![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | SV32_TGACCA_L001_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 31356637 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 53 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
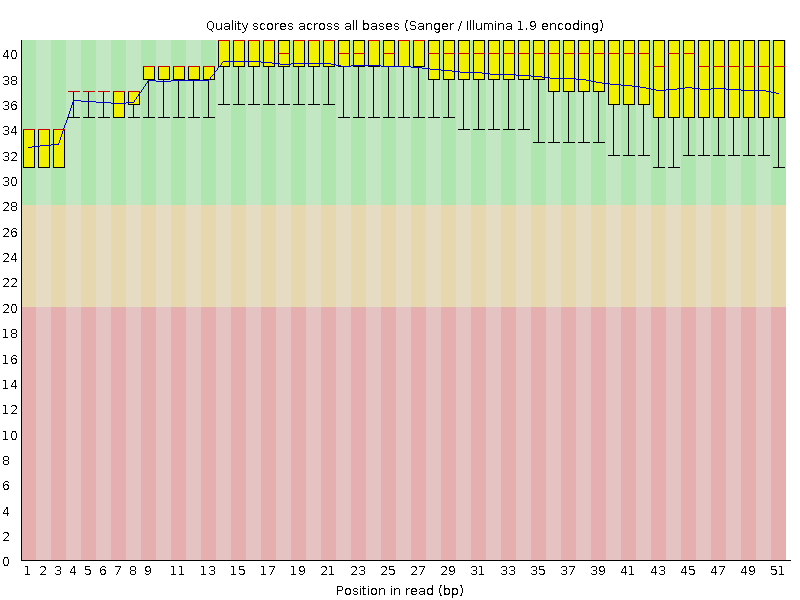
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
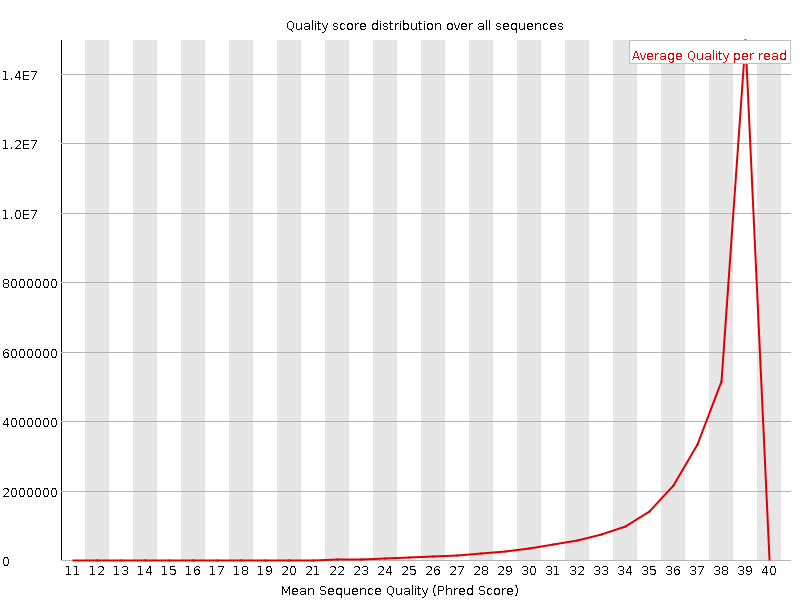
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
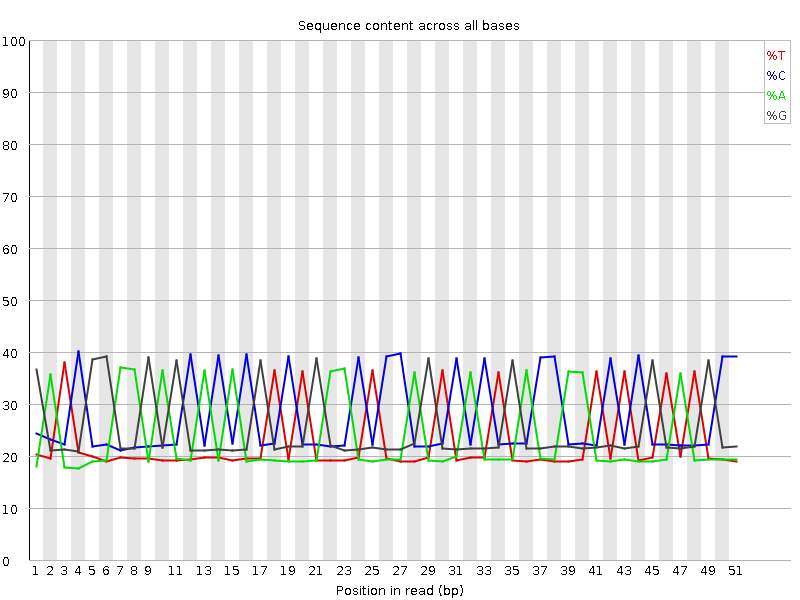
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
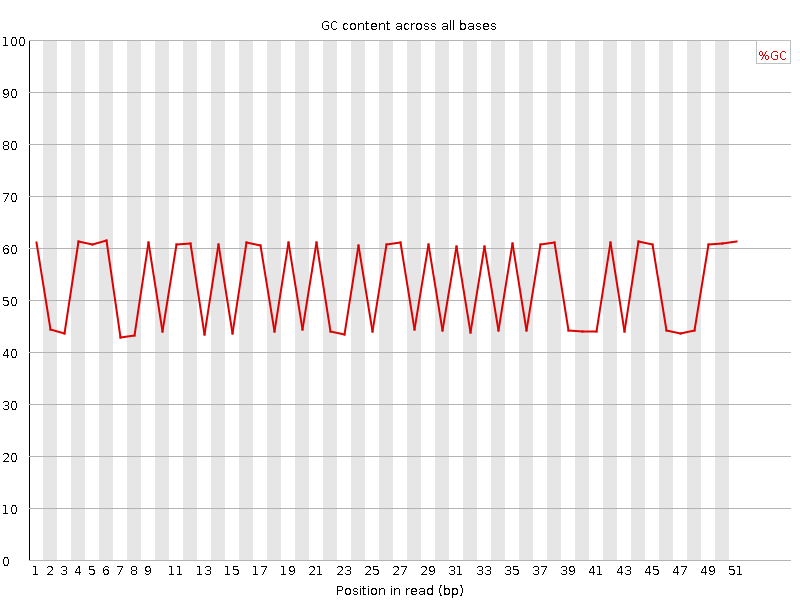
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
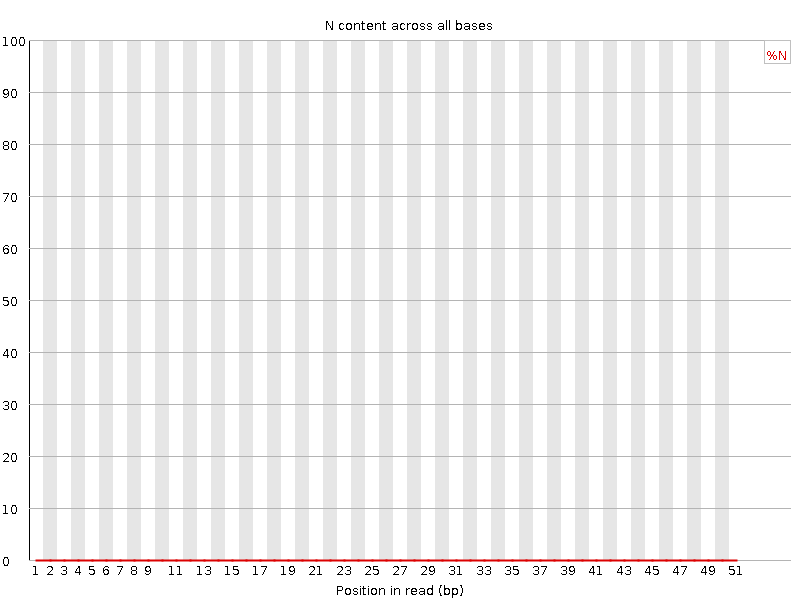
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
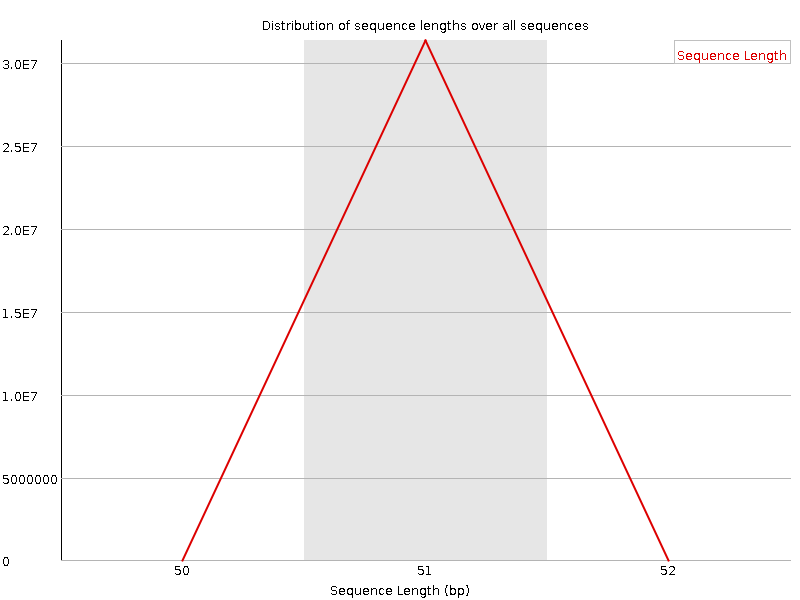
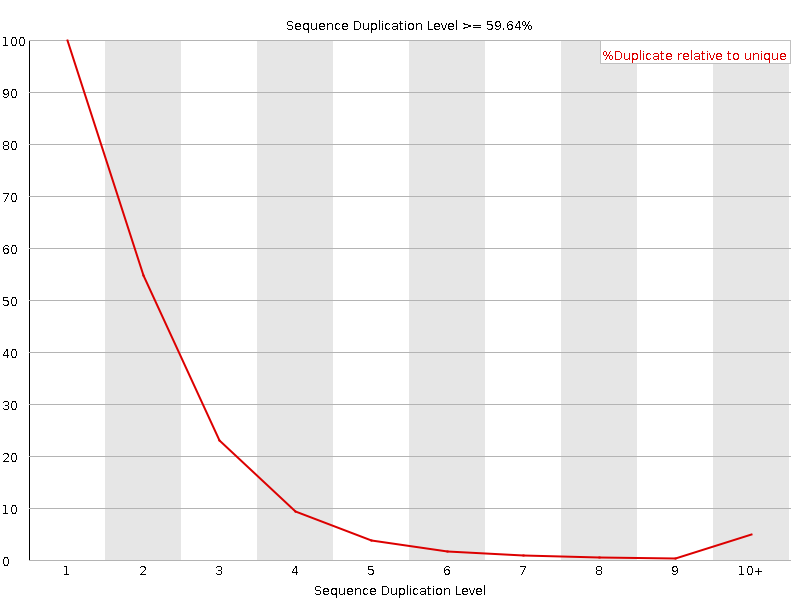
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACTGACCAATCTCGTATGCC | 5165245 | 16.472573254587218 | TruSeq Adapter, Index 4 (100% over 51bp) |
| ACTTCCAGGGATTTATAAGCCGATGACGTCATAACATCCCTGACCCTTTAA | 108763 | 0.34685798735368206 | No Hit |
| TATTCGTTGGAATTCCTCGGGGAATTCGGTATTCCCAGGCGGTCTCCCATC | 78588 | 0.25062636659664744 | No Hit |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
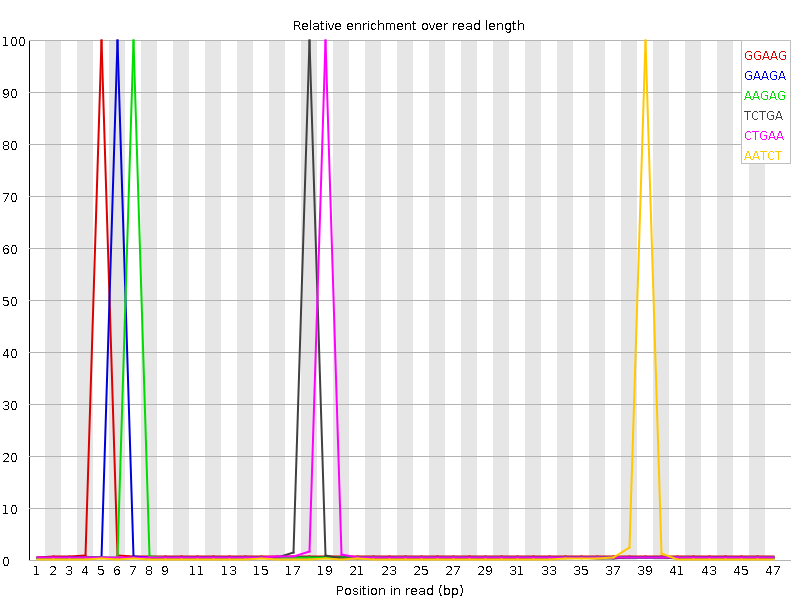
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGAAG | 7782440 | 5.6339736 | 189.18048 | 5 |
| GAAGA | 7180570 | 5.4864736 | 198.95348 | 6 |
| AAGAG | 7134335 | 5.4511466 | 198.58029 | 7 |
| TCTGA | 6992480 | 5.348 | 195.41475 | 18 |
| CTGAA | 7221795 | 5.278376 | 186.63635 | 19 |
| AATCT | 6437235 | 5.1963077 | 201.7654 | 39 |
| GTATG | 6135820 | 5.133541 | 209.63025 | 45 |
| ACTGA | 6892080 | 5.0373883 | 184.70325 | 32 |
| GTCTG | 6879505 | 4.985194 | 185.16617 | 17 |
| TCCAG | 7719330 | 4.8866944 | 160.79587 | 25 |
| GAACT | 6649960 | 4.860424 | 185.67366 | 21 |
| AGTCA | 6577800 | 4.8076825 | 184.63316 | 28 |
| AGAGC | 7198205 | 4.7636404 | 171.84045 | 8 |
| TCACT | 6810110 | 4.761349 | 176.75945 | 30 |
| TGAAC | 6429600 | 4.699364 | 185.70009 | 20 |
| TATGC | 6089145 | 4.6571097 | 191.5678 | 46 |
| GAGCA | 6977140 | 4.6173434 | 171.3792 | 9 |
| ATCTC | 6598640 | 4.6134977 | 175.01819 | 40 |
| CACTG | 7188905 | 4.5509105 | 160.21288 | 31 |
| AACTC | 6692490 | 4.4715557 | 169.76399 | 22 |
| CTCCA | 7580765 | 4.386966 | 147.09906 | 24 |
| TCGTA | 5727765 | 4.3807187 | 191.22127 | 43 |
| CGTAT | 5712485 | 4.3690324 | 191.28021 | 44 |
| CACAC | 7841465 | 4.336541 | 143.14676 | 12 |
| CTGAC | 6841125 | 4.33075 | 159.78922 | 33 |
| CCAGT | 6755750 | 4.2767034 | 160.09282 | 26 |
| CAGTC | 6739170 | 4.2662077 | 160.0564 | 27 |
| TCGGA | 6113680 | 4.233724 | 180.2207 | 3 |
| CGGAA | 6396805 | 4.2332883 | 172.26137 | 4 |
| CCAAT | 6267860 | 4.187841 | 167.2766 | 37 |
| ACCAA | 6557290 | 4.1868773 | 160.14998 | 36 |
| TGACC | 6603570 | 4.180366 | 159.2288 | 34 |
| TCTCG | 6302035 | 4.1746626 | 165.70154 | 41 |
| GCACA | 6866735 | 4.1541367 | 156.16566 | 11 |
| GTCAC | 6535430 | 4.137231 | 159.91983 | 29 |
| CAATC | 6189220 | 4.1352983 | 166.82379 | 38 |
| AGCAC | 6825735 | 4.1293335 | 156.42216 | 10 |
| GATCG | 5887250 | 4.0769205 | 179.89943 | 1 |
| ATCGG | 5862930 | 4.0600796 | 180.07613 | 2 |
| CGTCT | 6079365 | 4.0271597 | 168.92744 | 16 |
| ATGCC | 6339595 | 4.0132575 | 158.77509 | 47 |
| TTTTT | 3740245 | 3.994204 | 4.1087255 | 12 |
| CTCGT | 6017860 | 3.9864168 | 165.54941 | 42 |
| GACCA | 6526995 | 3.948606 | 152.03786 | 35 |
| CACGT | 6223300 | 3.939638 | 162.06291 | 14 |
| ACTCC | 6787380 | 3.927837 | 146.8721 | 23 |
| ACGTC | 6123290 | 3.8763273 | 161.49954 | 15 |
| ACACG | 6037995 | 3.652778 | 154.98677 | 13 |
| AAAAA | 3698195 | 3.1477108 | 3.308001 | 34 |
| TTTAA | 1734455 | 1.6915433 | 6.6704097 | 47 |
| TTTAT | 1524815 | 1.5561172 | 6.7626767 | 12 |
| AGGGA | 2005350 | 1.4517412 | 5.113919 | 7 |
| TTCCA | 2065435 | 1.4440672 | 5.117988 | 3 |
| CCTTT | 1922910 | 1.4068246 | 5.1570272 | 45 |
| ATTTA | 1304830 | 1.2725476 | 6.2660494 | 11 |
| GATTT | 1334455 | 1.2330728 | 5.9841347 | 10 |
| CTTTA | 1339700 | 1.1316395 | 5.444669 | 46 |
| TTATA | 904170 | 0.88180023 | 5.873424 | 13 |
| TATAA | 913330 | 0.8512219 | 5.61683 | 14 |
| GGATT | 994115 | 0.8317275 | 5.09584 | 9 |