![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | SV33_ACAGTG_L001_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 26524278 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 46 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
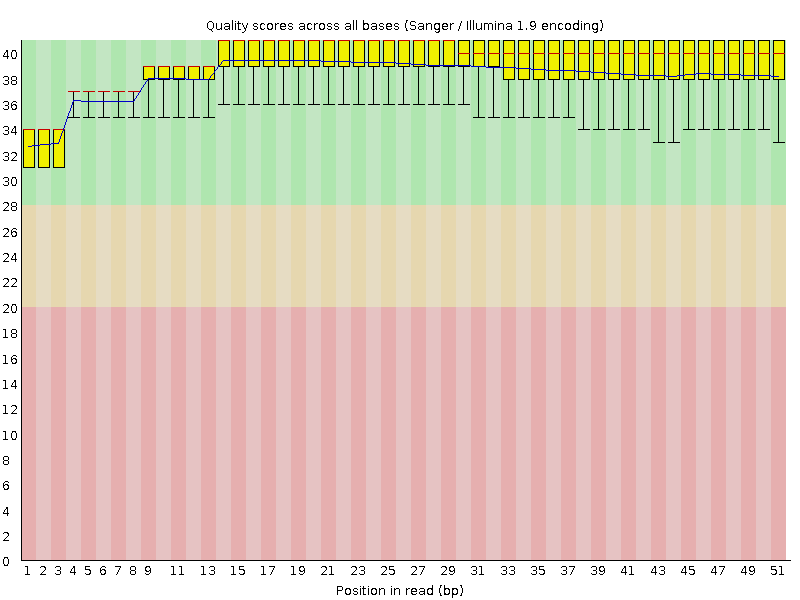
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
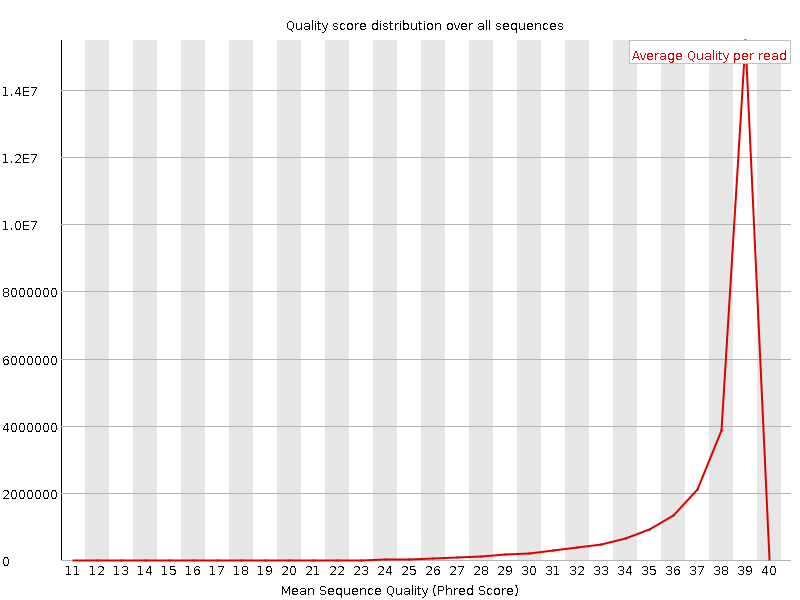
![[OK]](Icons/tick.png) Per base sequence content
Per base sequence content
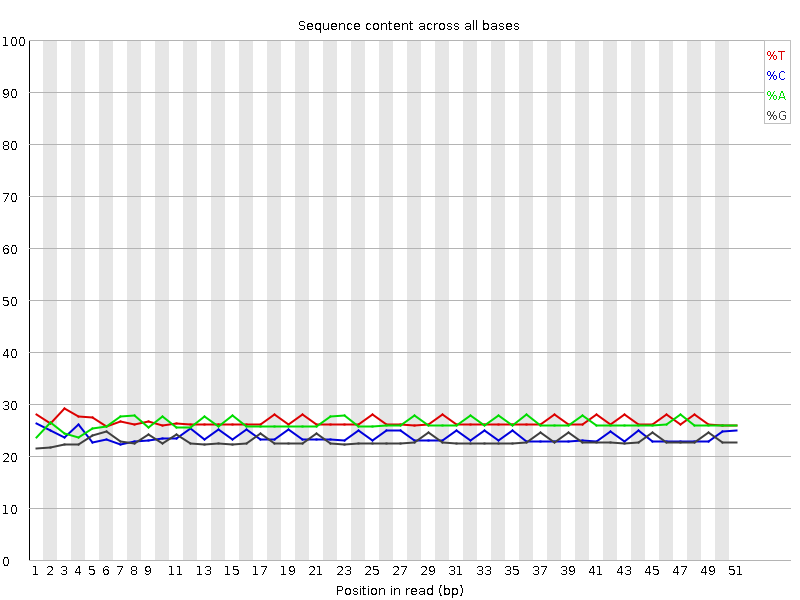
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
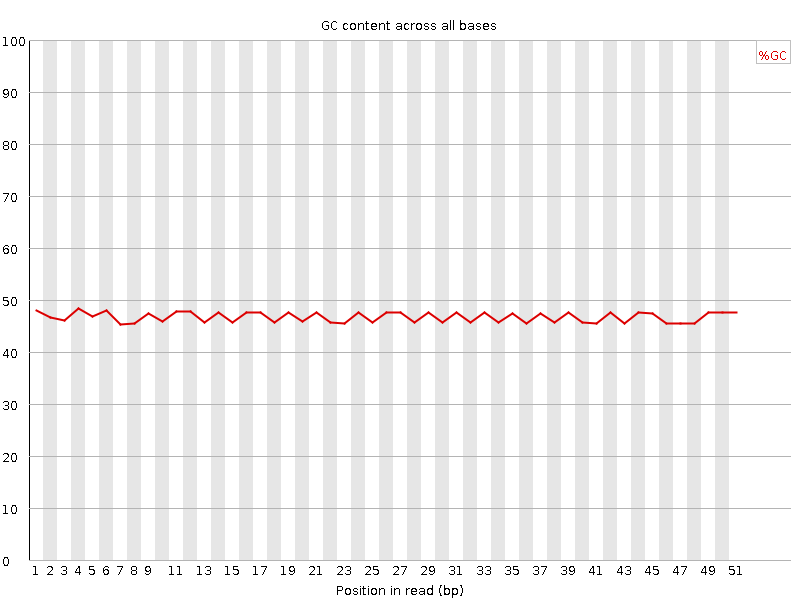
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
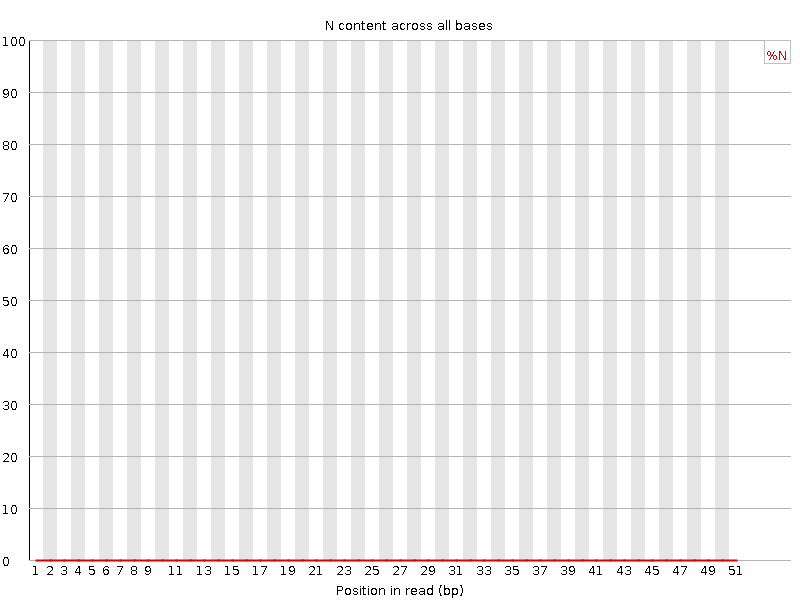
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
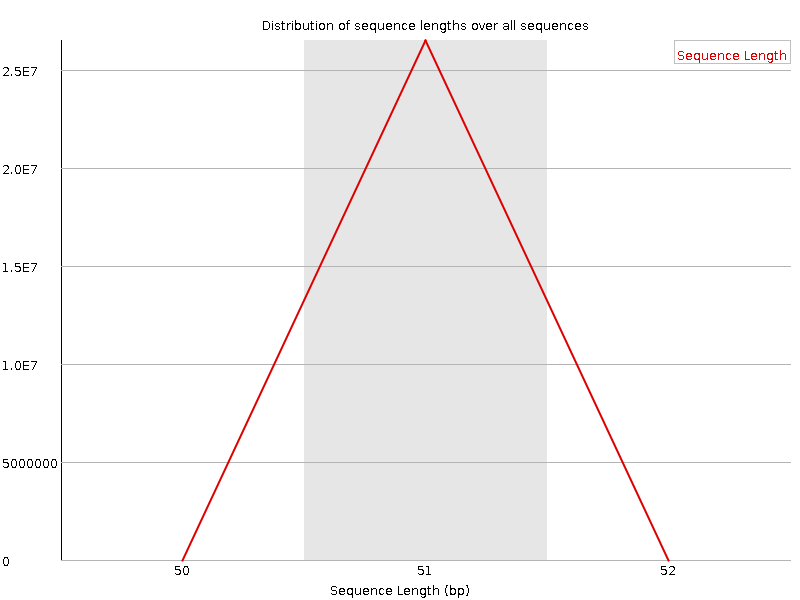
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
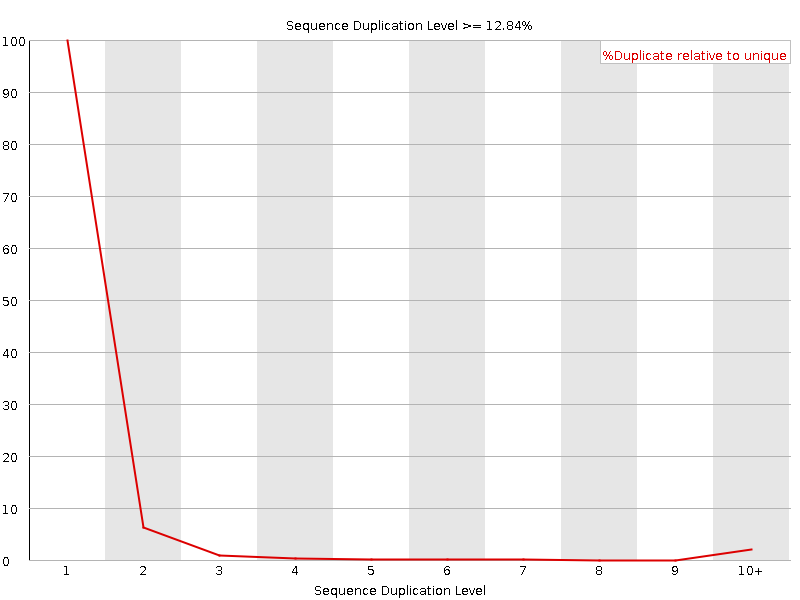
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACACAGTGATCTCGTATGCC | 499165 | 1.881917389042597 | TruSeq Adapter, Index 5 (100% over 51bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
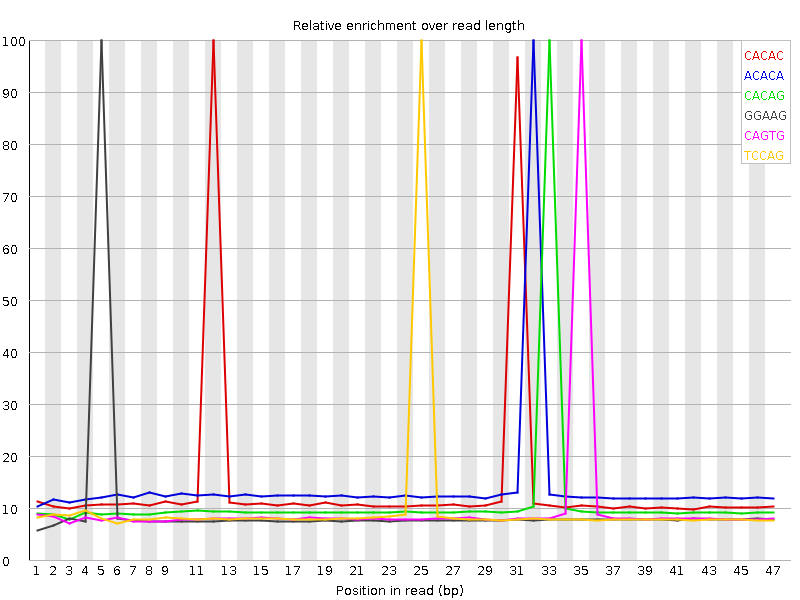
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CACAC | 4075100 | 3.4556932 | 23.990685 | 12 |
| ACACA | 3946095 | 3.0293443 | 21.351763 | 32 |
| CACAG | 3023850 | 2.6598623 | 23.622 | 33 |
| GGAAG | 2698025 | 2.5535765 | 26.216307 | 5 |
| CAGTG | 2647200 | 2.3815393 | 23.615805 | 35 |
| TCCAG | 2718525 | 2.3577752 | 23.397112 | 25 |
| CTCCA | 2645135 | 2.2116423 | 22.48026 | 24 |
| GAAGA | 2533400 | 2.0926127 | 22.72282 | 6 |
| CTGAA | 2661965 | 2.0900445 | 21.274727 | 19 |
| AAGAG | 2487540 | 2.0547318 | 22.649513 | 7 |
| AGAGC | 2246485 | 2.0497665 | 24.795954 | 8 |
| GAGCA | 2113775 | 1.9286777 | 24.591618 | 9 |
| TCTGA | 2469980 | 1.9121274 | 20.84612 | 18 |
| GCACA | 2152450 | 1.8933548 | 23.683756 | 11 |
| GTCTG | 2062235 | 1.8292763 | 23.552464 | 17 |
| TCACA | 2362025 | 1.787869 | 19.871096 | 30 |
| AGCAC | 1990675 | 1.751053 | 23.59853 | 10 |
| CCAGT | 2003050 | 1.7372439 | 22.706 | 26 |
| AGTGA | 2092940 | 1.704559 | 20.984344 | 36 |
| ACAGT | 2145695 | 1.6846946 | 20.264755 | 34 |
| CAGTC | 1883405 | 1.633476 | 22.578825 | 27 |
| AGTCA | 1961480 | 1.5400583 | 20.476751 | 28 |
| GAACT | 1881320 | 1.4771203 | 20.64963 | 21 |
| AACTC | 1934965 | 1.4646178 | 19.928726 | 22 |
| ACTCC | 1750890 | 1.463949 | 21.831879 | 23 |
| ATGCC | 1639040 | 1.4215384 | 22.268293 | 47 |
| GTCAC | 1616985 | 1.40241 | 22.317745 | 29 |
| ATCTC | 1869345 | 1.3951175 | 19.31232 | 40 |
| TGAAC | 1710110 | 1.3426946 | 20.54391 | 20 |
| GTGAT | 1659100 | 1.3322874 | 20.328472 | 37 |
| GATCT | 1573655 | 1.2182401 | 19.700016 | 39 |
| GTATG | 1431885 | 1.1498296 | 20.490898 | 45 |
| TGATC | 1404685 | 1.0874325 | 19.384748 | 38 |
| TATGC | 1359810 | 1.0526927 | 19.689964 | 46 |
| CACGT | 1174640 | 1.0187646 | 22.407228 | 14 |
| CGGAA | 966590 | 0.88194853 | 23.79601 | 4 |
| TCTCG | 981210 | 0.8390757 | 21.376343 | 41 |
| ACACG | 943105 | 0.8295813 | 22.516956 | 13 |
| ACGTC | 932100 | 0.80840975 | 22.159386 | 15 |
| CGTCT | 880690 | 0.7531166 | 21.753164 | 16 |
| TCGGA | 820835 | 0.7384599 | 23.32436 | 3 |
| CTCGT | 839470 | 0.7178676 | 21.265642 | 42 |
| ATCGG | 730995 | 0.6576358 | 23.206846 | 2 |
| GATCG | 722590 | 0.6500743 | 23.146025 | 1 |
| CGTAT | 704185 | 0.5451426 | 19.183174 | 44 |
| TCGTA | 679895 | 0.52633864 | 19.171896 | 43 |