![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | SV19_TGACCA_L007_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 15948282 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 43 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
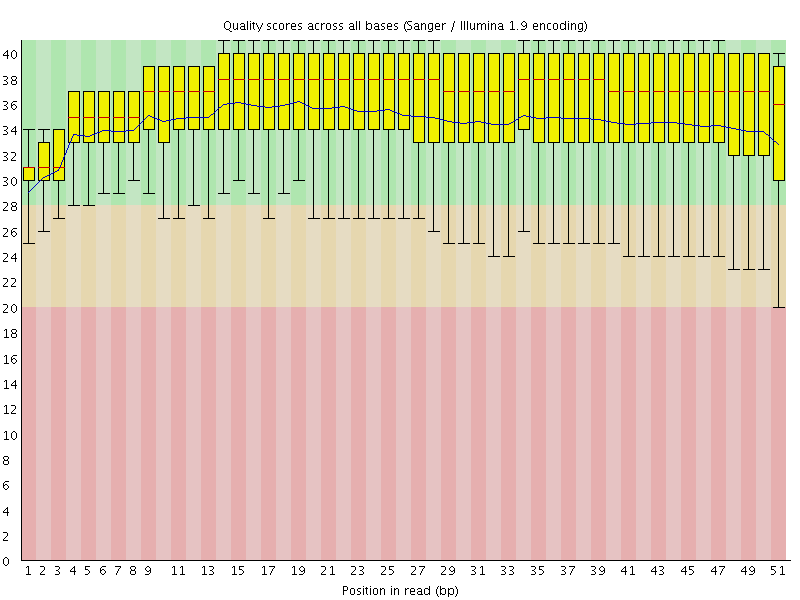
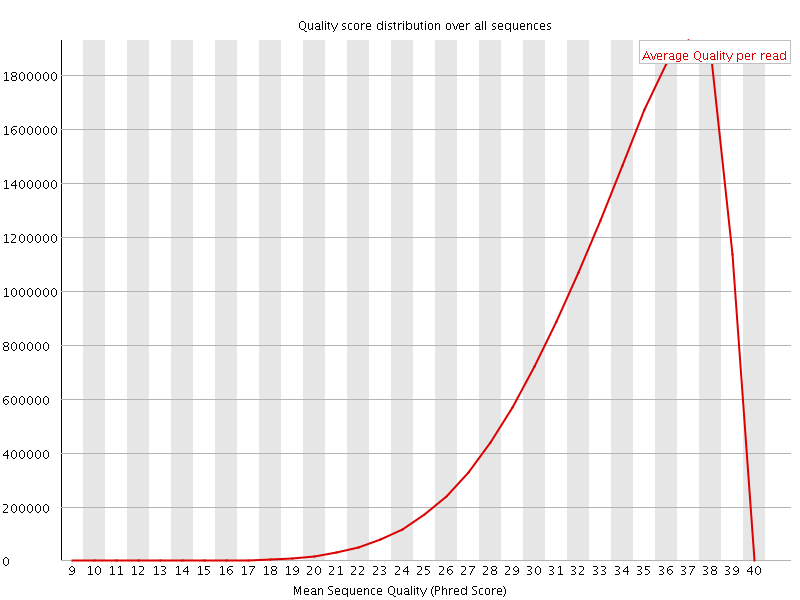
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
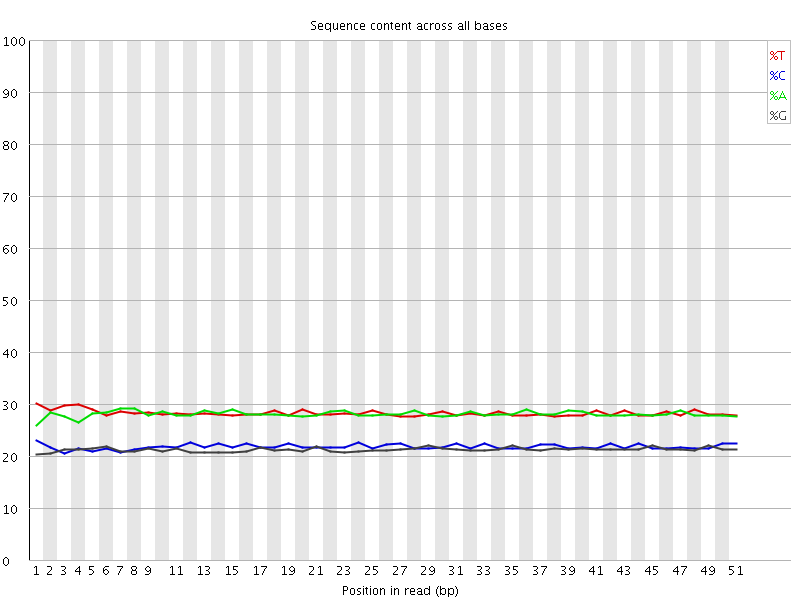
![[OK]](Icons/tick.png) Per base sequence content
Per base sequence content
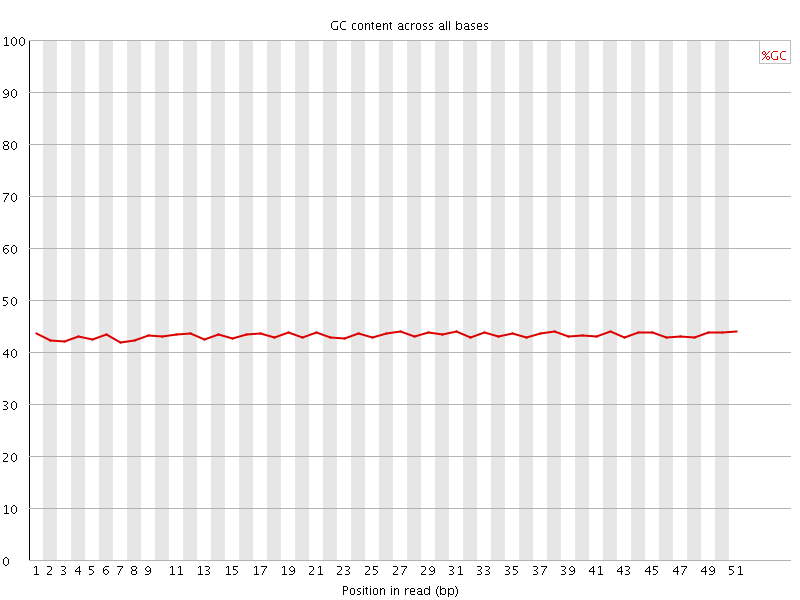
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
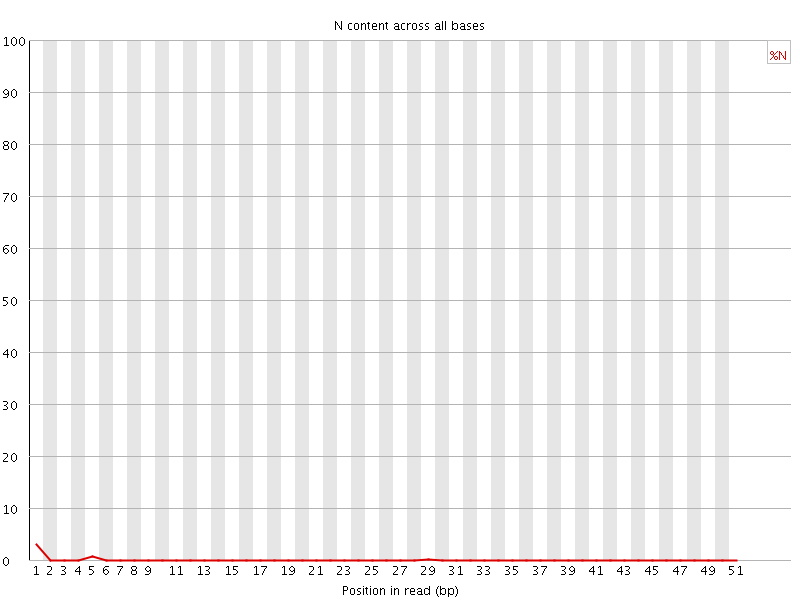
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
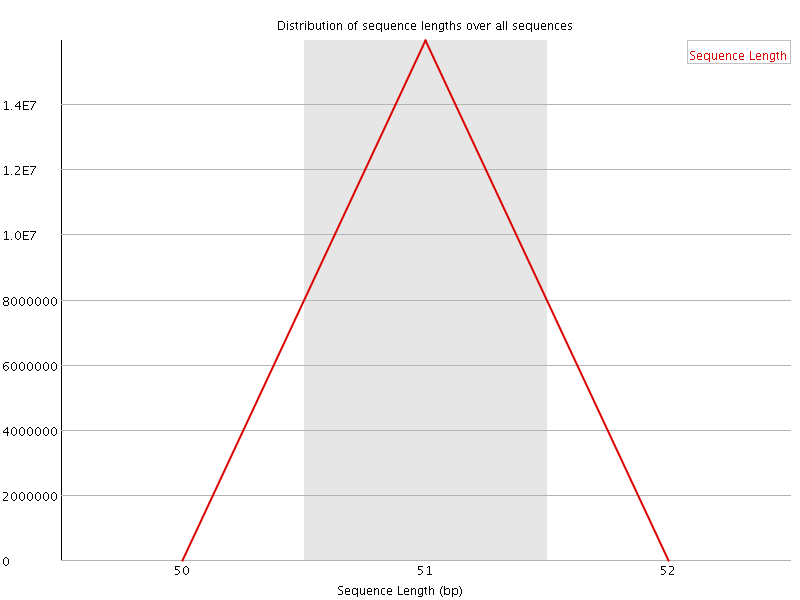
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
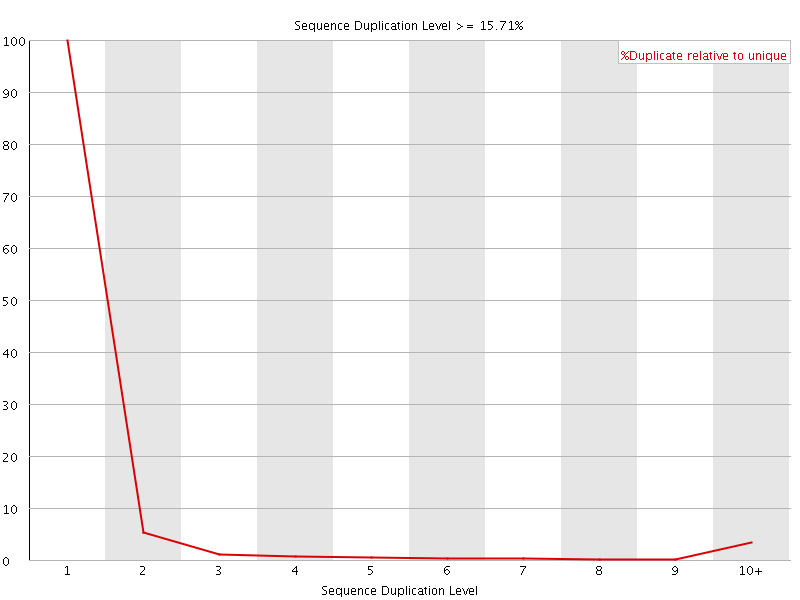
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACTGACCAATCTCGTATGCC | 72489 | 0.45452544669074707 | TruSeq Adapter, Index 4 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
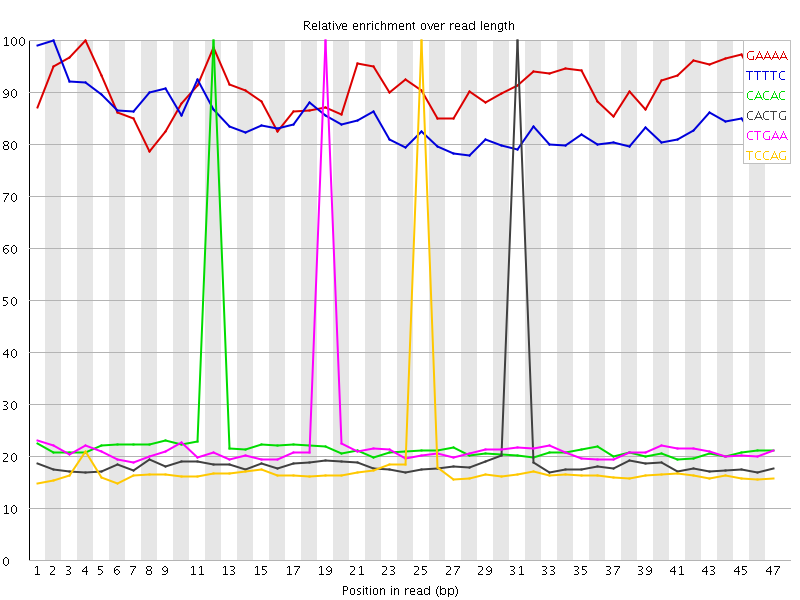
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GAAAA | 3183690 | 3.1320646 | 3.4597666 | 4 |
| TTTTC | 3298400 | 3.0791974 | 3.64541 | 2 |
| CACAC | 1919320 | 3.0337892 | 13.237965 | 12 |
| CACTG | 1509410 | 2.4376295 | 12.239217 | 31 |
| CTGAA | 1800630 | 2.2626133 | 10.038088 | 19 |
| TCCAG | 1372835 | 2.2170668 | 12.076814 | 25 |
| GGAAG | 1215675 | 2.0877993 | 12.607973 | 5 |
| ACTGA | 1604170 | 2.015748 | 9.740608 | 32 |
| CTCCA | 1228310 | 1.9295559 | 11.713528 | 24 |
| GCACA | 1073195 | 1.7439246 | 11.894466 | 11 |
| GAAGA | 1333765 | 1.733664 | 9.79687 | 6 |
| AGAGC | 965675 | 1.613211 | 11.709561 | 8 |
| AAGAG | 1222595 | 1.5891622 | 9.44194 | 7 |
| GAGCA | 933070 | 1.5587426 | 11.568068 | 9 |
| TCTGA | 1205195 | 1.5050623 | 9.212096 | 18 |
| GTCTG | 895700 | 1.4779013 | 11.456541 | 17 |
| CCAGT | 901895 | 1.45652 | 11.059375 | 26 |
| TCACT | 1194240 | 1.4507004 | 8.914417 | 30 |
| AGCAC | 870105 | 1.4139067 | 11.477219 | 10 |
| CTGAC | 839980 | 1.35653 | 11.173381 | 33 |
| CAGTC | 819250 | 1.3230519 | 10.912309 | 27 |
| CACGT | 773555 | 1.2492566 | 11.317137 | 14 |
| GACCA | 750320 | 1.219258 | 10.888327 | 35 |
| ACTCC | 775545 | 1.218306 | 11.1918545 | 23 |
| ATGCC | 751575 | 1.2137599 | 10.100684 | 47 |
| AACTC | 975955 | 1.1929014 | 8.918227 | 22 |
| TGACC | 732200 | 1.1824701 | 10.8707485 | 34 |
| AGTCA | 922555 | 1.1592528 | 8.547984 | 28 |
| ACCAA | 919990 | 1.1314791 | 8.472425 | 36 |
| GAACT | 896755 | 1.1268333 | 8.854343 | 21 |
| ATCTC | 923150 | 1.1213944 | 8.615064 | 40 |
| GTCAC | 684870 | 1.1060343 | 10.822155 | 29 |
| TGAAC | 817085 | 1.0267226 | 8.734669 | 20 |
| ACGTC | 559050 | 0.9028406 | 10.938102 | 15 |
| CCAAT | 730260 | 0.89259046 | 8.1795 | 37 |
| AATCT | 920115 | 0.86966825 | 6.591361 | 39 |
| GTATG | 667325 | 0.8567333 | 7.691782 | 45 |
| TATGC | 635230 | 0.79328305 | 7.4475327 | 46 |
| CAATC | 626525 | 0.76579607 | 8.108763 | 38 |
| CGGAA | 441960 | 0.7383175 | 11.200039 | 4 |
| TCTCG | 454415 | 0.7293306 | 10.543736 | 41 |
| ACACG | 349220 | 0.5674769 | 10.71649 | 13 |
| CGTCT | 309285 | 0.49639872 | 10.448015 | 16 |
| CTCGT | 291310 | 0.46754906 | 9.895157 | 42 |
| TCGGA | 270090 | 0.44841486 | 10.906171 | 3 |
| ATCGG | 233040 | 0.3869029 | 10.914029 | 2 |
| GATCG | 228910 | 0.38004613 | 11.02283 | 1 |
| CGTAT | 237630 | 0.29675525 | 7.2514977 | 44 |
| TCGTA | 212925 | 0.26590335 | 7.2821865 | 43 |