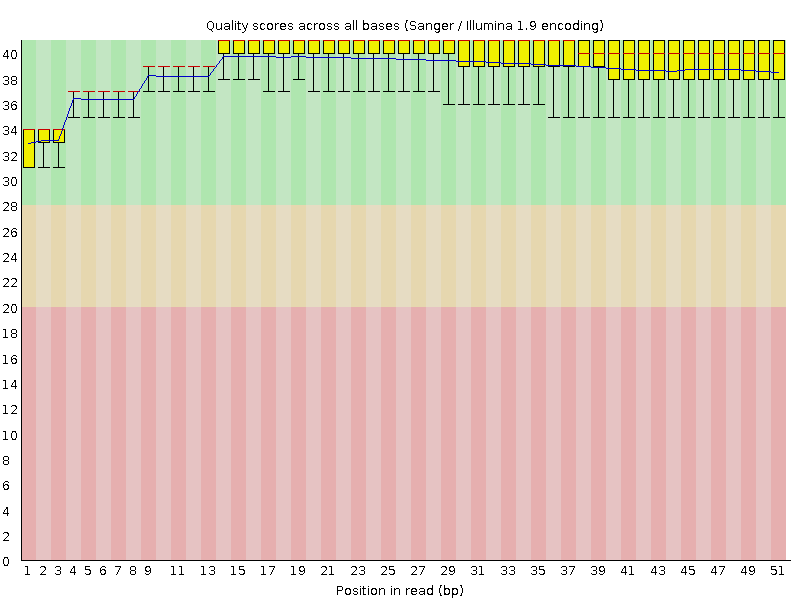
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
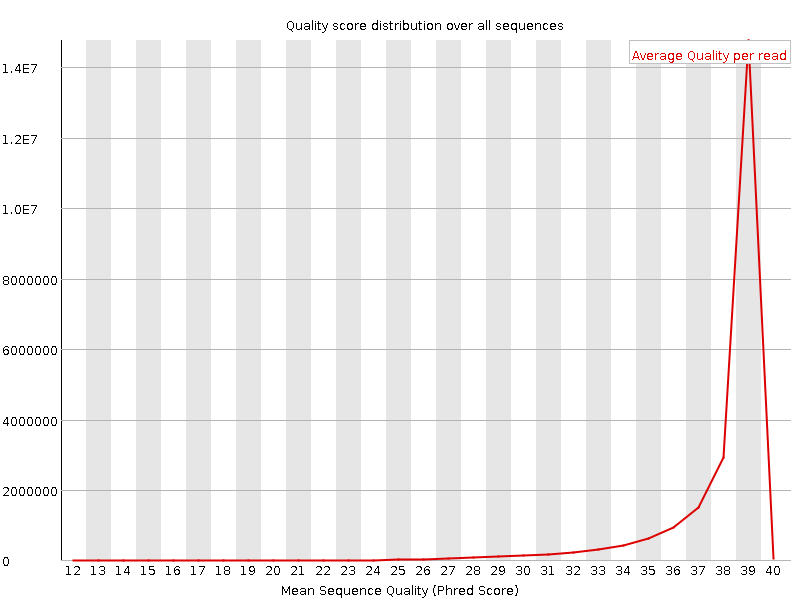
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
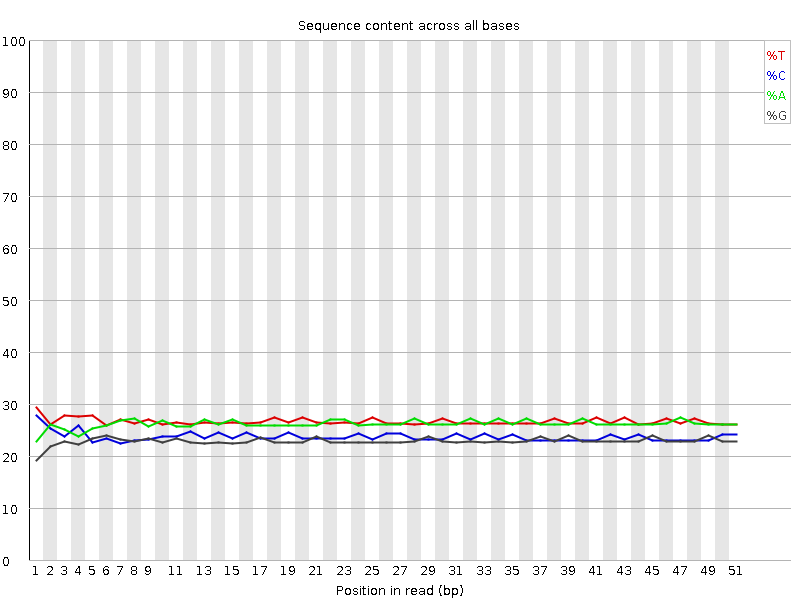
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
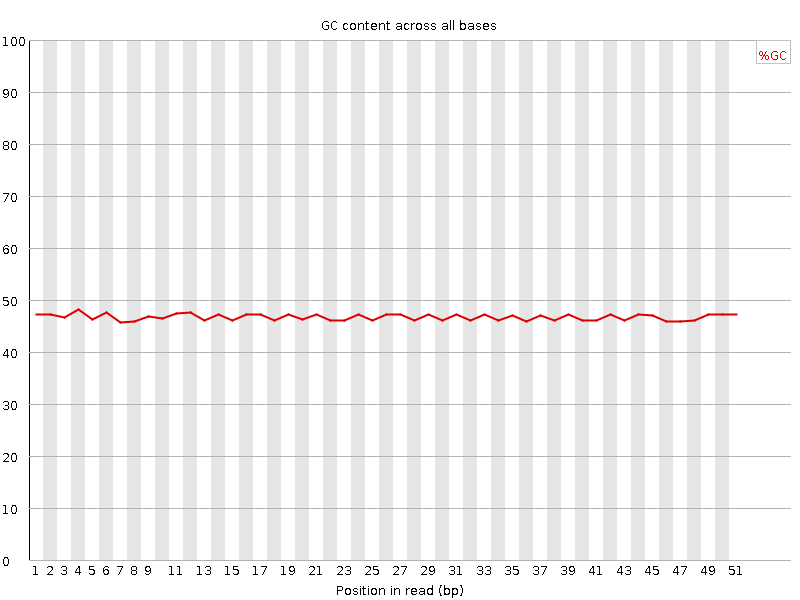
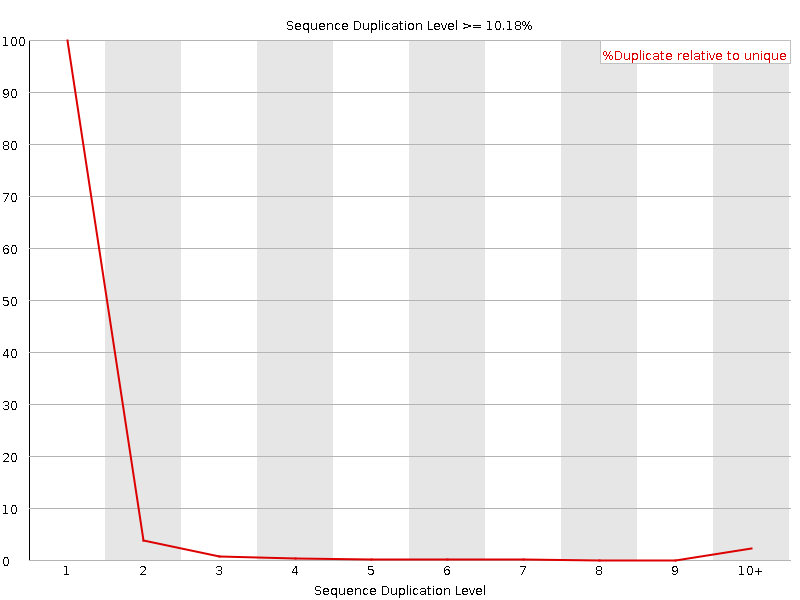
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
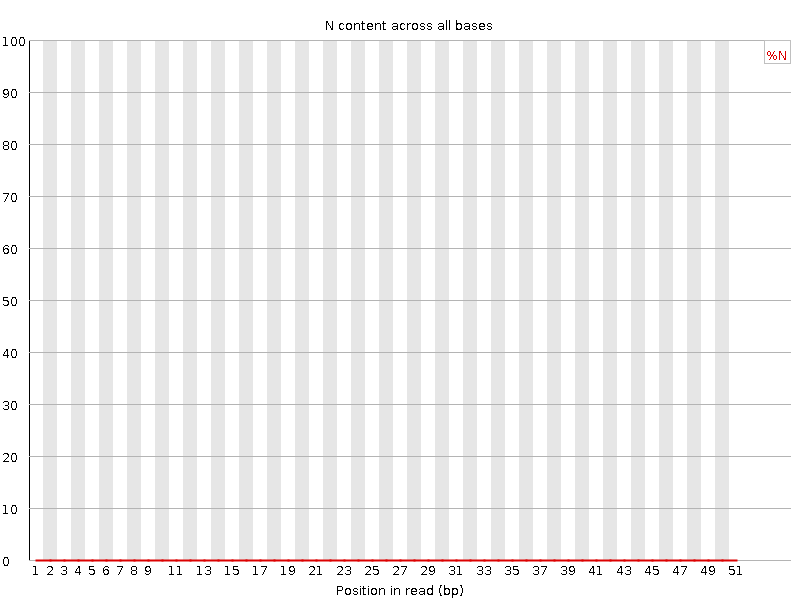
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
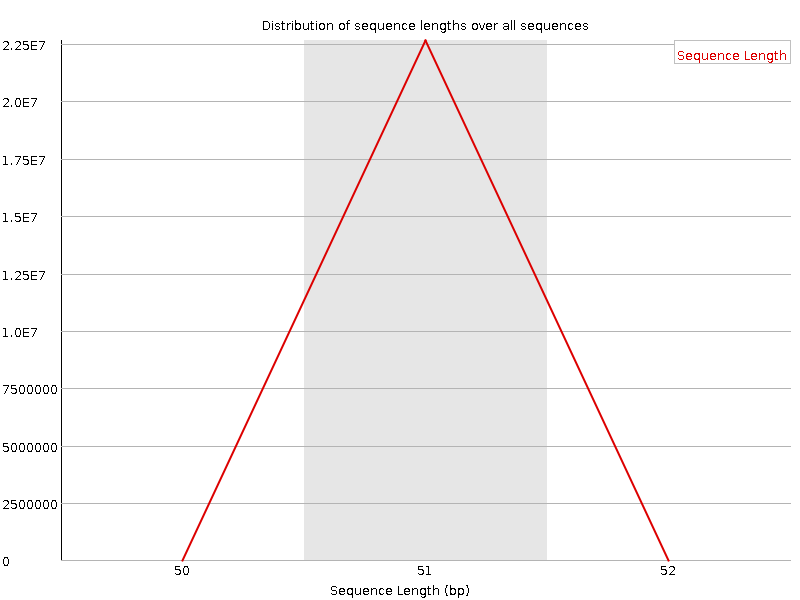
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACACAGTGATCTCGTATGCC | 251246 | 1.1092886355378027 | TruSeq Adapter, Index 5 (100% over 51bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
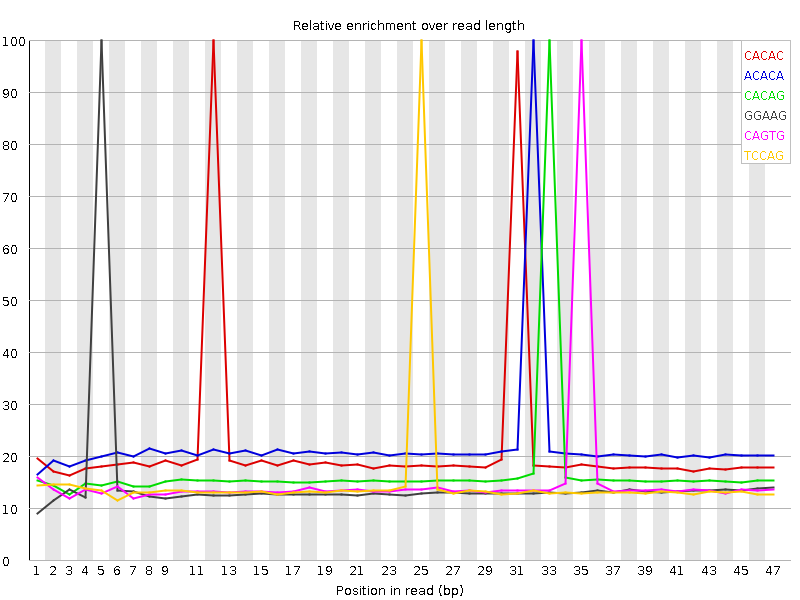
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CACAC | 3293890 | 3.283319 | 15.105426 | 12 |
| ACACA | 3380945 | 3.0518303 | 13.800446 | 32 |
| CACAG | 2447270 | 2.5299625 | 14.799822 | 33 |
| GGAAG | 2129240 | 2.3676279 | 16.005913 | 5 |
| CAGTG | 2135400 | 2.2511246 | 14.627436 | 35 |
| TCCAG | 2137295 | 2.1724832 | 14.32009 | 25 |
| CTCCA | 2059705 | 2.0186858 | 13.717065 | 24 |
| CTGAA | 2140960 | 1.9706923 | 13.075667 | 19 |
| GAAGA | 1967045 | 1.909823 | 13.817488 | 6 |
| AAGAG | 1926095 | 1.8700644 | 13.752465 | 7 |
| AGAGC | 1708565 | 1.831859 | 14.966205 | 8 |
| GCACA | 1691855 | 1.7490225 | 14.345056 | 11 |
| TCTGA | 1930810 | 1.7474692 | 12.665328 | 18 |
| GAGCA | 1608835 | 1.7249322 | 14.828389 | 9 |
| GTCTG | 1600050 | 1.6584924 | 14.175734 | 17 |
| TCACA | 1849180 | 1.6411989 | 12.196201 | 30 |
| AGCAC | 1541455 | 1.5935405 | 14.249826 | 10 |
| ACAGT | 1698165 | 1.5631123 | 12.424166 | 34 |
| AGTGA | 1628735 | 1.5548517 | 12.875232 | 36 |
| CCAGT | 1514825 | 1.539765 | 13.619038 | 26 |
| CAGTC | 1423695 | 1.4471346 | 13.516089 | 27 |
| AGTCA | 1494670 | 1.375801 | 12.318395 | 28 |
| GAACT | 1440805 | 1.3262197 | 12.381555 | 21 |
| AACTC | 1468205 | 1.3030727 | 11.954479 | 22 |
| ACTCC | 1306660 | 1.2806377 | 13.022449 | 23 |
| ATGCC | 1209945 | 1.2298654 | 13.305239 | 47 |
| ATCTC | 1403405 | 1.2246859 | 11.560341 | 40 |
| GTCAC | 1188975 | 1.2085502 | 13.337182 | 29 |
| TGAAC | 1292180 | 1.1894146 | 12.274053 | 20 |
| GTGAT | 1266545 | 1.1888274 | 12.304329 | 37 |
| GATCT | 1159065 | 1.0490055 | 11.7416725 | 39 |
| GTATG | 1058370 | 0.9934266 | 12.122515 | 45 |
| TGATC | 1022135 | 0.9250778 | 11.602369 | 38 |
| TATGC | 976065 | 0.8833824 | 11.628157 | 46 |
| CACGT | 856175 | 0.870271 | 13.182642 | 14 |
| CGGAA | 659350 | 0.7069302 | 13.911282 | 4 |
| TCTCG | 655075 | 0.6547006 | 12.493748 | 41 |
| ACACG | 627410 | 0.64861 | 13.195022 | 13 |
| ACGTC | 622250 | 0.6324947 | 12.953337 | 15 |
| CGTCT | 557780 | 0.55746114 | 12.64512 | 16 |
| TCGGA | 508535 | 0.53609425 | 13.520286 | 3 |
| CTCGT | 522425 | 0.5221264 | 12.362227 | 42 |
| ATCGG | 435050 | 0.45862684 | 13.408113 | 2 |
| GATCG | 428360 | 0.45157427 | 13.352653 | 1 |
| CGTAT | 408875 | 0.37005013 | 11.114205 | 44 |
| TCGTA | 386065 | 0.34940606 | 11.08545 | 43 |