![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | SV17_CAGATC_L007_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 17208101 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 49 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
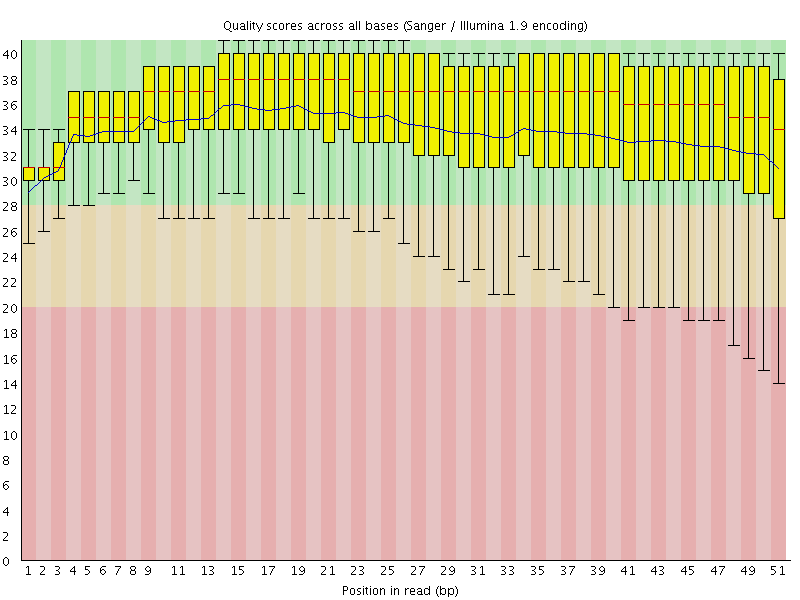
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
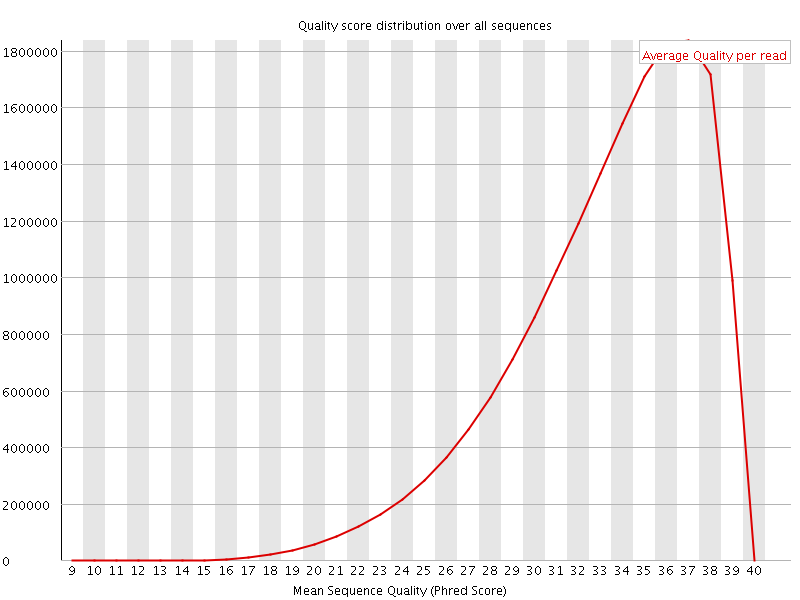
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
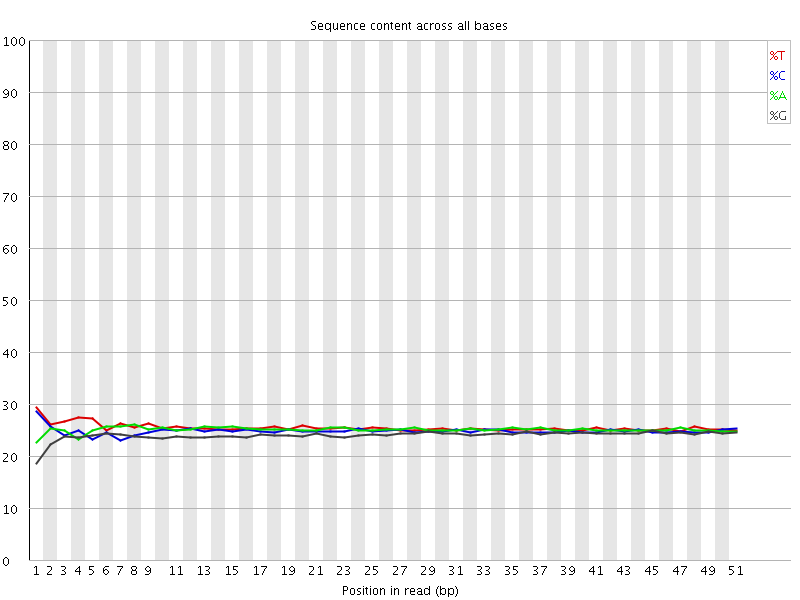
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
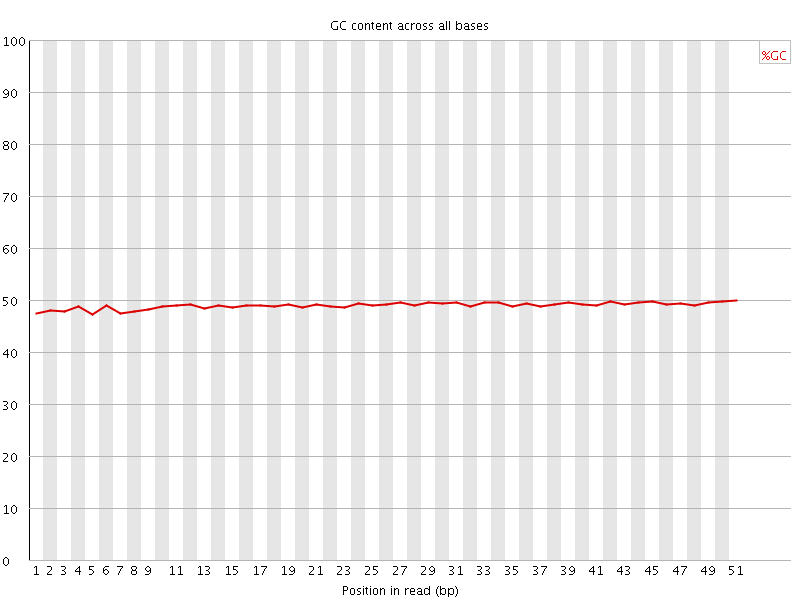
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
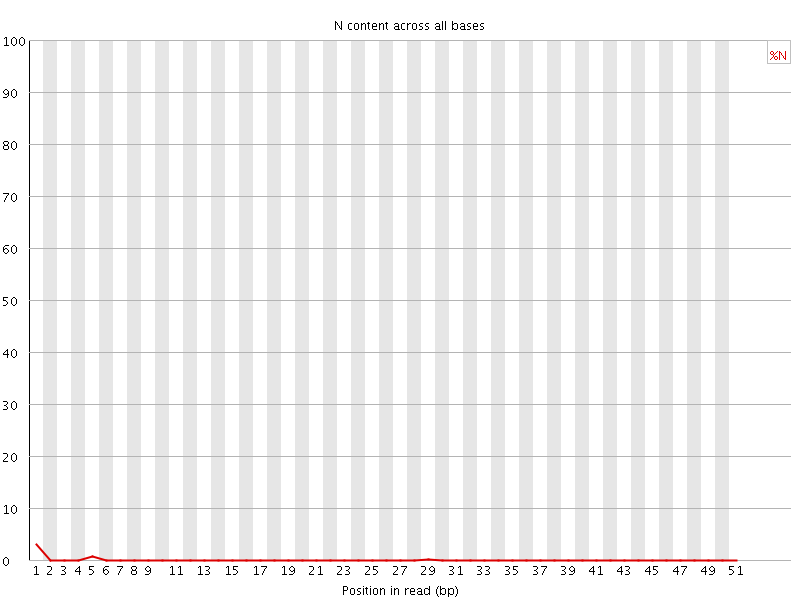
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
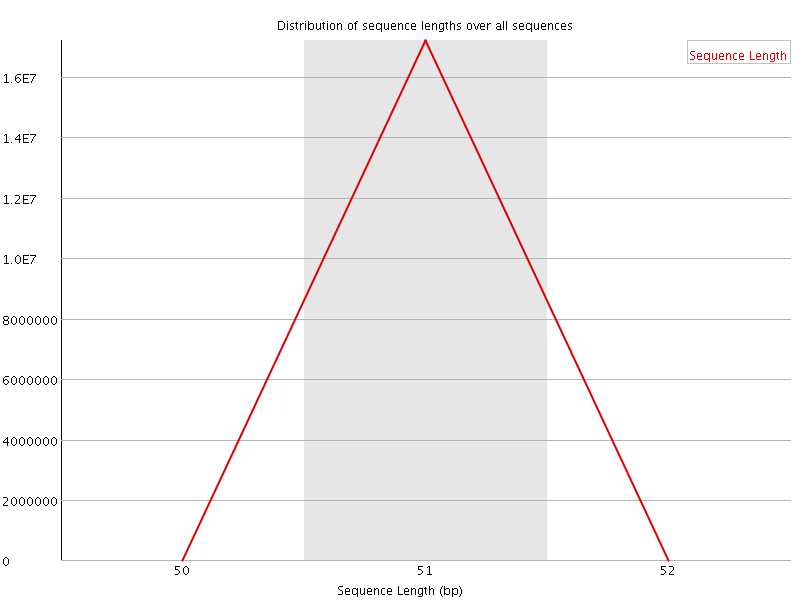
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
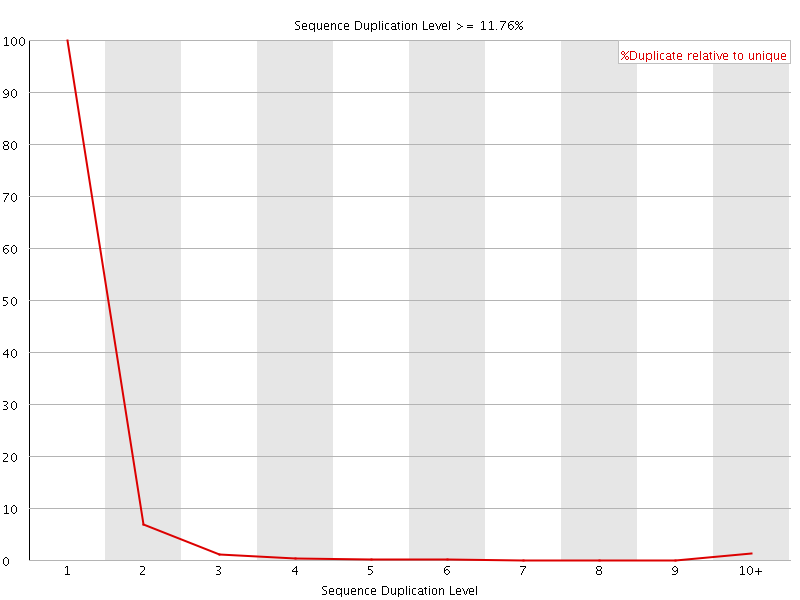
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCAGATCATCTCGTATGCC | 49234 | 0.2861094318309731 | TruSeq Adapter, Index 7 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
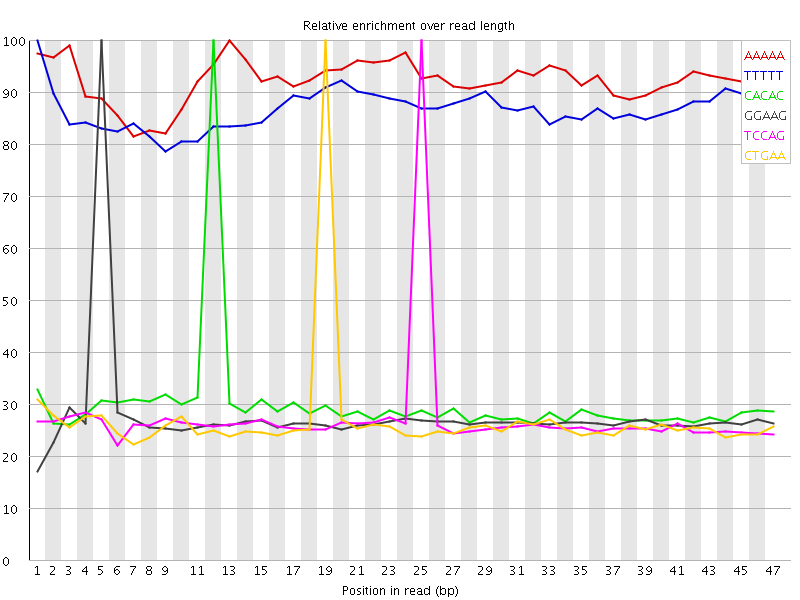
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 2652955 | 3.2004888 | 3.467678 | 13 |
| TTTTT | 2727700 | 3.0536678 | 3.5147133 | 1 |
| CACAC | 1649675 | 2.0600798 | 6.8407583 | 12 |
| GGAAG | 1415000 | 1.9532465 | 7.0259066 | 5 |
| TCCAG | 1397080 | 1.7771317 | 6.479143 | 25 |
| CTGAA | 1376260 | 1.7306098 | 6.3960986 | 19 |
| CTCCA | 1367085 | 1.681856 | 6.256872 | 24 |
| GAAGA | 1258295 | 1.660651 | 6.4596305 | 6 |
| CCAGA | 1267420 | 1.6364816 | 6.3407354 | 33 |
| AAGAG | 1199960 | 1.5836626 | 6.2720156 | 7 |
| AGAGC | 1119025 | 1.4939477 | 6.3271136 | 8 |
| TCTGA | 1143610 | 1.4167209 | 5.98451 | 18 |
| GAGCA | 1018220 | 1.3593684 | 6.1357336 | 9 |
| CACCA | 1078340 | 1.3466085 | 5.9562483 | 31 |
| CATCT | 1103490 | 1.3221169 | 5.7619834 | 39 |
| GCACA | 1014330 | 1.3096942 | 6.1659822 | 11 |
| GTCTG | 957660 | 1.2408592 | 5.9498224 | 17 |
| ACCAG | 934945 | 1.2071929 | 5.906946 | 32 |
| AGCAC | 930725 | 1.2017441 | 5.997441 | 10 |
| CAGAT | 935750 | 1.1766804 | 5.7905903 | 34 |
| CCAGT | 907685 | 1.1546052 | 5.7413516 | 26 |
| AACTC | 923630 | 1.1232895 | 5.722442 | 22 |
| TCACC | 907435 | 1.1163718 | 5.5994563 | 30 |
| CAGTC | 863335 | 1.0981907 | 5.6904693 | 27 |
| GAACT | 861980 | 1.0839165 | 5.713288 | 21 |
| AGTCA | 856620 | 1.0771766 | 5.5594344 | 28 |
| ACTCC | 865805 | 1.0651566 | 5.7329497 | 23 |
| ATCTC | 864880 | 1.0362326 | 5.557646 | 40 |
| ATCAT | 821935 | 0.97350717 | 5.3109345 | 37 |
| TCATC | 808550 | 0.9687422 | 5.38505 | 38 |
| TGAAC | 754990 | 0.94937956 | 5.5720367 | 20 |
| GTCAC | 727655 | 0.9256012 | 5.5304136 | 29 |
| AGATC | 729510 | 0.91733915 | 5.476209 | 35 |
| ATGCC | 706895 | 0.89919376 | 5.4055066 | 47 |
| GATCA | 590230 | 0.7421983 | 5.300993 | 36 |
| GTATG | 573660 | 0.7347946 | 5.2023735 | 45 |
| CACGT | 570895 | 0.7261972 | 5.466672 | 14 |
| CGGAA | 527060 | 0.70364827 | 5.6929483 | 4 |
| TCTCG | 507545 | 0.6360344 | 5.3186035 | 41 |
| ACGTC | 419790 | 0.5339868 | 5.2936854 | 15 |
| ACACG | 401085 | 0.5178775 | 5.3570733 | 13 |
| TCGGA | 381185 | 0.50134784 | 5.4080095 | 3 |
| CGTCT | 399185 | 0.50024223 | 5.179295 | 16 |
| CTCGT | 356260 | 0.44645032 | 5.071432 | 42 |
| GATCG | 274000 | 0.3603744 | 5.2625365 | 1 |
| ATCGG | 262485 | 0.3452295 | 5.227152 | 2 |