![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | DGY955_1_GCCAAT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 24478116 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 38 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
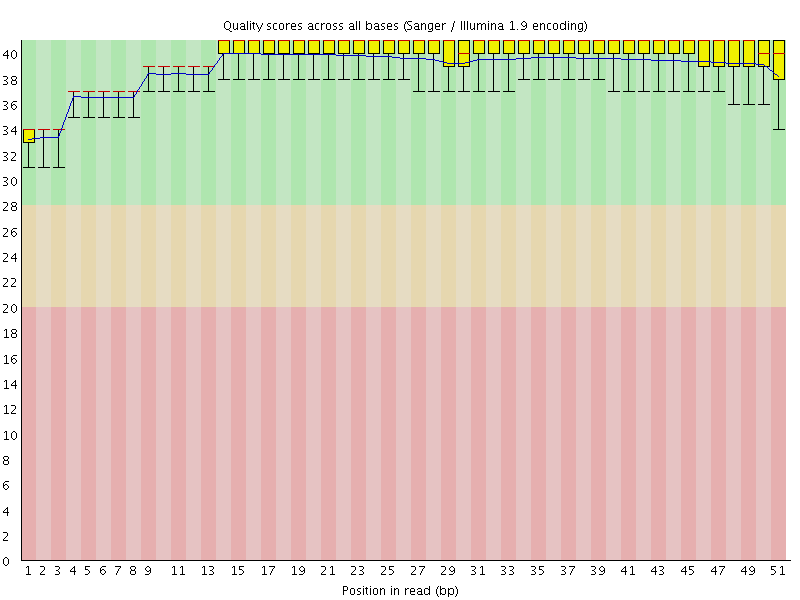
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
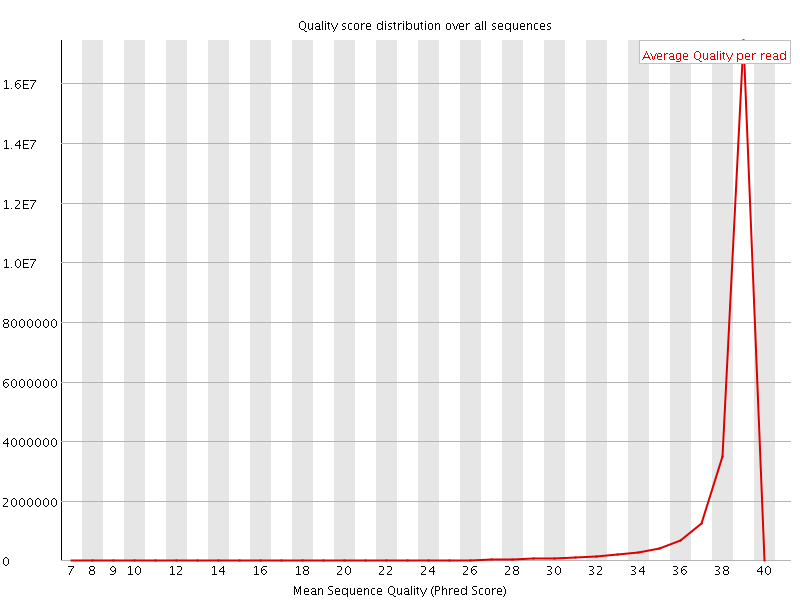
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
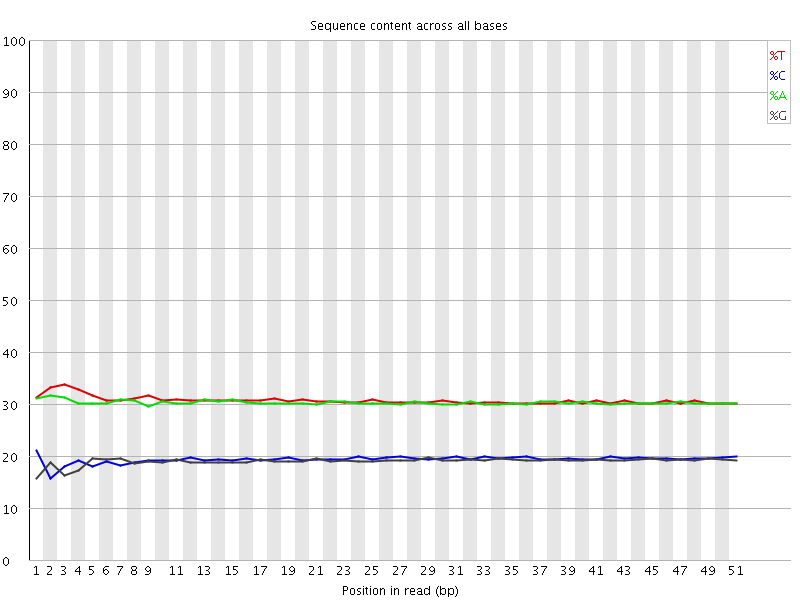
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
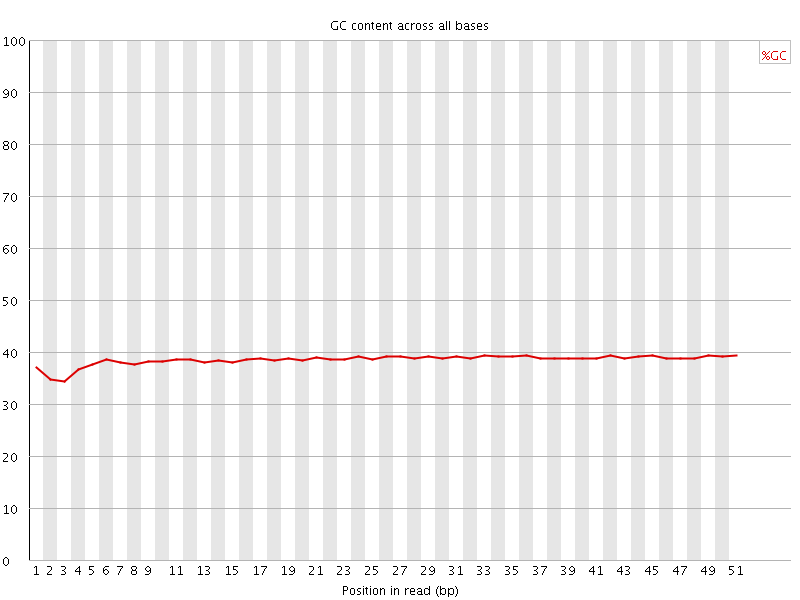
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
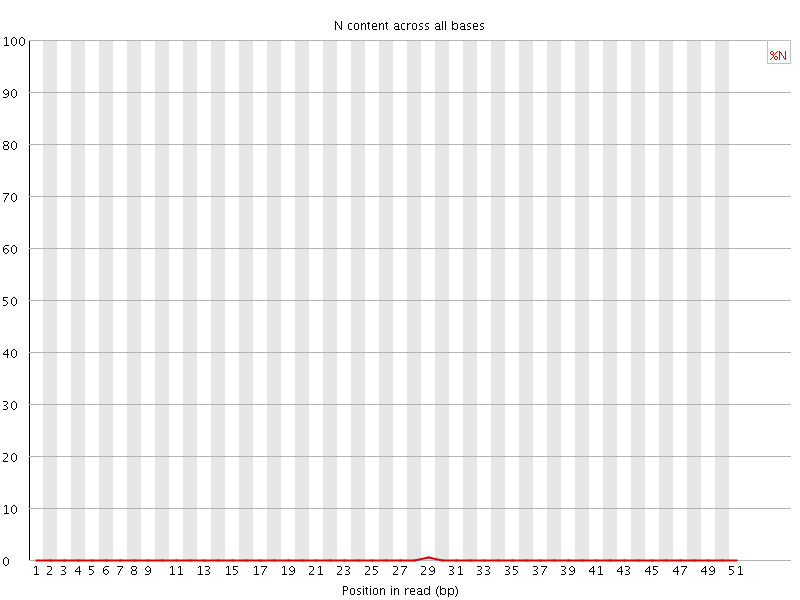
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
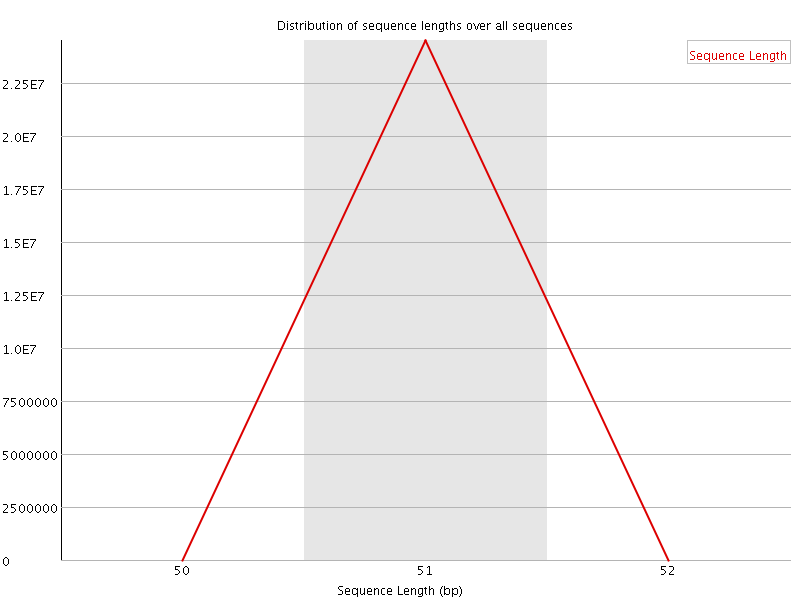
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
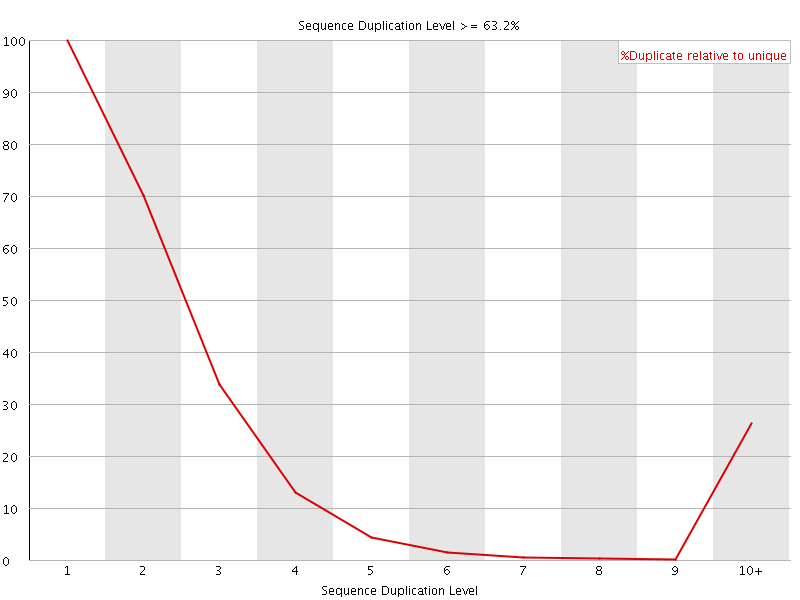
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGCCAATATCTCGTATGCC | 108341 | 0.44260350755752603 | TruSeq Adapter, Index 6 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
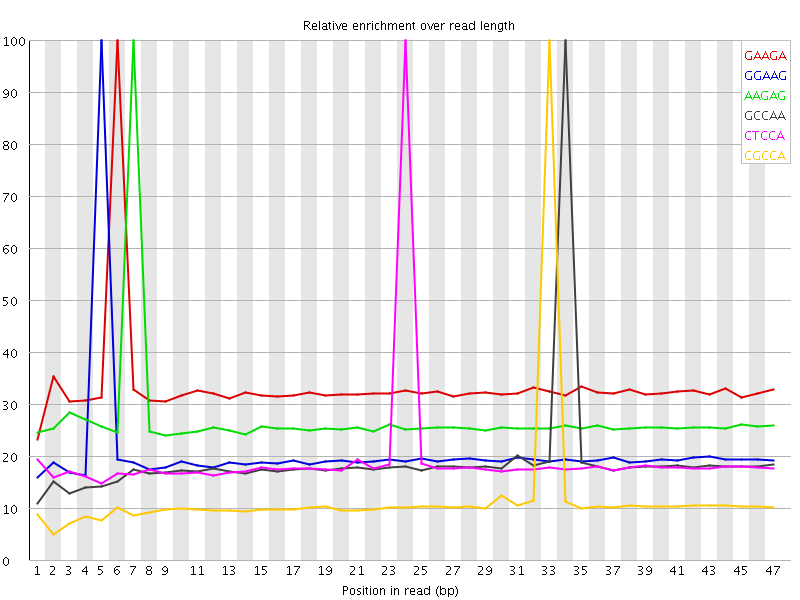
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GAAGA | 2671980 | 2.2401228 | 6.691985 | 6 |
| GGAAG | 1404935 | 1.8715601 | 9.035239 | 5 |
| AAGAG | 1967710 | 1.6496805 | 6.08366 | 7 |
| GCCAA | 1241215 | 1.5956583 | 8.329102 | 34 |
| CTCCA | 1249265 | 1.5567942 | 8.041578 | 24 |
| CGCCA | 708195 | 1.4211087 | 11.938006 | 33 |
| TCCAG | 1106790 | 1.4040067 | 8.088461 | 25 |
| AGAGC | 1019565 | 1.3342437 | 8.265487 | 8 |
| CTGAA | 1612105 | 1.3101339 | 5.5650573 | 19 |
| CCAAT | 1623900 | 1.2964457 | 5.4646635 | 35 |
| CCAGT | 1003320 | 1.272751 | 7.901128 | 26 |
| GAGCA | 957975 | 1.2536445 | 8.248592 | 9 |
| TGAAC | 1520470 | 1.2356634 | 5.5033445 | 20 |
| CGGAA | 942640 | 1.2335765 | 8.16842 | 4 |
| ATGCC | 964035 | 1.2229164 | 7.858192 | 47 |
| AGCAC | 915855 | 1.1773881 | 7.9971986 | 10 |
| CACAC | 907630 | 1.146237 | 7.8626866 | 12 |
| ACTCC | 896515 | 1.1172084 | 7.6244917 | 23 |
| TCTGA | 1382815 | 1.1089131 | 5.300598 | 18 |
| GAACT | 1326230 | 1.0778074 | 5.3414025 | 21 |
| ACGCC | 507790 | 1.0189635 | 11.500196 | 32 |
| ATCTC | 1267390 | 0.99842715 | 5.0736732 | 40 |
| GCACA | 774700 | 0.9959246 | 7.8333316 | 11 |
| AACTC | 1213360 | 0.96868986 | 5.1269855 | 22 |
| GTCTG | 758530 | 0.9665293 | 7.660772 | 17 |
| CACGC | 477065 | 0.9573087 | 11.455292 | 31 |
| GTCAC | 748330 | 0.9492861 | 7.6147475 | 29 |
| TCTCG | 756370 | 0.9467803 | 7.430053 | 41 |
| CGTCT | 755125 | 0.9452219 | 7.485845 | 16 |
| ATCGG | 706715 | 0.9125898 | 7.8744407 | 2 |
| TATGC | 1130665 | 0.9067078 | 5.0195575 | 46 |
| CTCGT | 723820 | 0.9060361 | 7.3793135 | 42 |
| ACGTC | 706015 | 0.8956078 | 7.582365 | 15 |
| CAGTC | 699700 | 0.887597 | 7.548313 | 27 |
| TCGGA | 679875 | 0.87793094 | 7.8671656 | 3 |
| AGTCA | 1059410 | 0.86096686 | 5.104779 | 28 |
| GTATG | 1052230 | 0.858957 | 5.0951724 | 45 |
| GATCG | 646555 | 0.83490443 | 7.639897 | 1 |
| CACGT | 625150 | 0.7930274 | 7.4666696 | 14 |
| TCACG | 617680 | 0.7835514 | 7.4009476 | 30 |
| ACACG | 560065 | 0.71999806 | 7.5318565 | 13 |