![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | SV37_TGACCA_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 18341618 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 54 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
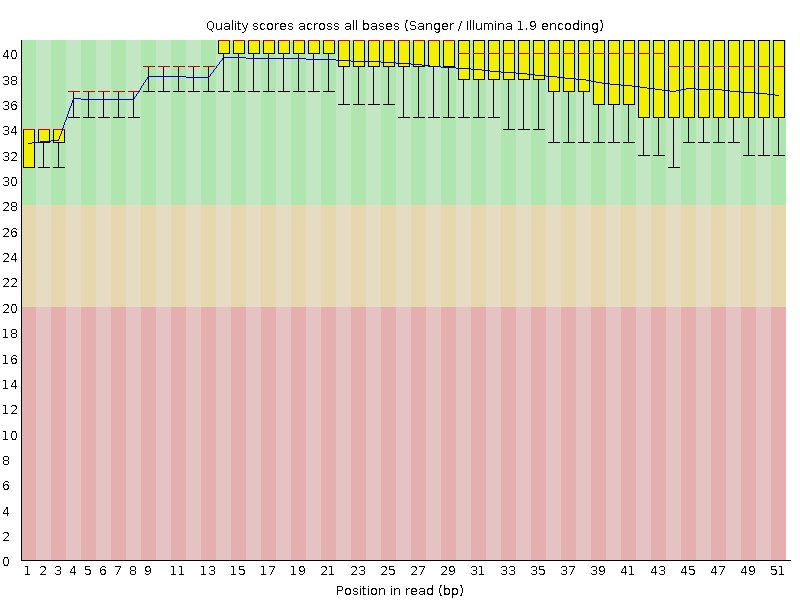
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
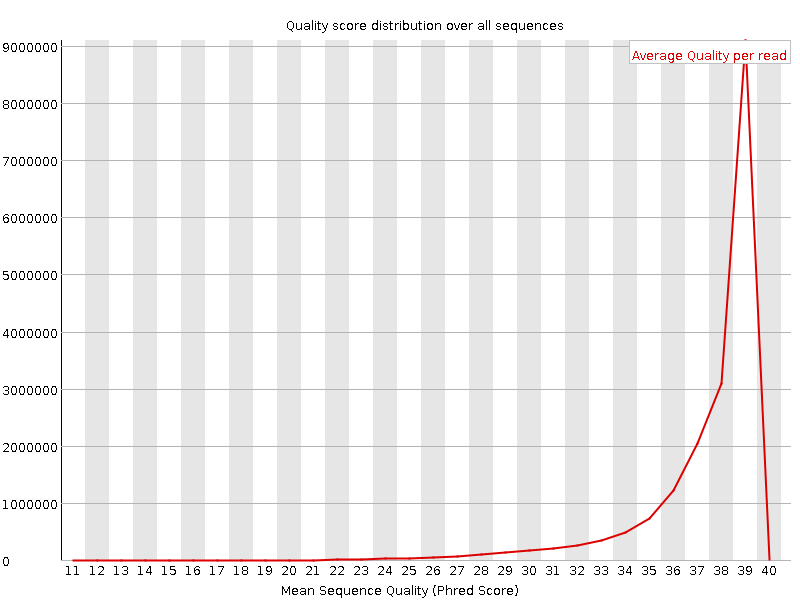
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
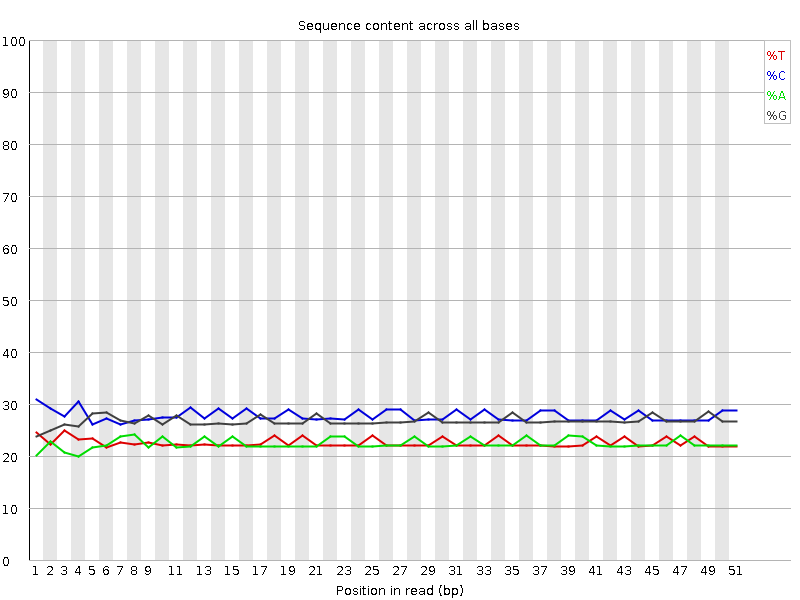
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
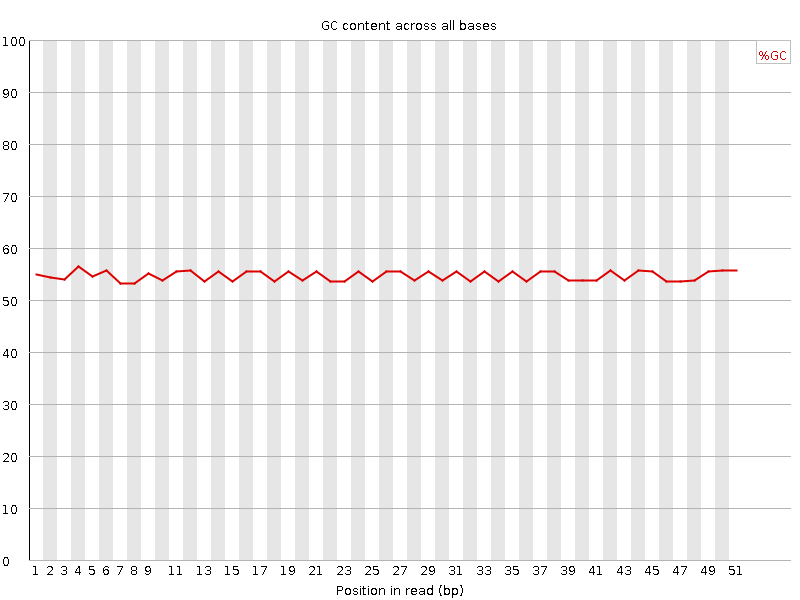
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
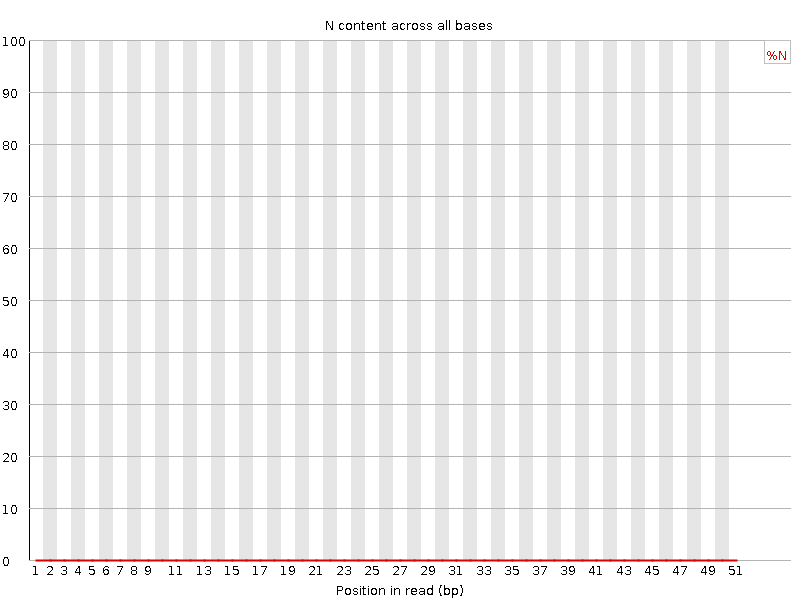
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
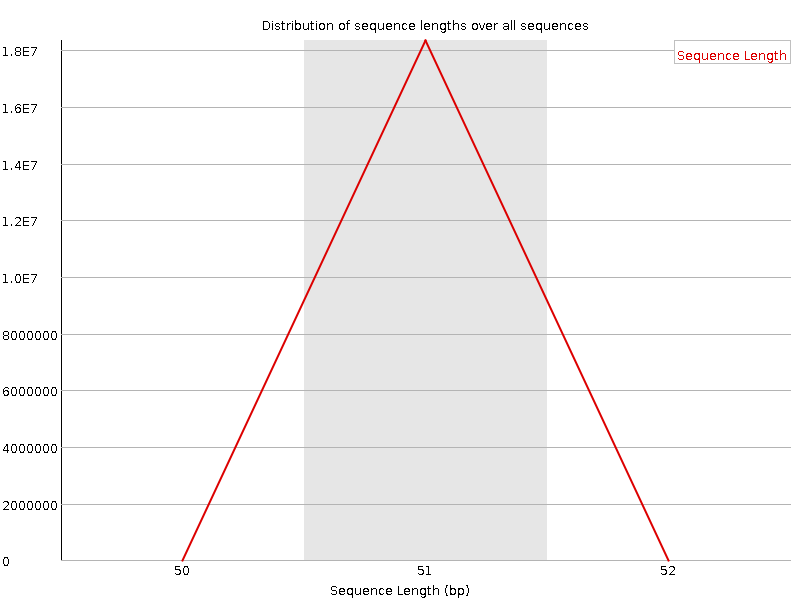
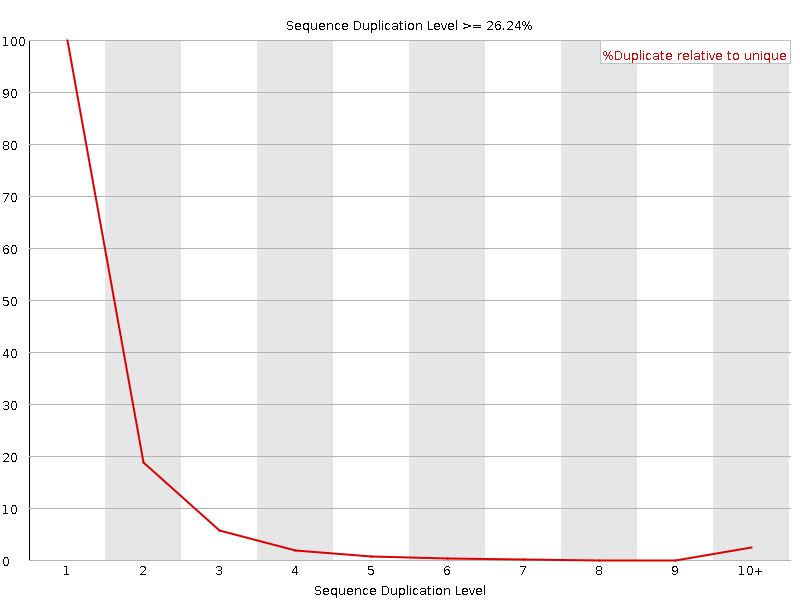
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACTGACCAATCTCGTATGCC | 338475 | 1.8453933562458884 | TruSeq Adapter, Index 4 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
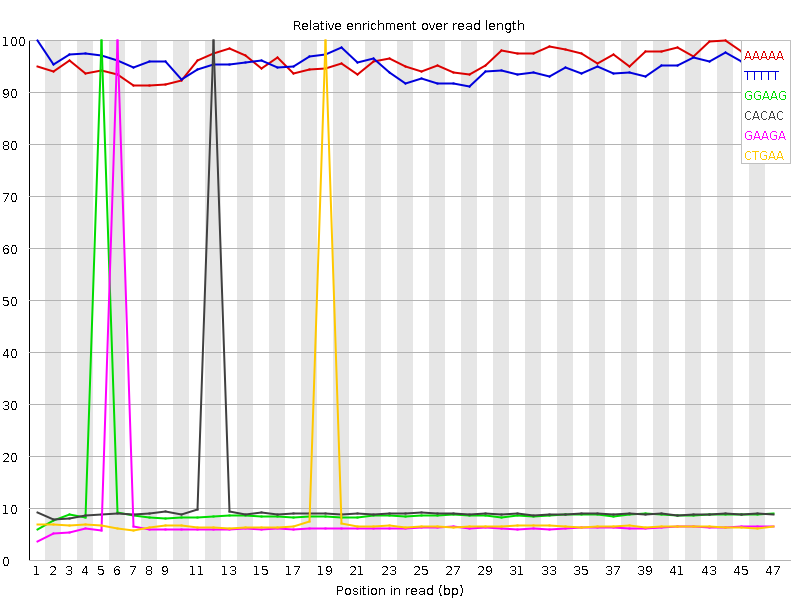
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 2102390 | 4.2326603 | 4.4126506 | 44 |
| TTTTT | 2064800 | 3.9745064 | 4.1769733 | 1 |
| GGAAG | 1962670 | 2.3184721 | 21.92639 | 5 |
| CACAC | 2016235 | 2.1207051 | 19.331879 | 12 |
| GAAGA | 1476840 | 2.0838666 | 25.4321 | 6 |
| CTGAA | 1528475 | 2.0563195 | 23.8482 | 19 |
| AAGAG | 1439795 | 2.031595 | 25.326326 | 7 |
| TCCAG | 1845730 | 1.9999238 | 19.436052 | 25 |
| CTCCA | 1841725 | 1.9198395 | 18.663794 | 24 |
| TCTGA | 1364050 | 1.8187106 | 23.416986 | 18 |
| AGAGC | 1563295 | 1.7766031 | 20.519934 | 8 |
| ACTGA | 1294005 | 1.7408777 | 23.340794 | 32 |
| CACTG | 1551495 | 1.6811082 | 19.038588 | 31 |
| GAGCA | 1426720 | 1.6213926 | 20.340204 | 9 |
| TCACT | 1237325 | 1.5871278 | 22.162539 | 30 |
| GAACT | 1137010 | 1.5296661 | 23.222874 | 21 |
| AATCT | 943875 | 1.5032446 | 26.789421 | 39 |
| GTCTG | 1346700 | 1.5032191 | 19.554626 | 17 |
| GCACA | 1359455 | 1.4863089 | 19.413757 | 11 |
| AACTC | 1135475 | 1.4696189 | 22.320738 | 22 |
| AGTCA | 1079410 | 1.4521743 | 23.030025 | 28 |
| AGCAC | 1327825 | 1.4517277 | 19.45307 | 10 |
| ACCAA | 1069390 | 1.3965684 | 22.16236 | 36 |
| ATCTC | 1080665 | 1.3861786 | 21.762548 | 40 |
| TGAAC | 994295 | 1.3376656 | 23.056581 | 20 |
| CCAGT | 1229055 | 1.3317313 | 18.737879 | 26 |
| CAGTC | 1226795 | 1.3292826 | 18.726677 | 27 |
| CTGAC | 1219510 | 1.3213888 | 18.681097 | 33 |
| ACTCC | 1260325 | 1.3137802 | 18.102596 | 23 |
| GACCA | 1086525 | 1.1879112 | 18.62913 | 35 |
| GTCAC | 1077320 | 1.1673203 | 18.548185 | 29 |
| TGACC | 1064840 | 1.1537976 | 18.456522 | 34 |
| CCAAT | 872755 | 1.1295866 | 21.697245 | 37 |
| CGGAA | 991445 | 1.1267254 | 20.023441 | 4 |
| CAATC | 820920 | 1.0624977 | 21.595661 | 38 |
| GTATG | 755630 | 1.0472435 | 23.121254 | 45 |
| ATGCC | 957365 | 1.0373441 | 18.322216 | 47 |
| TCTCG | 930715 | 0.9994545 | 18.050262 | 41 |
| TATGC | 725645 | 0.9675147 | 22.214014 | 46 |
| CACGT | 885320 | 0.9592804 | 18.642141 | 14 |
| TCGGA | 795385 | 0.8958347 | 19.647532 | 3 |
| CGTCT | 787670 | 0.8458447 | 18.271881 | 16 |
| ACACG | 767545 | 0.8391666 | 18.695675 | 13 |
| ACGTC | 738220 | 0.7998916 | 18.406616 | 15 |
| CTCGT | 731640 | 0.7856765 | 17.841316 | 42 |
| GATCG | 634065 | 0.7141415 | 19.333542 | 1 |
| ATCGG | 607340 | 0.6840413 | 19.367832 | 2 |
| TCGTA | 492055 | 0.6560651 | 21.876953 | 43 |
| CGTAT | 474805 | 0.6330654 | 21.851694 | 44 |