![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | SV35_CAGATC_L001_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 29368635 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
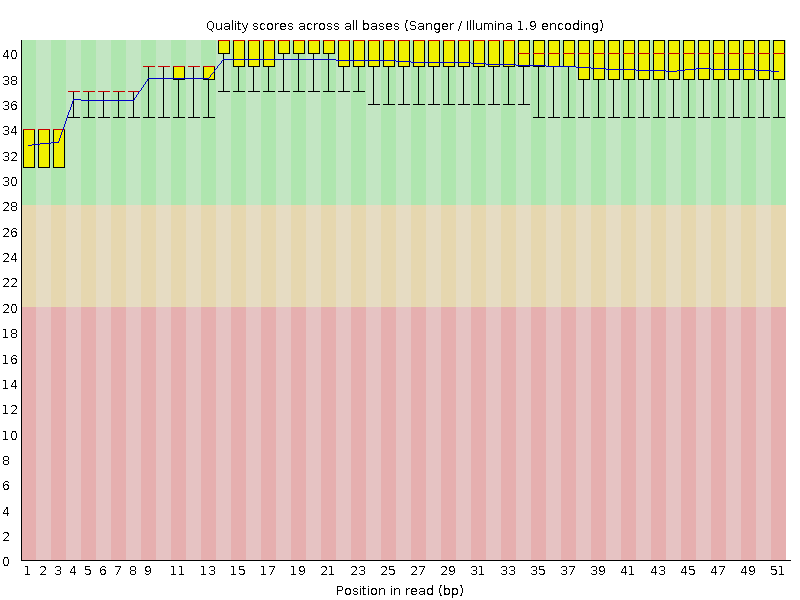
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
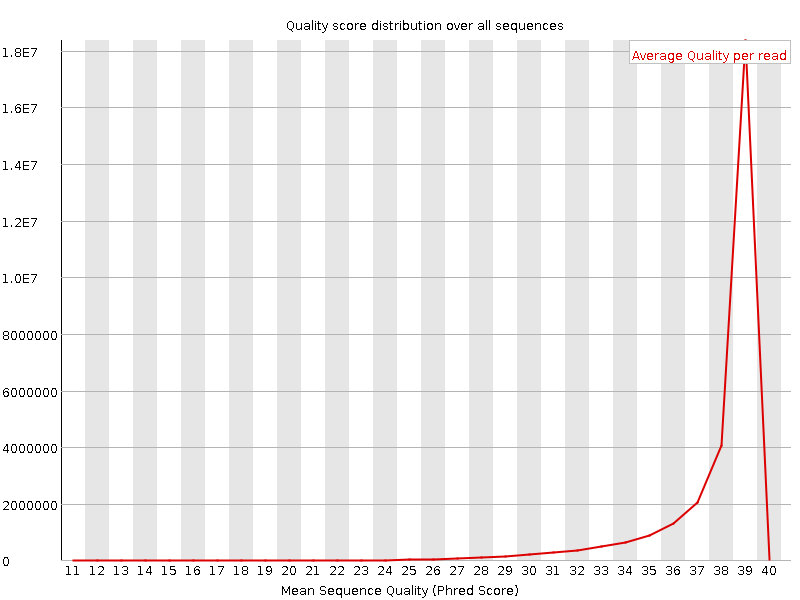
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
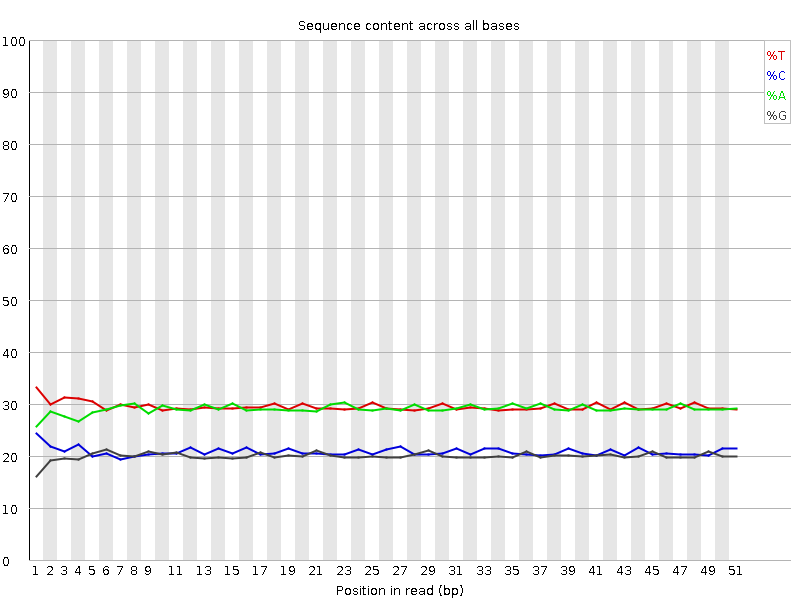
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
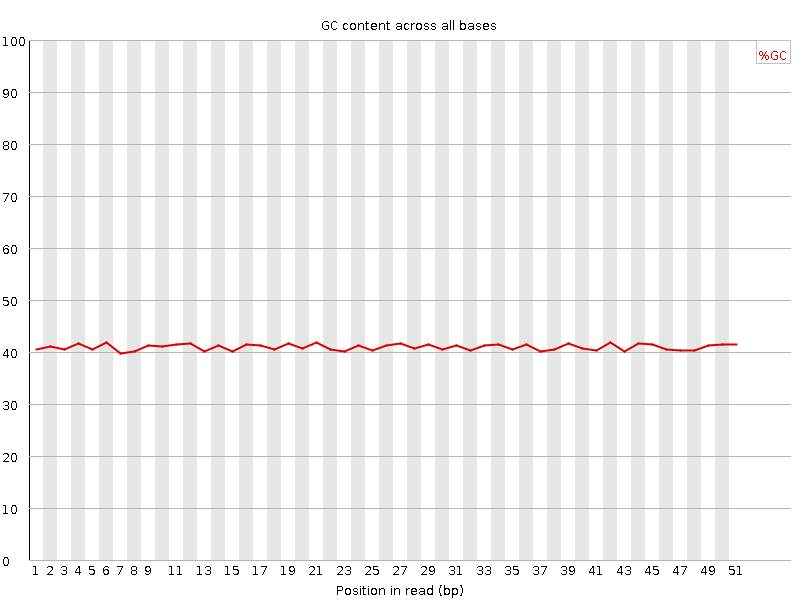
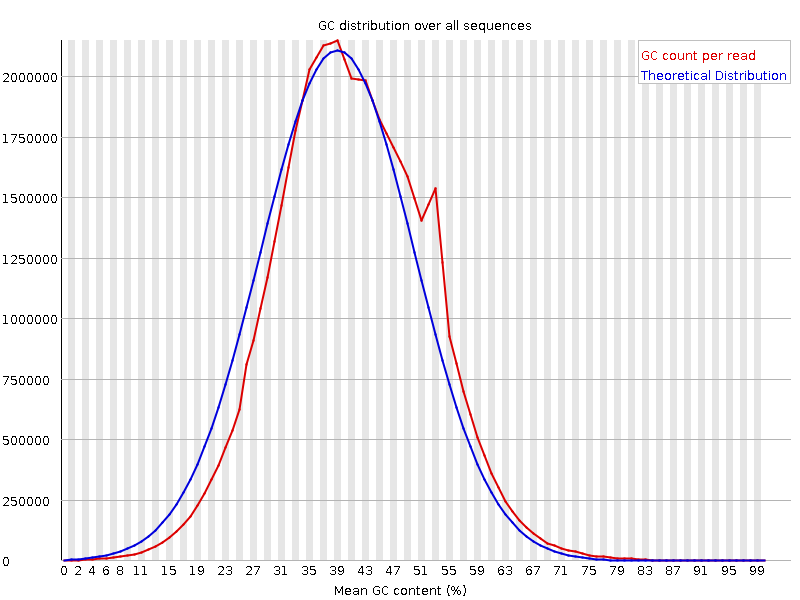
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
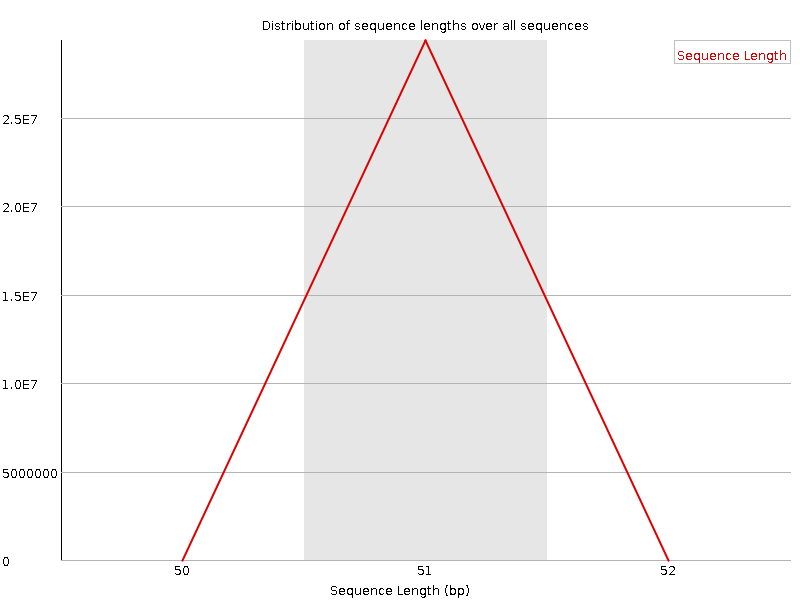
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
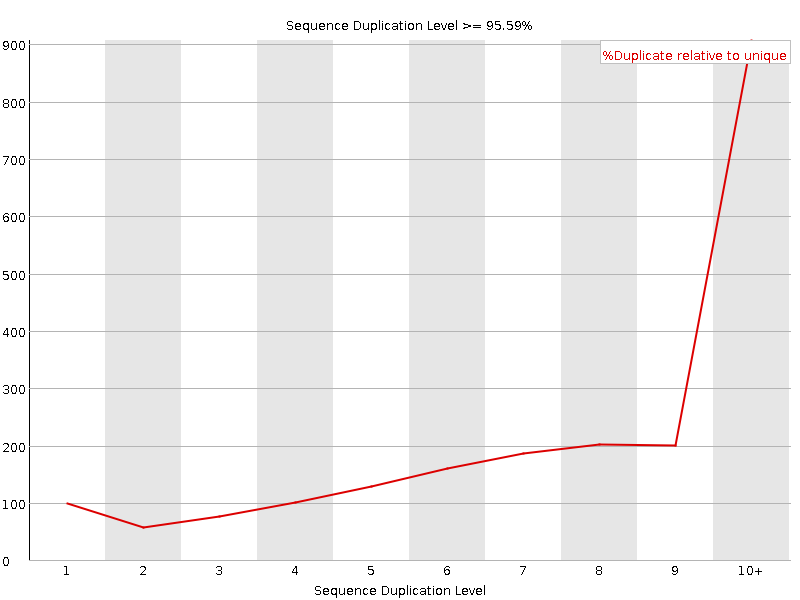
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCAGATCATCTCGTATGCC | 319919 | 1.0893219926632614 | TruSeq Adapter, Index 7 (100% over 51bp) |
| ACTTCCAGGGATTTATAAGCCGATGACGTCATAACATCCCTGACCCTTTAA | 63339 | 0.21566885897148438 | No Hit |
| TATTCGTTGGAATTCCTCGGGGAATTCGGTATTCCCAGGCGGTCTCCCATC | 46402 | 0.15799849056655169 | No Hit |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
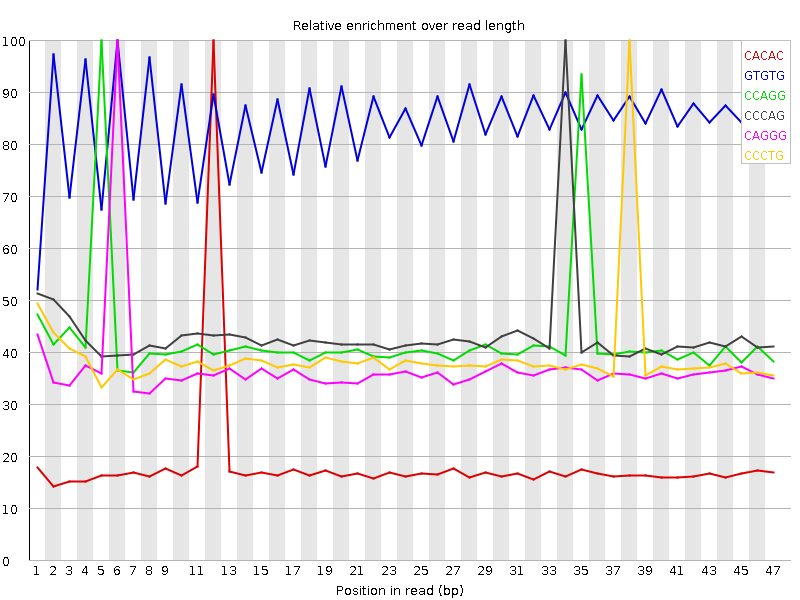
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CACAC | 3554820 | 3.269973 | 17.756695 | 12 |
| GTGTG | 2979945 | 3.000077 | 3.5846097 | 6 |
| CCAGG | 2100400 | 2.918682 | 6.8517795 | 5 |
| CCCAG | 1997250 | 2.6631181 | 6.139478 | 34 |
| CAGGG | 1722745 | 2.4947817 | 6.7160892 | 6 |
| CCCTG | 1835130 | 2.40621 | 6.1448503 | 38 |
| TCCAG | 2509965 | 2.3660812 | 16.973505 | 25 |
| GGAAG | 2215585 | 2.3067224 | 19.192852 | 5 |
| CTCCC | 1715915 | 2.1589155 | 5.4748793 | 44 |
| CCAGA | 2201655 | 2.1105838 | 16.653774 | 33 |
| CAGGC | 1513480 | 2.1031075 | 5.7523127 | 36 |
| CTCCA | 2276655 | 2.0593607 | 16.093704 | 24 |
| GGGGA | 1356095 | 2.046579 | 5.9186006 | 19 |
| CTGAA | 3012790 | 2.041861 | 12.727611 | 19 |
| AGAGC | 1857810 | 1.8560147 | 17.915575 | 8 |
| GCACA | 1895730 | 1.8173133 | 17.082783 | 11 |
| CACCA | 1954580 | 1.79796 | 15.843722 | 31 |
| GAAGA | 2500665 | 1.7961019 | 13.4273205 | 6 |
| GAGCA | 1755960 | 1.754263 | 17.768845 | 9 |
| AAGAG | 2438455 | 1.7514197 | 13.329213 | 7 |
| GTCTG | 1737700 | 1.6786964 | 16.892147 | 17 |
| TCTGA | 2464000 | 1.6421267 | 12.109779 | 18 |
| ACCAG | 1702875 | 1.6324358 | 16.27397 | 32 |
| CCAGT | 1714615 | 1.6163247 | 16.239388 | 26 |
| AGCAC | 1675555 | 1.606246 | 16.979132 | 10 |
| CAGTC | 1570750 | 1.4807067 | 16.092299 | 27 |
| CATCT | 2308845 | 1.4765015 | 11.205574 | 39 |
| ATGCC | 1460410 | 1.3766918 | 15.780189 | 47 |
| CAGAT | 2028260 | 1.3746146 | 11.678244 | 34 |
| ACTCC | 1474165 | 1.3334639 | 15.4292345 | 23 |
| TCACC | 1467135 | 1.327105 | 15.192867 | 30 |
| AGTCA | 1856485 | 1.2581973 | 11.73255 | 28 |
| GTCAC | 1329165 | 1.2529705 | 15.818056 | 29 |
| AACTC | 1913420 | 1.2443452 | 11.434317 | 22 |
| GAACT | 1752270 | 1.1875677 | 11.845467 | 21 |
| ATCTC | 1837980 | 1.1753843 | 10.954461 | 40 |
| TCATC | 1739645 | 1.1124994 | 10.8123 | 38 |
| GACCC | 825930 | 1.1012889 | 5.079583 | 42 |
| TGAAC | 1623095 | 1.1000216 | 11.75596 | 20 |
| CACGT | 1155940 | 1.0896757 | 16.07545 | 14 |
| AGATC | 1456570 | 0.98716253 | 11.241073 | 35 |
| GTATG | 1394605 | 0.96859866 | 11.589314 | 45 |
| ATCAT | 2095185 | 0.96328956 | 7.975155 | 37 |
| TATGC | 1330025 | 0.8863918 | 11.080639 | 46 |
| GATCA | 1295185 | 0.8777869 | 11.170838 | 36 |
| ACGTC | 910060 | 0.8578908 | 15.742267 | 15 |
| CGGGG | 358685 | 0.7529328 | 6.2281513 | 18 |
| CGGAA | 741630 | 0.7409133 | 16.97433 | 4 |
| TCTCG | 786050 | 0.728653 | 14.927383 | 41 |
| GGCGG | 326735 | 0.68586504 | 6.2686553 | 38 |
| ACACG | 671445 | 0.64367074 | 15.834199 | 13 |
| CGTCT | 612470 | 0.5677478 | 15.218372 | 16 |
| CTCGT | 599245 | 0.55548847 | 14.742528 | 42 |
| TCGGA | 543405 | 0.53384155 | 16.49211 | 3 |
| ATCGG | 494890 | 0.48618034 | 16.427753 | 2 |
| GATCG | 479505 | 0.47106612 | 16.355238 | 1 |
| CGTAT | 542310 | 0.3614211 | 10.592573 | 44 |
| TCGTA | 493585 | 0.3289485 | 10.548565 | 43 |