![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | SV25_CAGATC_L008_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 7110737 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
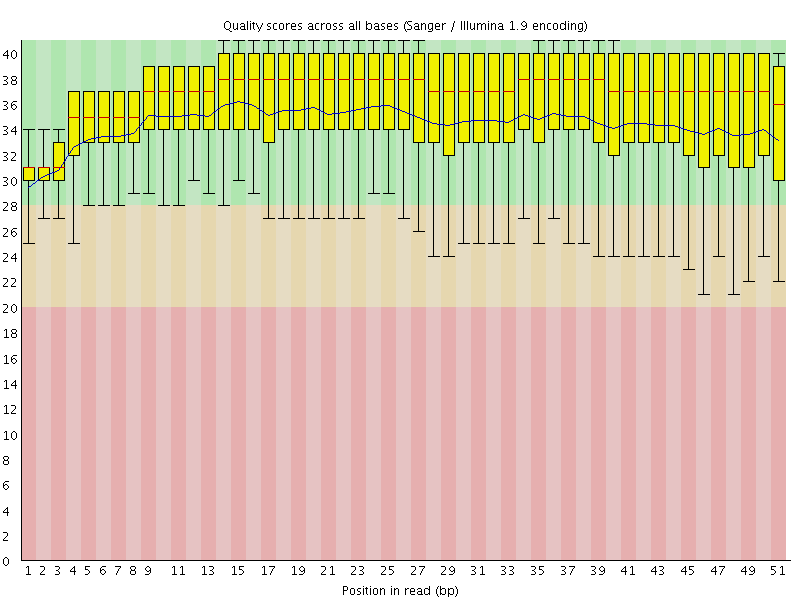
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
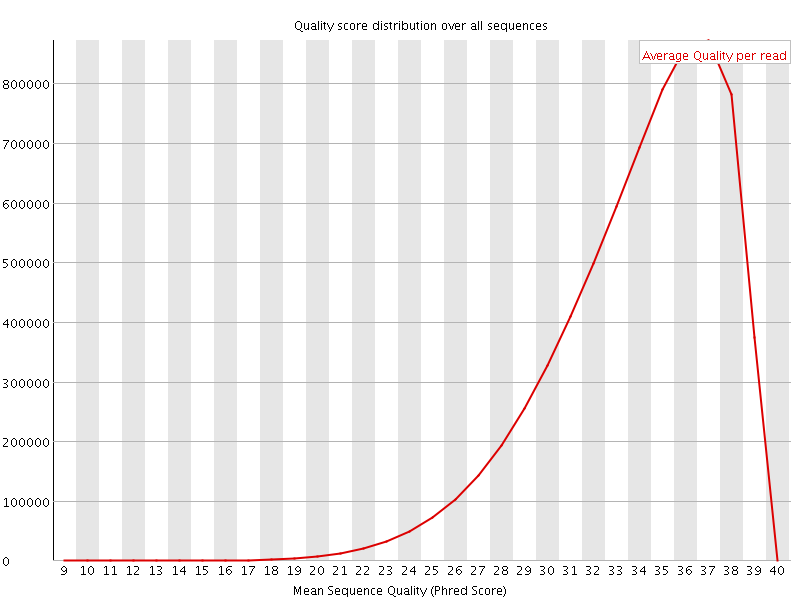
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
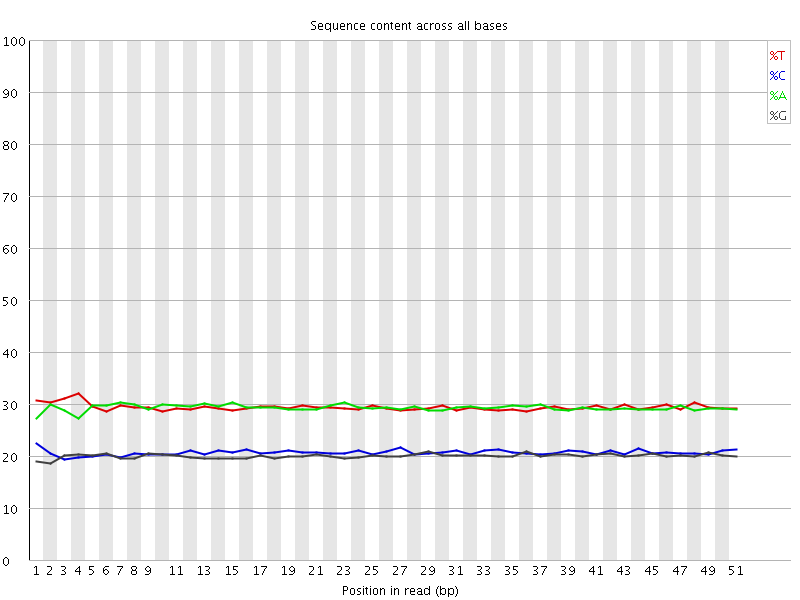
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
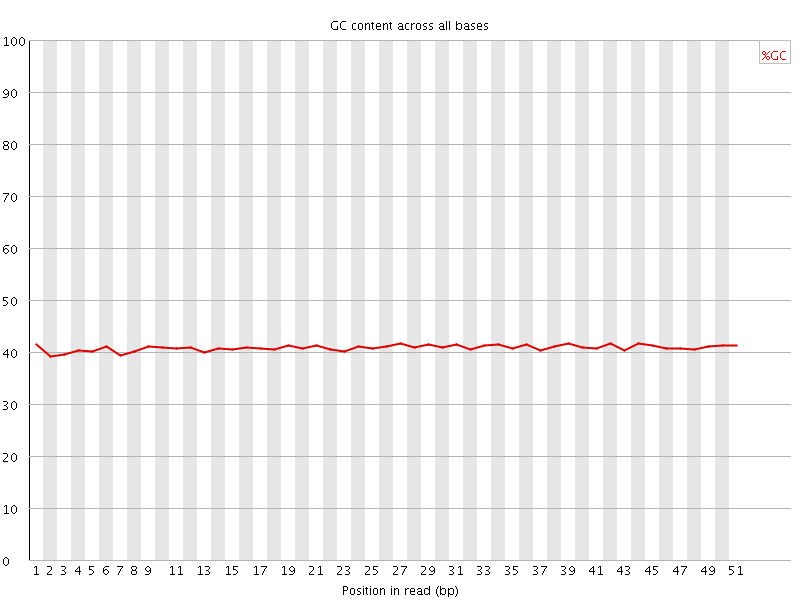
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
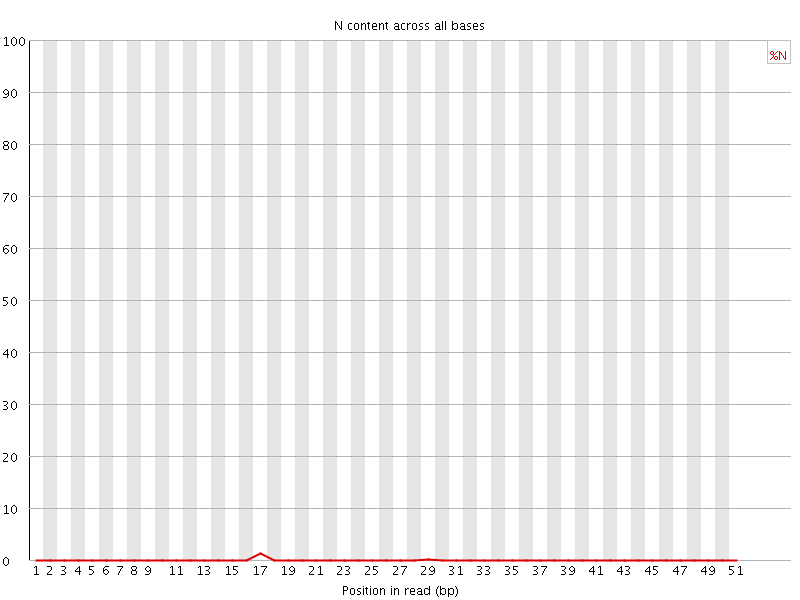
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
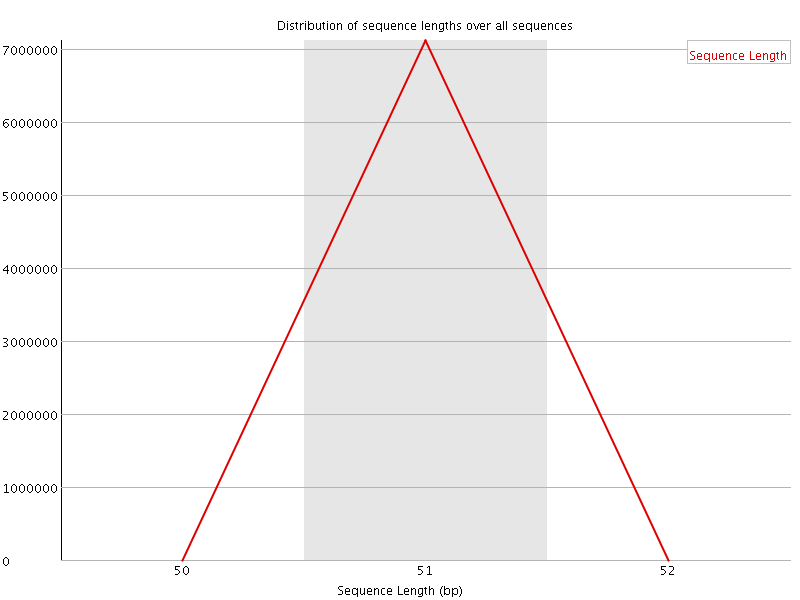
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
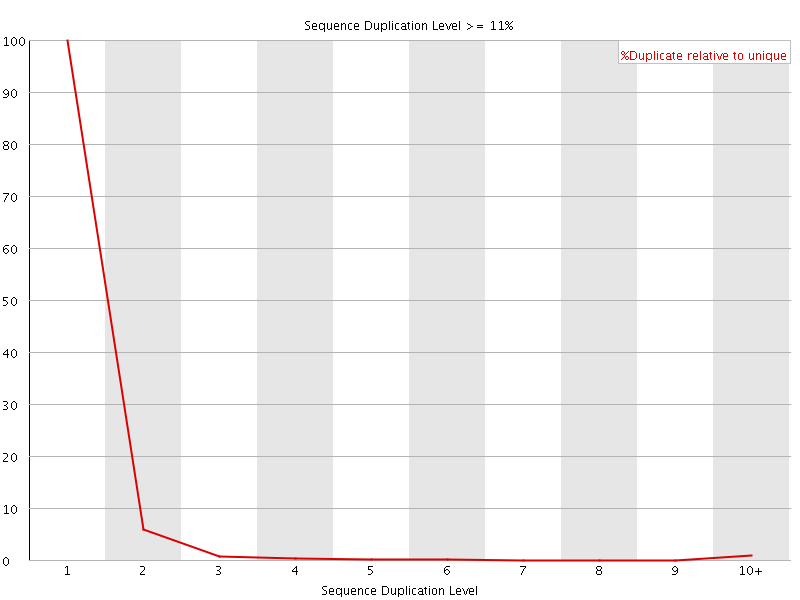
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCAGATCATCTCGTATGCC | 23240 | 0.3268296943059489 | TruSeq Adapter, Index 7 (100% over 51bp) |
| ACTTCCAGGGATTTATAAGCCGATGACGTCATAACATCCCTGACCCTTTAA | 11030 | 0.15511753563660138 | No Hit |
| TATTCGTTGGAATTCCTCGGGGAATTCGGTATTCCCAGGCGGTCTCCCATC | 8359 | 0.11755462197519048 | No Hit |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
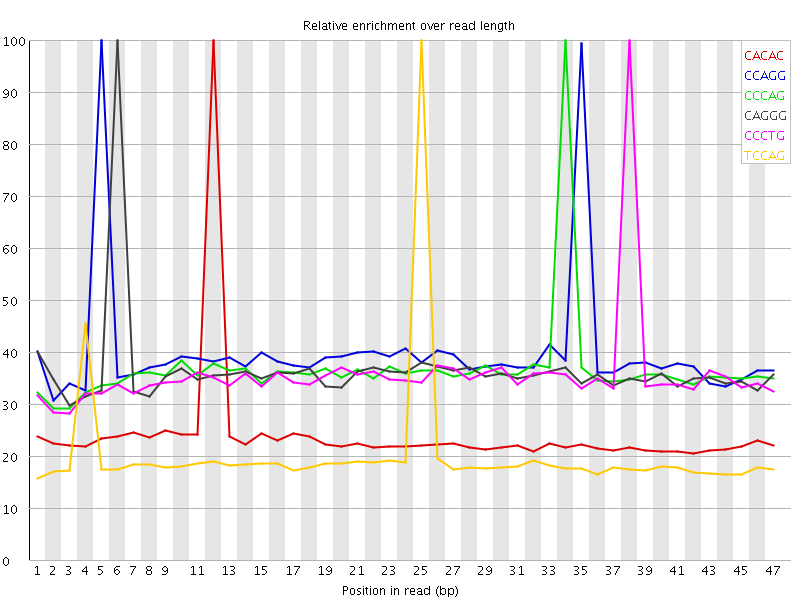
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CACAC | 746670 | 2.8640375 | 11.838825 | 12 |
| CCAGG | 472520 | 2.7297623 | 6.783471 | 5 |
| CCCAG | 452295 | 2.533338 | 6.870746 | 34 |
| CAGGG | 407460 | 2.427856 | 6.6234717 | 6 |
| CCCTG | 417165 | 2.328358 | 6.4997187 | 38 |
| TCCAG | 541880 | 2.1362739 | 10.492001 | 25 |
| CTCCC | 389285 | 2.106572 | 6.3039412 | 44 |
| GGAAG | 491285 | 2.0676801 | 10.8225155 | 5 |
| CAGGC | 352015 | 2.0336013 | 6.238462 | 36 |
| GGGGA | 327920 | 2.0152972 | 6.290072 | 19 |
| CTGAA | 707175 | 1.9692125 | 7.6808558 | 19 |
| CCAGA | 483650 | 1.9134384 | 10.753026 | 33 |
| CTCCA | 500030 | 1.9112461 | 10.450109 | 24 |
| GCACA | 434390 | 1.7185539 | 10.689144 | 11 |
| AGAGC | 407690 | 1.6635914 | 10.564649 | 8 |
| CACCA | 431600 | 1.6555086 | 10.110297 | 31 |
| GAAGA | 566620 | 1.6331286 | 7.8772197 | 6 |
| GAGCA | 388255 | 1.5842861 | 10.571347 | 9 |
| AAGAG | 547890 | 1.5791446 | 7.826525 | 7 |
| TCTGA | 545965 | 1.5149597 | 7.1180396 | 18 |
| GTCTG | 366660 | 1.4856658 | 9.463361 | 17 |
| ACCAG | 374755 | 1.4826229 | 10.259815 | 32 |
| AGCAC | 374020 | 1.4797151 | 10.420082 | 10 |
| CCAGT | 370195 | 1.4594337 | 9.666598 | 26 |
| CATCT | 536835 | 1.4442545 | 7.285849 | 39 |
| CAGTC | 340195 | 1.3411634 | 9.5952 | 27 |
| CAGAT | 470175 | 1.309258 | 7.5189104 | 34 |
| ATGCC | 318140 | 1.2542152 | 9.282432 | 47 |
| TCACC | 322620 | 1.2331384 | 9.697449 | 30 |
| ACTCC | 322315 | 1.2319725 | 9.987052 | 23 |
| AACTC | 435225 | 1.175023 | 7.2943945 | 22 |
| AGTCA | 419430 | 1.1679525 | 6.9326677 | 28 |
| GTCAC | 288120 | 1.1358662 | 9.584073 | 29 |
| ATCTC | 413795 | 1.1132385 | 7.0414457 | 40 |
| TCATC | 406960 | 1.0948502 | 6.97533 | 38 |
| GACCC | 194560 | 1.089745 | 5.3296366 | 42 |
| GAACT | 381460 | 1.0622206 | 6.7973266 | 21 |
| TGAAC | 356540 | 0.99282795 | 6.7299647 | 20 |
| CACGT | 246410 | 0.9714314 | 9.6038 | 14 |
| ATCAT | 497290 | 0.94498444 | 5.0879946 | 37 |
| AGATC | 329515 | 0.9175736 | 7.0488634 | 35 |
| GTATG | 290200 | 0.8305521 | 6.2928047 | 45 |
| CGGGG | 94800 | 0.8248362 | 6.9011135 | 18 |
| GATCA | 295925 | 0.8240382 | 6.8803177 | 36 |
| TATGC | 289460 | 0.8032021 | 6.3008246 | 46 |
| ACGTC | 187745 | 0.7401541 | 9.258811 | 15 |
| GGCGG | 82875 | 0.7210791 | 7.050219 | 38 |
| CGGAA | 148395 | 0.6055303 | 8.899575 | 4 |
| TCTCG | 151530 | 0.59528226 | 8.973797 | 41 |
| ACACG | 129745 | 0.51330316 | 9.236896 | 13 |
| CGTCT | 114005 | 0.4478661 | 8.960011 | 16 |
| CTCGT | 110545 | 0.4342736 | 8.56555 | 42 |
| TCGGA | 95485 | 0.38825965 | 8.806305 | 3 |
| ATCGG | 83720 | 0.34042096 | 8.812026 | 2 |
| GATCG | 82740 | 0.3364361 | 9.021382 | 1 |
| CGTAT | 94325 | 0.26173577 | 5.8062787 | 44 |
| TCGTA | 85705 | 0.23781674 | 5.7963967 | 43 |