![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Undetermined_Undetermined_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 4790109 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
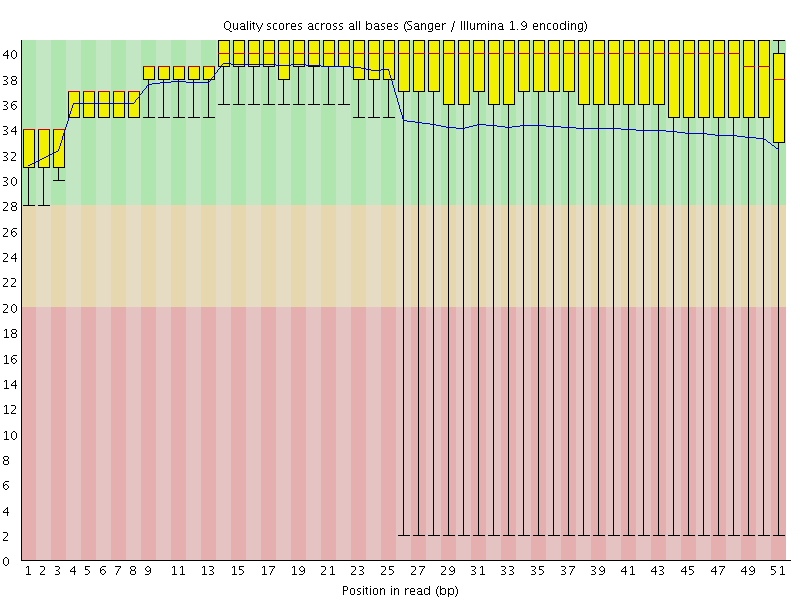
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
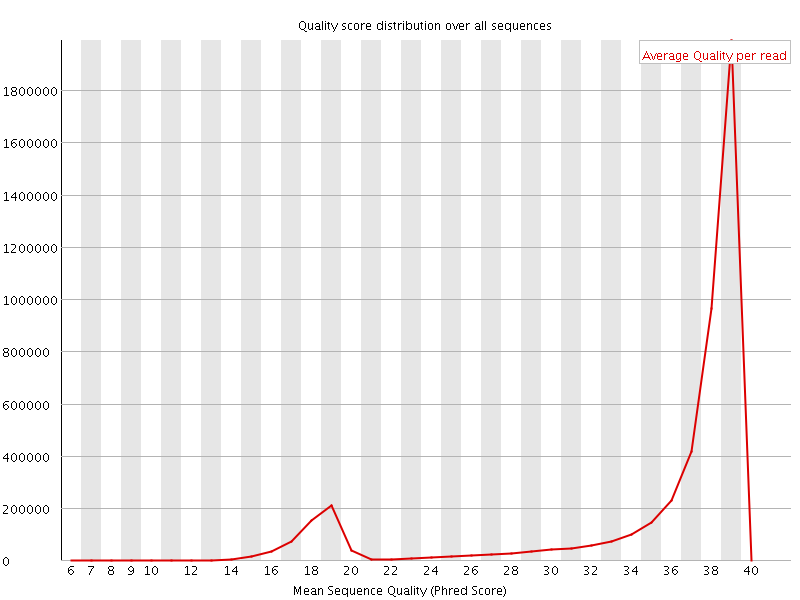
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
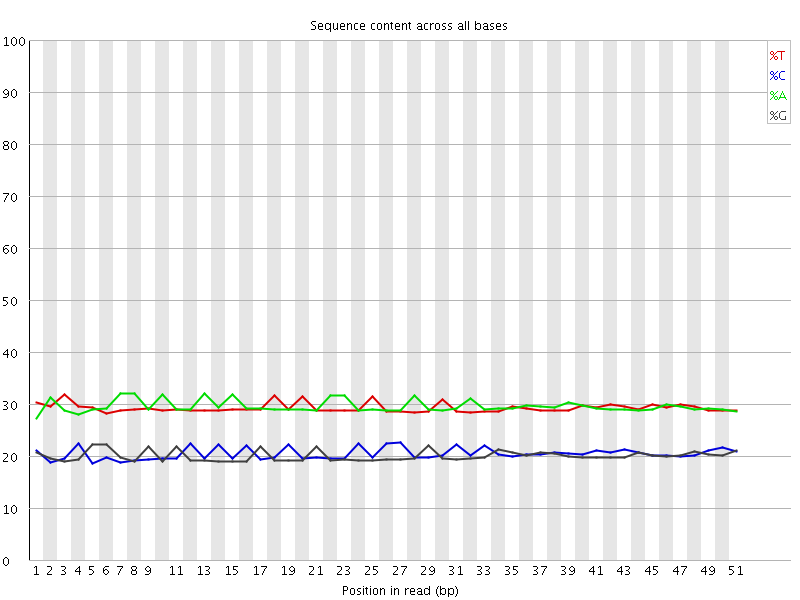
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
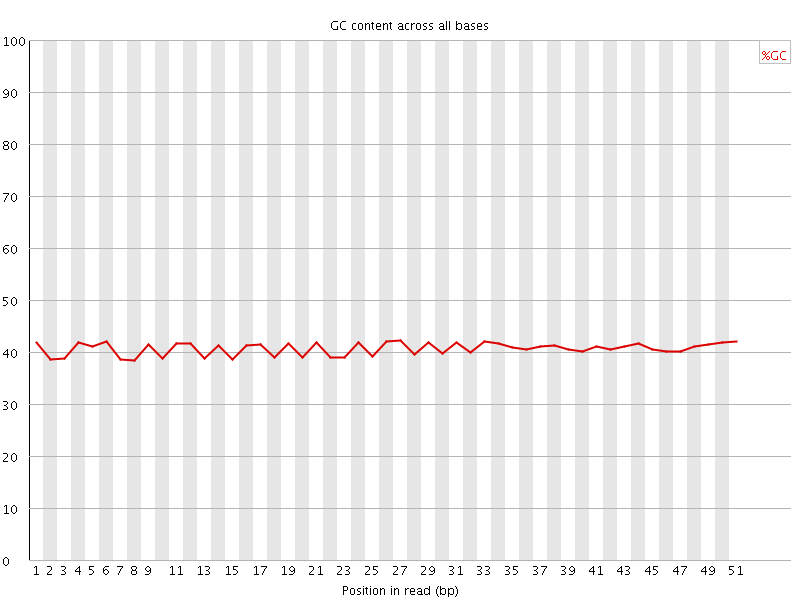
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
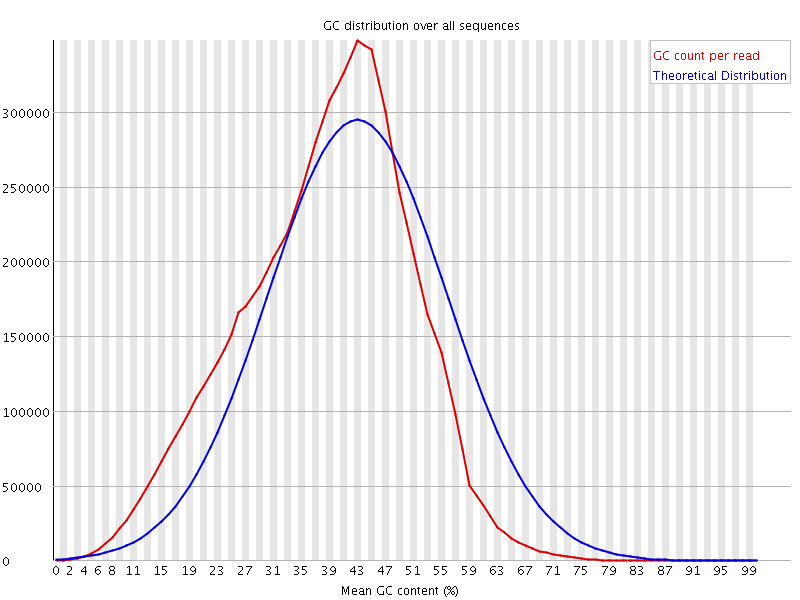
![[WARN]](Icons/warning.png) Per base N content
Per base N content
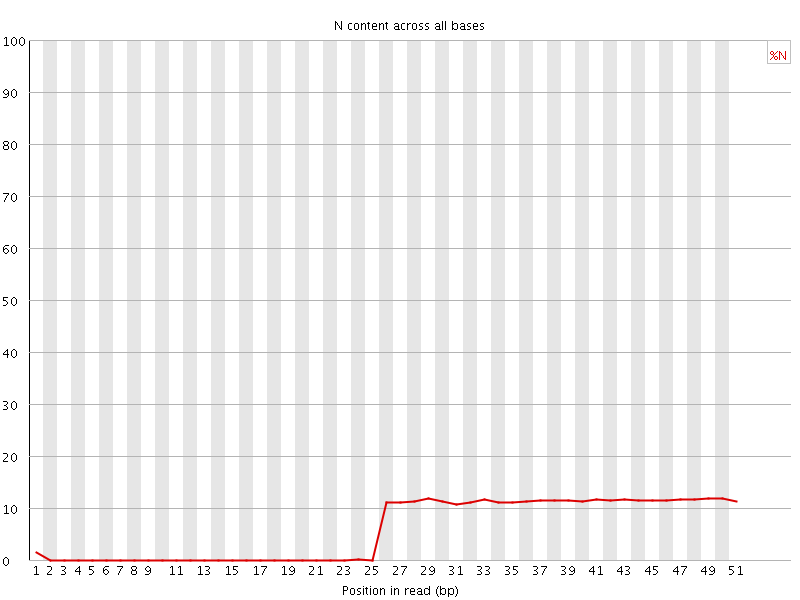
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
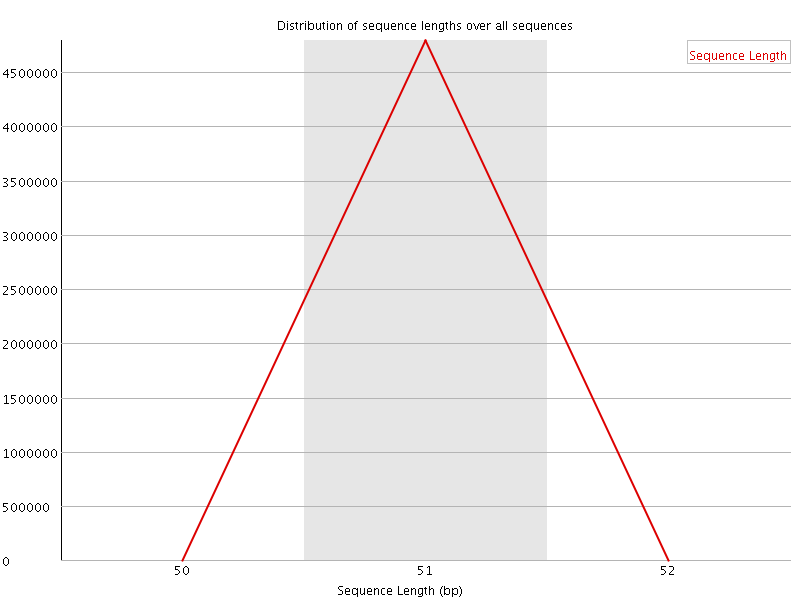
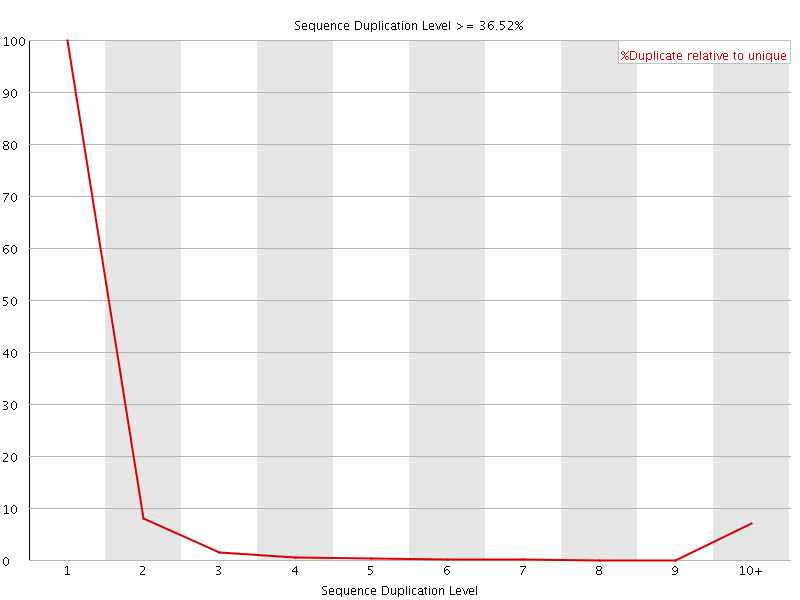
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGTAGCATCTCGTATGCCG | 9815 | 0.20490139159672566 | TruSeq Adapter, Index 3 (97% over 37bp) |
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGGAGCATCTCGTATGCCG | 8979 | 0.18744876160438104 | TruSeq Adapter, Index 2 (97% over 36bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
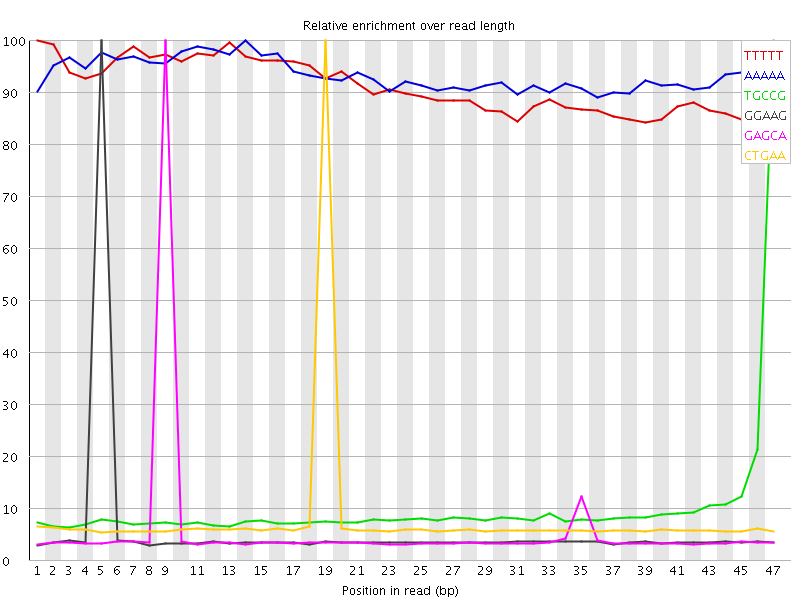
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 1488025 | 3.202745 | 3.5019643 | 1 |
| AAAAA | 1535920 | 3.131264 | 3.3451025 | 14 |
| TGCCG | 268045 | 2.5018182 | 24.855434 | 47 |
| GGAAG | 348985 | 2.2809954 | 39.65257 | 5 |
| GAGCA | 351380 | 2.247895 | 38.363155 | 9 |
| CTGAA | 485010 | 2.12719 | 26.339956 | 19 |
| CGGAA | 328270 | 2.1000524 | 38.996037 | 4 |
| CTCCA | 334785 | 2.074148 | 35.80666 | 24 |
| GAAGA | 466345 | 2.067138 | 27.310621 | 6 |
| AGAGC | 308890 | 1.9760722 | 38.253696 | 8 |
| TCCAG | 308950 | 1.9556028 | 34.385723 | 25 |
| AGCAC | 311140 | 1.9482116 | 37.36059 | 10 |
| CACGT | 296740 | 1.8783154 | 37.142067 | 14 |
| GCACA | 294805 | 1.8459294 | 37.081768 | 11 |
| AAGAG | 412950 | 1.8304573 | 27.09671 | 7 |
| TCTGA | 410195 | 1.8186882 | 26.382643 | 18 |
| CACAC | 295290 | 1.8097154 | 36.138634 | 12 |
| TCGGA | 276570 | 1.7886122 | 39.02567 | 3 |
| CGTCT | 277600 | 1.7763319 | 37.238293 | 16 |
| TCTCG | 273260 | 1.7485609 | 16.805853 | 40 |
| GTCTG | 266565 | 1.7427156 | 37.99579 | 17 |
| ATCGG | 260280 | 1.683263 | 38.305023 | 2 |
| ATGCC | 262835 | 1.6637024 | 16.58591 | 46 |
| TGAAC | 373600 | 1.6385604 | 25.761738 | 20 |
| CCAGT | 256775 | 1.6253436 | 33.113327 | 26 |
| ACGTC | 254310 | 1.6097406 | 36.857265 | 15 |
| CTCGT | 244550 | 1.5648485 | 16.510597 | 41 |
| CAGTC | 246125 | 1.557931 | 31.488798 | 27 |
| ACACG | 247875 | 1.552076 | 36.580524 | 13 |
| ACTCC | 250435 | 1.5515606 | 36.104168 | 23 |
| GTCAC | 231990 | 1.4684587 | 27.341125 | 29 |
| ATCTC | 334450 | 1.4513774 | 11.650958 | 39 |
| GATCG | 221815 | 1.434505 | 37.737232 | 1 |
| GGAGC | 149810 | 1.4131728 | 6.3243656 | 34 |
| TCACG | 217815 | 1.3787332 | 18.803131 | 30 |
| AACTC | 318760 | 1.3683611 | 25.466831 | 22 |
| GAACT | 309290 | 1.3565055 | 25.431723 | 21 |
| GCATC | 208725 | 1.321195 | 9.510385 | 37 |
| AGTCA | 297995 | 1.3069668 | 21.13508 | 28 |
| TCACC | 206995 | 1.2824298 | 6.507498 | 30 |
| GTATG | 271995 | 1.2321042 | 11.886431 | 44 |
| CATCT | 281190 | 1.2202506 | 7.4120584 | 38 |
| TATGC | 265440 | 1.1768855 | 11.576866 | 45 |
| CGGAG | 120960 | 1.1410279 | 6.4493556 | 33 |
| AGCAT | 257860 | 1.1309402 | 6.577876 | 36 |
| CGTAT | 253170 | 1.1224838 | 11.465209 | 43 |
| TCGTA | 249640 | 1.1068329 | 11.498303 | 42 |
| CACGG | 105285 | 0.9720805 | 13.141196 | 31 |
| GTAGC | 147360 | 0.9529953 | 5.667349 | 34 |