![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | SV20_CGATGT_L007_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 12933943 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
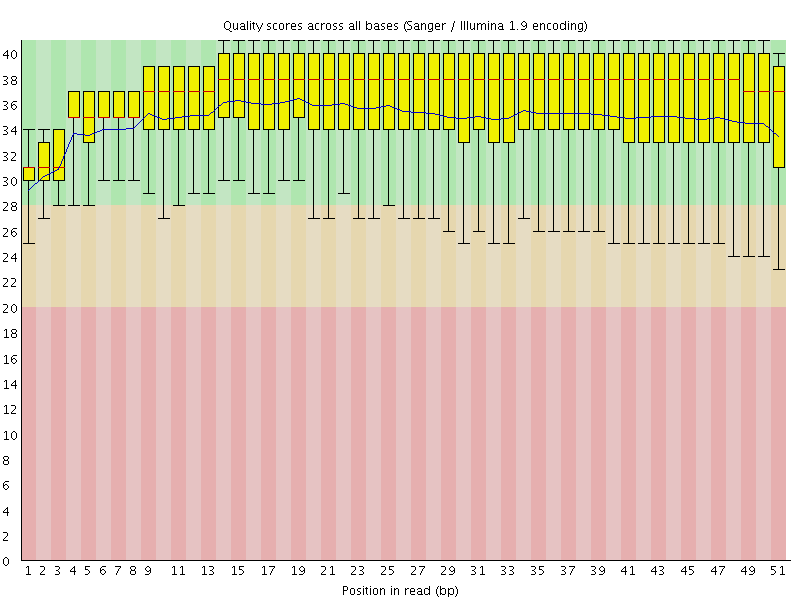
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
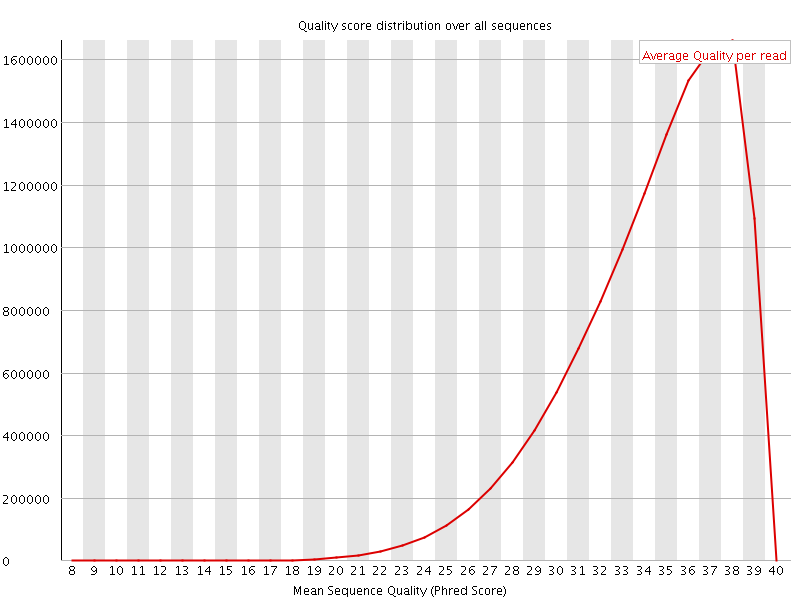
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
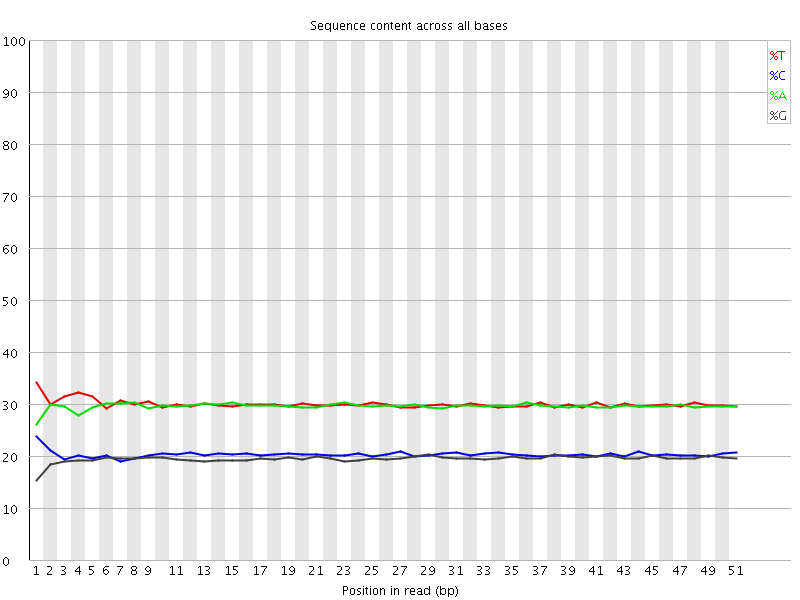
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
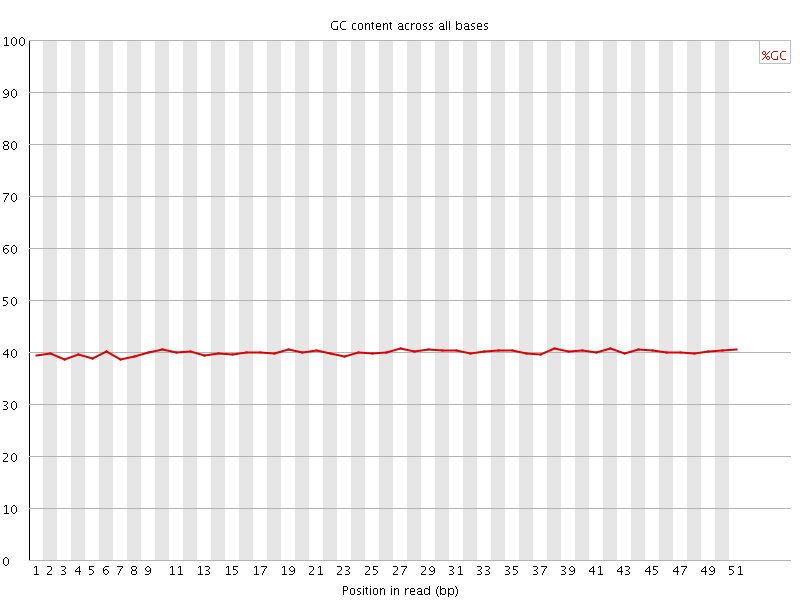
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
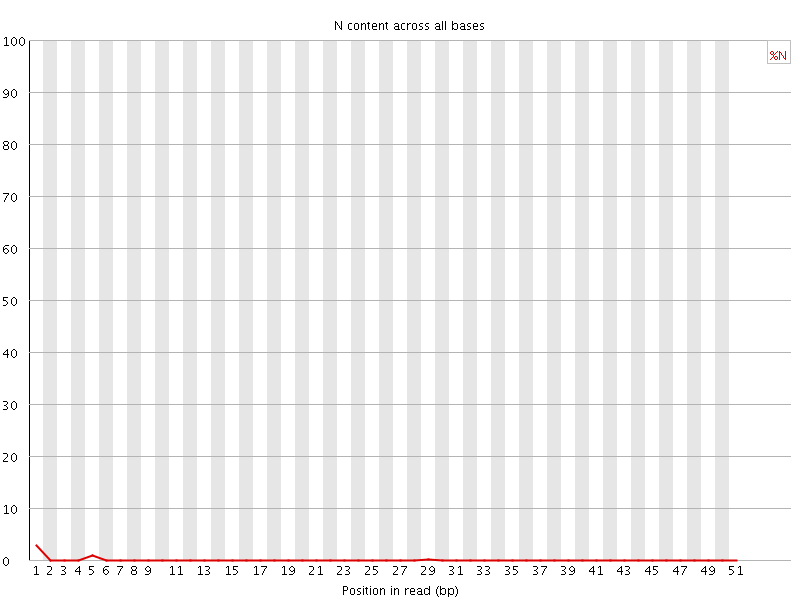
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
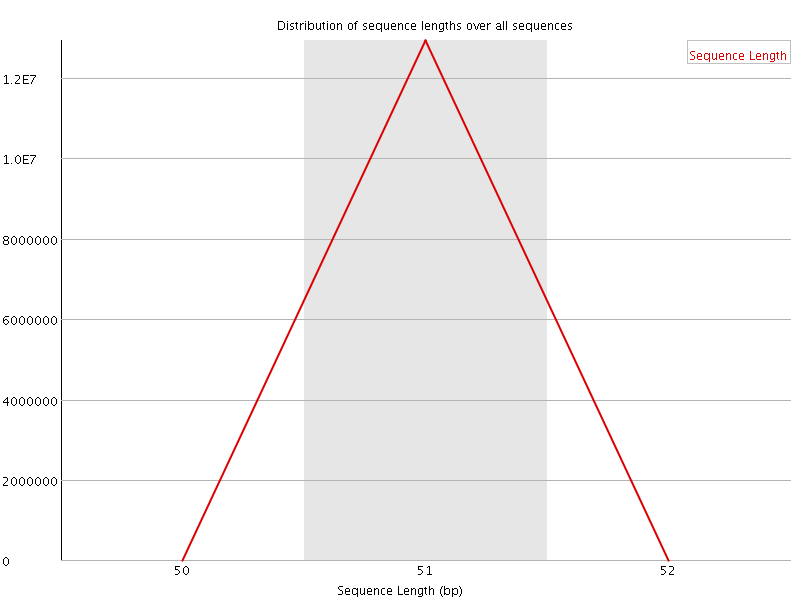
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
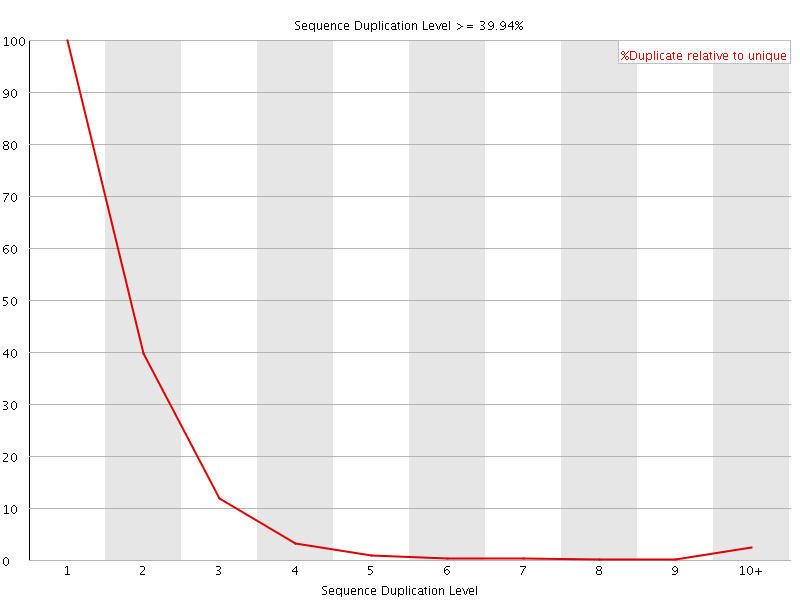
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCGATGTATCTCGTATGCC | 36733 | 0.28400465349197845 | TruSeq Adapter, Index 2 (100% over 51bp) |
| ACTTCCAGGGATTTATAAGCCGATGACGTCATAACATCCCTGACCCTTTAA | 17519 | 0.13544980057512238 | No Hit |
| TATTCGTTGGAATTCCTCGGGGAATTCGGTATTCCCAGGCGGTCTCCCATC | 15104 | 0.1167780003360151 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
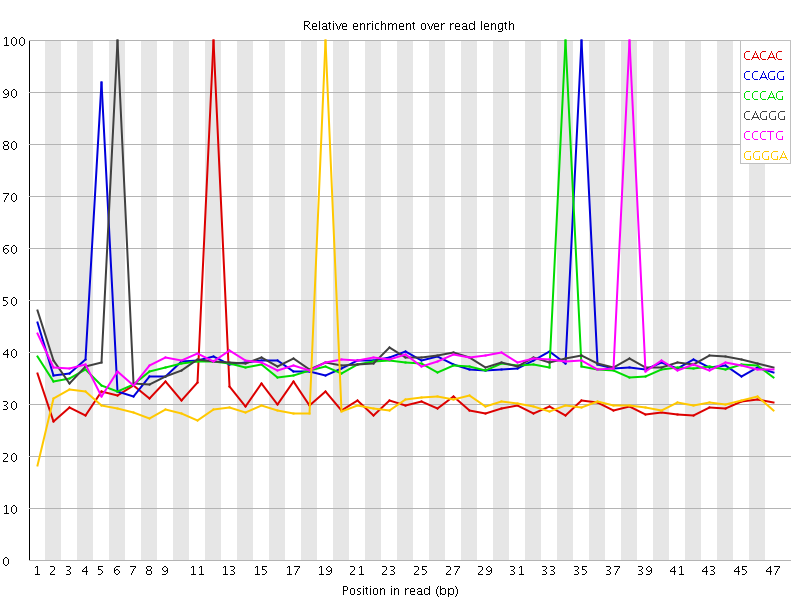
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CACAC | 1330635 | 2.8951218 | 9.067362 | 12 |
| CCAGG | 834430 | 2.865789 | 7.161762 | 35 |
| CCCAG | 795905 | 2.6230245 | 6.865413 | 34 |
| CAGGG | 702595 | 2.5146208 | 6.363669 | 6 |
| CCCTG | 725560 | 2.3579075 | 5.994901 | 38 |
| GGGGA | 569110 | 2.1226425 | 6.822893 | 19 |
| CTCCC | 678730 | 2.1165926 | 6.2107844 | 44 |
| CAGGC | 616240 | 2.1164315 | 6.5833545 | 36 |
| TCCAG | 943800 | 2.11015 | 8.255485 | 25 |
| GGAAG | 825845 | 2.0335083 | 8.811857 | 5 |
| CTGAA | 1233075 | 1.8967183 | 6.3255773 | 19 |
| CTCCA | 861420 | 1.8481411 | 7.8981724 | 24 |
| GCACA | 728925 | 1.6527379 | 8.013619 | 11 |
| AGAGC | 683240 | 1.6143868 | 8.050581 | 8 |
| GAAGA | 986755 | 1.6040709 | 6.0969043 | 6 |
| AAGAG | 950495 | 1.5451266 | 5.8921103 | 7 |
| GAGCA | 647340 | 1.5295608 | 7.9336004 | 9 |
| TCTGA | 970695 | 1.4723412 | 5.7015157 | 18 |
| GTCTG | 632560 | 1.4533179 | 7.7689056 | 17 |
| CCAGT | 635850 | 1.4216348 | 7.531525 | 26 |
| AGCAC | 622385 | 1.4111729 | 7.738184 | 10 |
| CAGTC | 575430 | 1.2865477 | 7.37686 | 27 |
| ATGCC | 546505 | 1.221877 | 7.238317 | 47 |
| TCACC | 546725 | 1.1729759 | 7.259432 | 30 |
| ACTCC | 542355 | 1.1636004 | 7.391007 | 23 |
| AGTCA | 719895 | 1.1073438 | 5.2998824 | 28 |
| AACTC | 746915 | 1.1024816 | 5.372021 | 22 |
| GTCAC | 487090 | 1.089037 | 7.266795 | 29 |
| GATGT | 671315 | 1.0611218 | 5.4026 | 35 |
| ATCTC | 725330 | 1.0557184 | 5.1901603 | 40 |
| GAACT | 664670 | 1.0223968 | 5.30242 | 21 |
| TGAAC | 623505 | 0.9590767 | 5.246485 | 20 |
| CACGT | 417365 | 0.9331456 | 7.242962 | 14 |
| CGGGG | 162330 | 0.88003415 | 7.6583414 | 18 |
| GTATG | 513845 | 0.8122149 | 5.016237 | 45 |
| GGCGG | 142445 | 0.7722321 | 7.646377 | 38 |
| ACGTC | 314245 | 0.7025897 | 6.9693046 | 15 |
| CACCG | 170450 | 0.5617436 | 9.659985 | 31 |
| CGGAA | 237125 | 0.5602885 | 7.226479 | 4 |
| TCTCG | 253925 | 0.5598235 | 6.764032 | 41 |
| AGGCG | 139150 | 0.49802446 | 5.0129223 | 37 |
| ACACG | 207620 | 0.47074994 | 6.897584 | 13 |
| CGATG | 182275 | 0.42469165 | 6.7204123 | 34 |
| CGTCT | 180245 | 0.39738268 | 6.5905848 | 16 |
| CTCGT | 174475 | 0.38466164 | 6.5049624 | 42 |
| CCGAT | 154990 | 0.34652698 | 6.390657 | 33 |
| TCGGA | 147995 | 0.34482092 | 6.934999 | 3 |
| ACCGA | 148855 | 0.33750835 | 6.498005 | 32 |
| ATCGG | 129530 | 0.30179837 | 6.9063096 | 2 |
| GATCG | 124560 | 0.29021856 | 6.9161882 | 1 |