![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | SV34_GCCAAT_L001_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 28532577 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 44 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
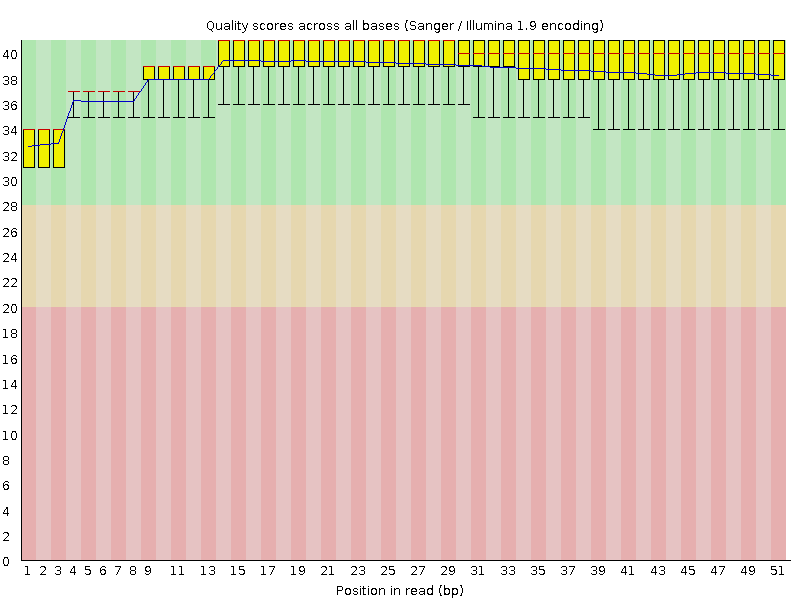
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
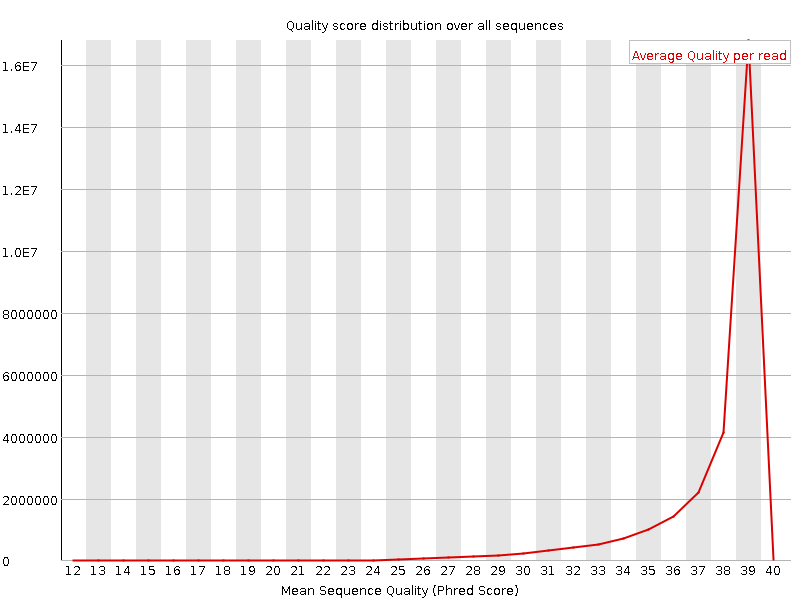
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
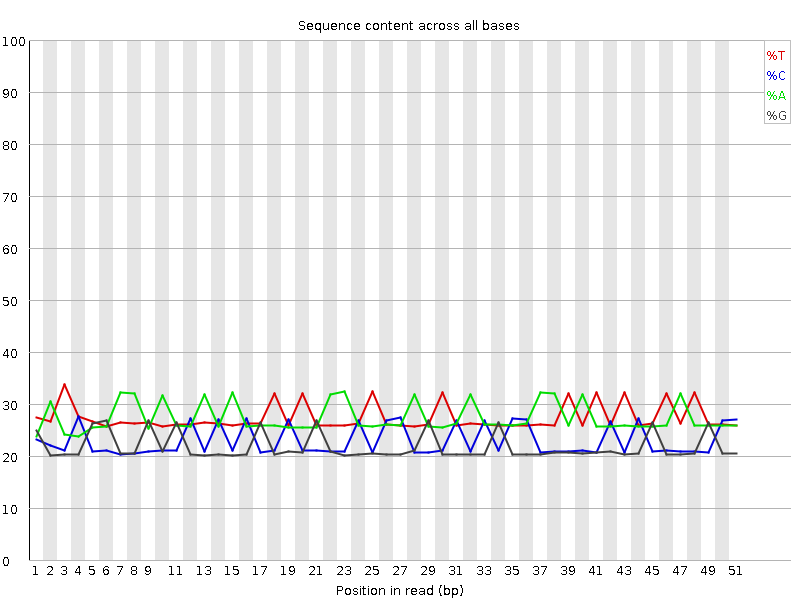
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
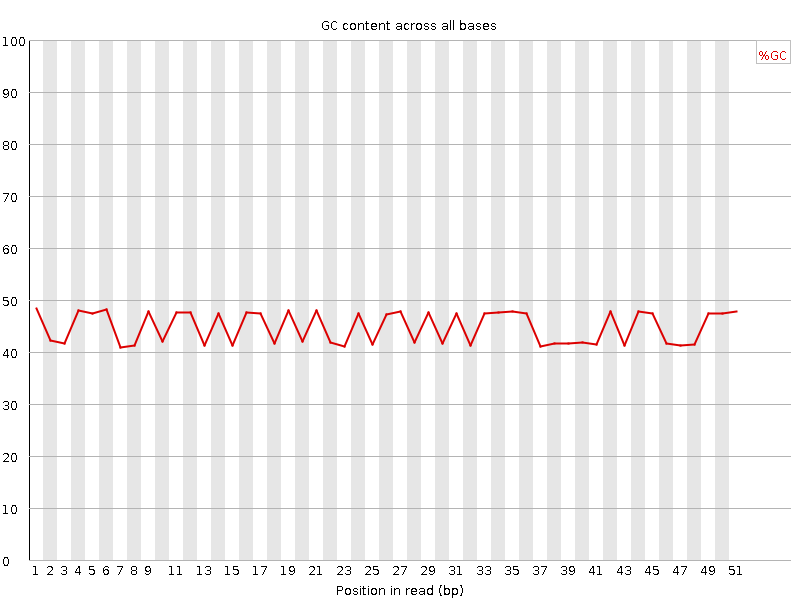
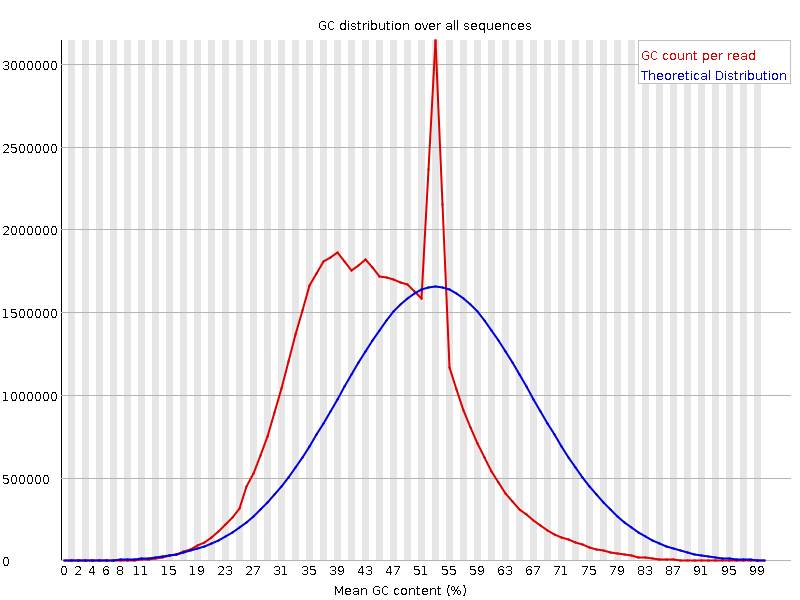
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
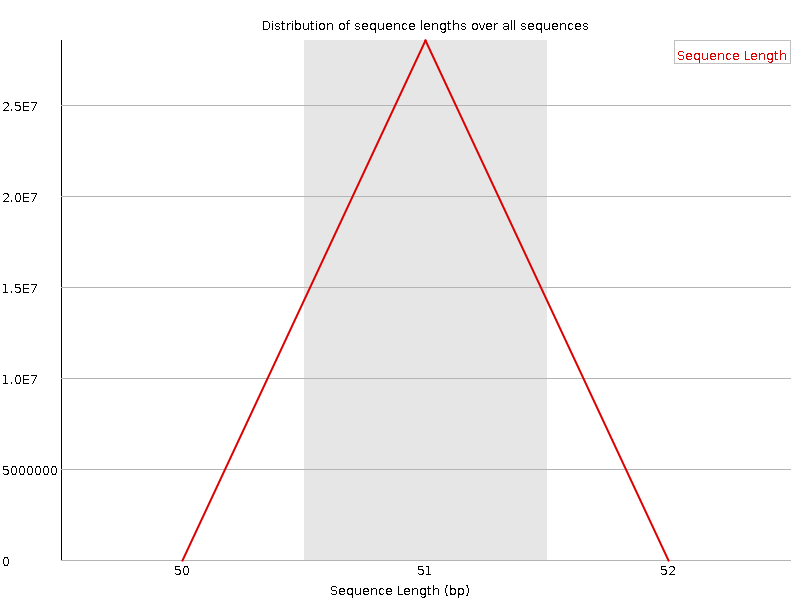
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
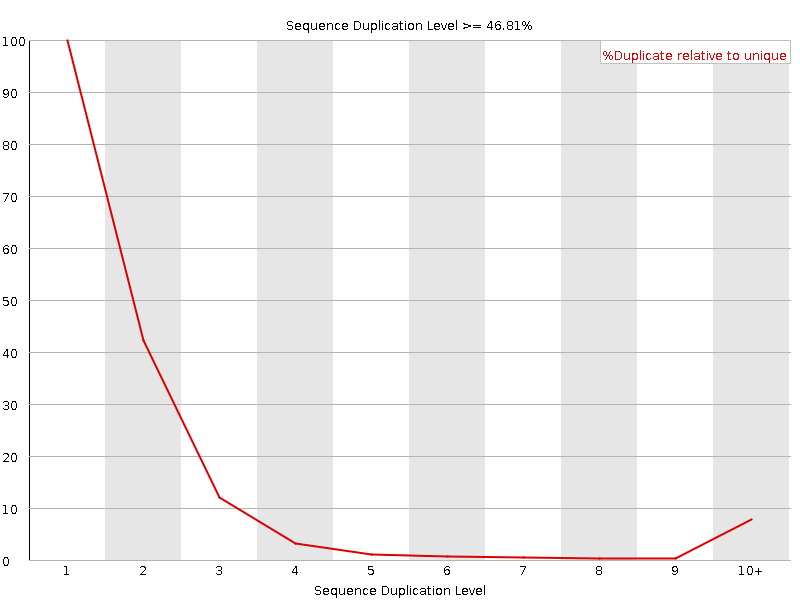
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGCCAATATCTCGTATGCC | 1676239 | 5.874825116567634 | TruSeq Adapter, Index 6 (100% over 51bp) |
| ACTTCCAGGGATTTATAAGCCGATGACGTCATAACATCCCTGACCCTTTAA | 80538 | 0.28226682784383617 | No Hit |
| TATTCGTTGGAATTCCTCGGGGAATTCGGTATTCCCAGGCGGTCTCCCATC | 59834 | 0.20970415676088422 | No Hit |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
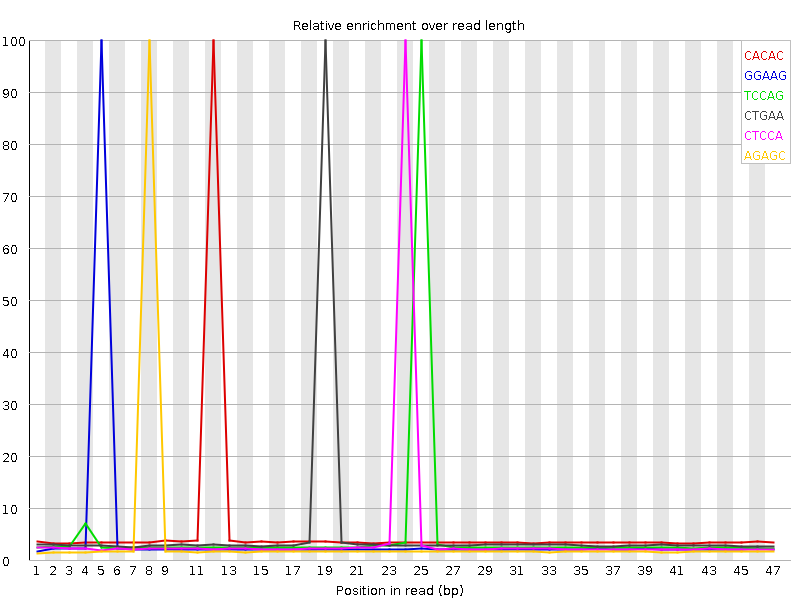
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CACAC | 4924435 | 3.9533322 | 70.12864 | 12 |
| GGAAG | 3777630 | 3.5418096 | 81.691 | 5 |
| TCCAG | 4081090 | 3.45047 | 71.98228 | 25 |
| CTGAA | 4389590 | 3.1098325 | 61.057396 | 19 |
| CTCCA | 3774405 | 3.0302906 | 68.22253 | 24 |
| AGAGC | 3323040 | 2.9585238 | 76.83801 | 8 |
| GAAGA | 3861290 | 2.8805995 | 64.973816 | 6 |
| GAGCA | 3231970 | 2.8774438 | 76.63215 | 9 |
| GCACA | 3367940 | 2.8473284 | 72.70277 | 11 |
| GTCTG | 3185125 | 2.8361144 | 75.80774 | 17 |
| AAGAG | 3724180 | 2.7783127 | 64.73795 | 7 |
| CCAGT | 3174175 | 2.683694 | 71.18995 | 26 |
| AGCAC | 3102580 | 2.6229873 | 72.62845 | 10 |
| TCTGA | 3672770 | 2.6021698 | 60.655716 | 18 |
| CAGTC | 3031480 | 2.5630481 | 71.063774 | 27 |
| CCAGG | 2407050 | 2.5574884 | 6.4306087 | 5 |
| CGCCA | 2531695 | 2.55431 | 83.07075 | 33 |
| ATGCC | 2871850 | 2.4280846 | 70.59932 | 47 |
| ACTCC | 2942535 | 2.3624213 | 67.72049 | 23 |
| GCCAA | 2786970 | 2.356164 | 69.740906 | 34 |
| GTCAC | 2766990 | 2.339428 | 70.73238 | 29 |
| CACGT | 2759185 | 2.332829 | 71.92244 | 14 |
| TCACG | 2679260 | 2.2652543 | 70.22277 | 30 |
| AGTCA | 3160680 | 2.2392037 | 59.613342 | 28 |
| CCCAG | 2215190 | 2.2349777 | 5.7783647 | 34 |
| GAACT | 3121665 | 2.211563 | 60.0372 | 21 |
| CACGC | 2180740 | 2.20022 | 83.154915 | 31 |
| AACTC | 3208350 | 2.158383 | 57.06558 | 22 |
| CAGGG | 1892860 | 2.1179385 | 6.2631536 | 6 |
| ATCTC | 3139405 | 2.1121411 | 56.187496 | 40 |
| ACGTC | 2495950 | 2.1102695 | 71.51194 | 15 |
| TGAAC | 2950760 | 2.0904844 | 59.972107 | 20 |
| ACGCC | 2066600 | 2.0850604 | 82.92755 | 32 |
| CGGAA | 2296180 | 2.044304 | 76.36502 | 4 |
| GTATG | 2732925 | 2.0390878 | 62.20599 | 45 |
| CCCTG | 1996645 | 2.0146148 | 5.7503295 | 38 |
| TCTCG | 2328815 | 1.9690917 | 69.969185 | 41 |
| AATAT | 4063280 | 1.9194387 | 39.53125 | 37 |
| TATGC | 2651930 | 1.8789011 | 58.993874 | 46 |
| CCAAT | 2783805 | 1.8727747 | 55.591568 | 35 |
| ACACG | 2169470 | 1.8341163 | 71.45614 | 13 |
| TCGGA | 2047355 | 1.8228946 | 76.223564 | 3 |
| CTCCC | 1900850 | 1.8212631 | 5.1590514 | 44 |
| GGGGA | 1517855 | 1.7885104 | 5.8859253 | 19 |
| CAGGC | 1675165 | 1.7798612 | 5.5528364 | 36 |
| TATCT | 3156450 | 1.7795669 | 46.86222 | 39 |
| CGTCT | 2103085 | 1.7782295 | 71.194084 | 16 |
| CTCGT | 2086175 | 1.7639315 | 69.794556 | 42 |
| ATCGG | 1980805 | 1.7636406 | 76.15033 | 2 |
| AGGGA | 1878035 | 1.7607977 | 5.2352858 | 7 |
| GATCG | 1970390 | 1.7543676 | 76.03596 | 1 |
| CTTCC | 2153825 | 1.7293187 | 5.0001607 | 2 |
| CAATA | 2816275 | 1.5875696 | 46.51569 | 36 |
| ATATC | 2791260 | 1.5735728 | 46.547127 | 38 |
| CGTAT | 1995580 | 1.4138751 | 58.545315 | 44 |
| TCGTA | 1952715 | 1.3835051 | 58.478886 | 43 |
| CGGGG | 574575 | 0.80797315 | 5.7639866 | 18 |
| GGCGG | 559675 | 0.78702056 | 5.7901244 | 38 |