![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Undetermined_Undetermined_L003_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 4846604 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
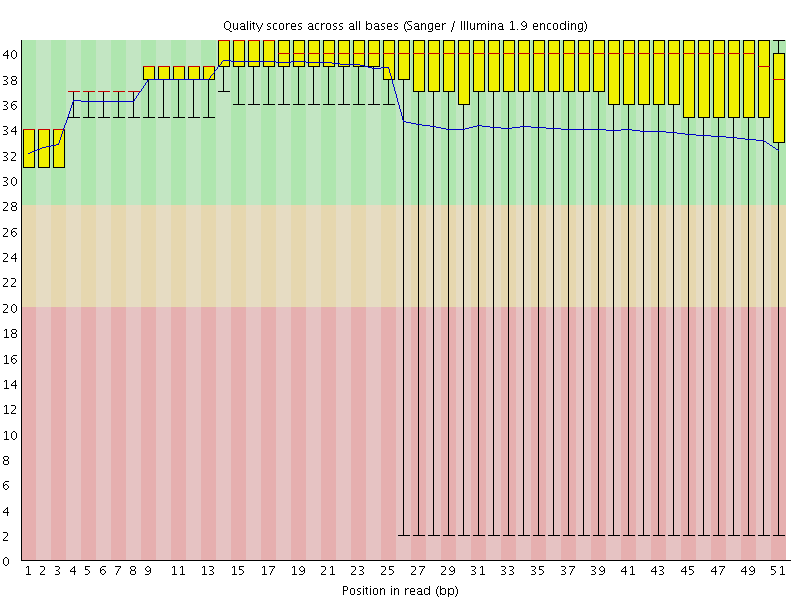
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
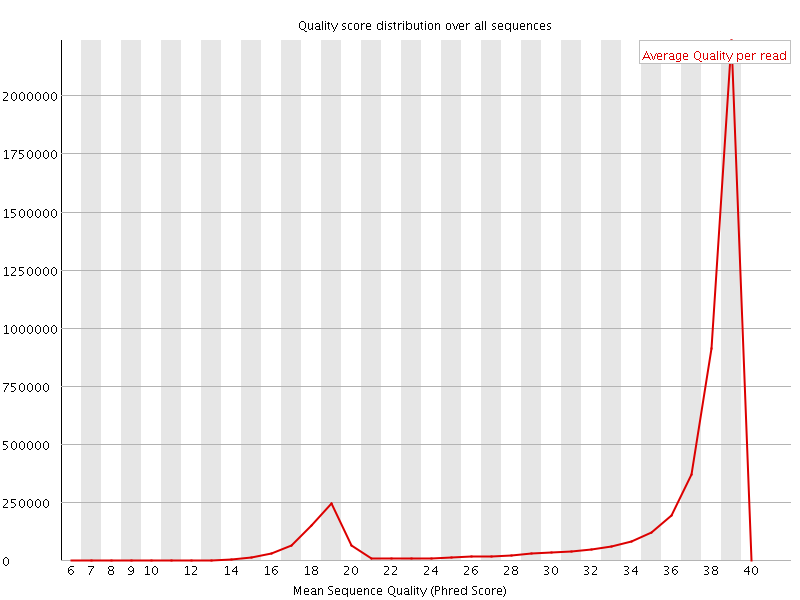
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
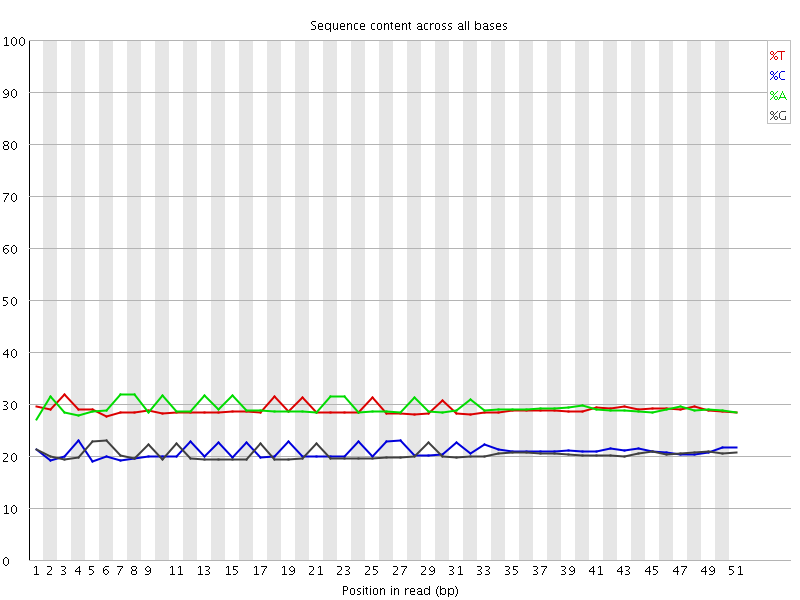
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
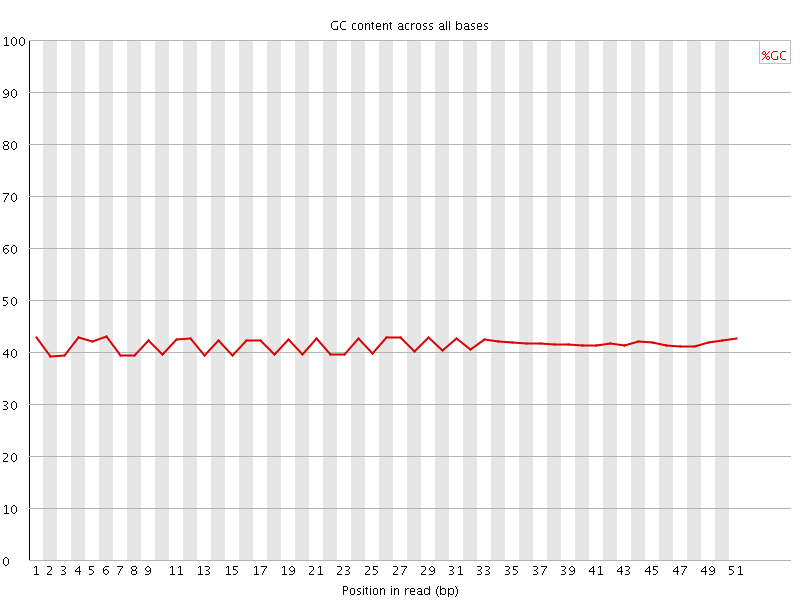
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
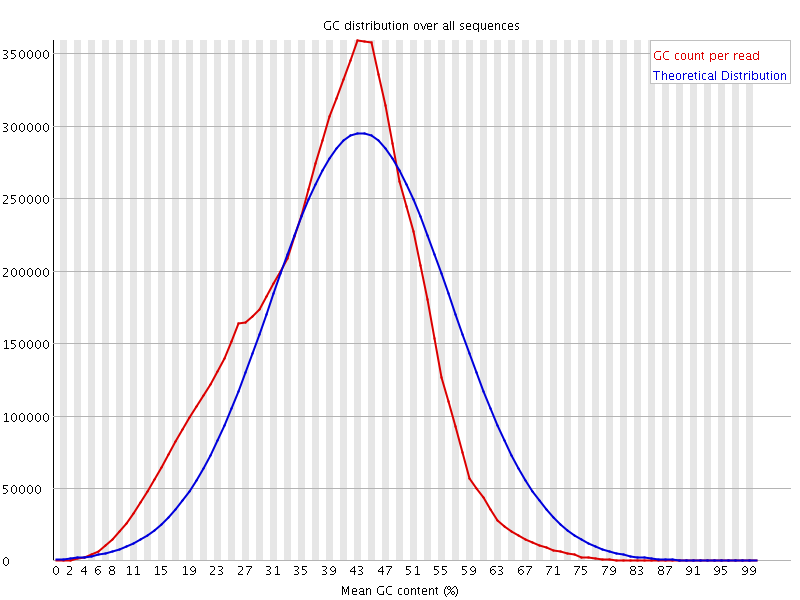
![[WARN]](Icons/warning.png) Per base N content
Per base N content
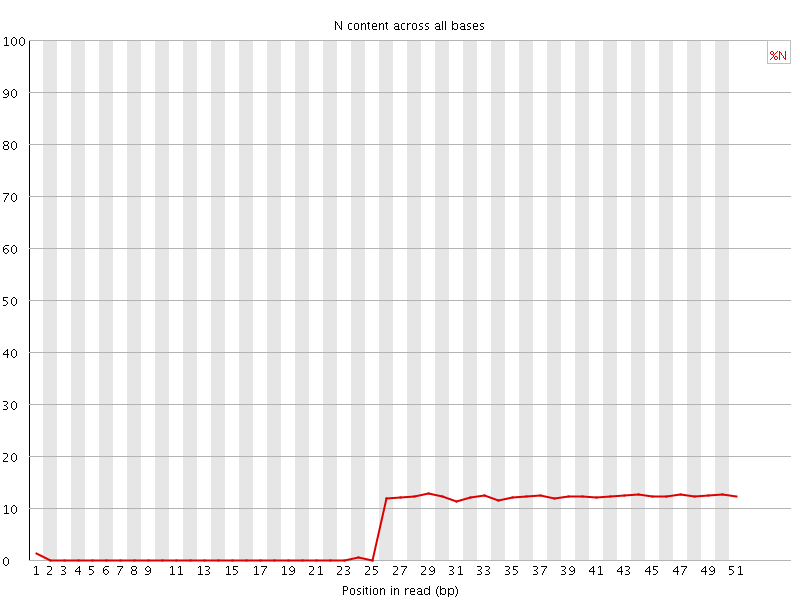
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
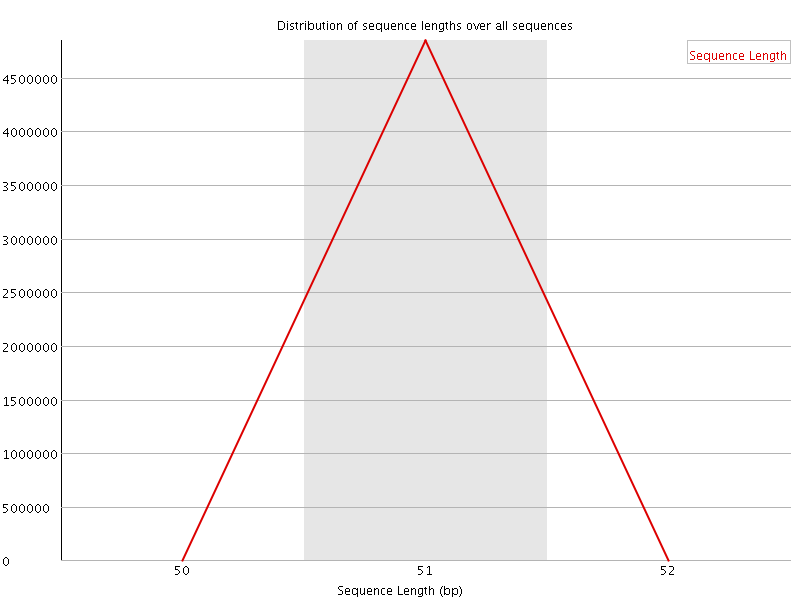
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
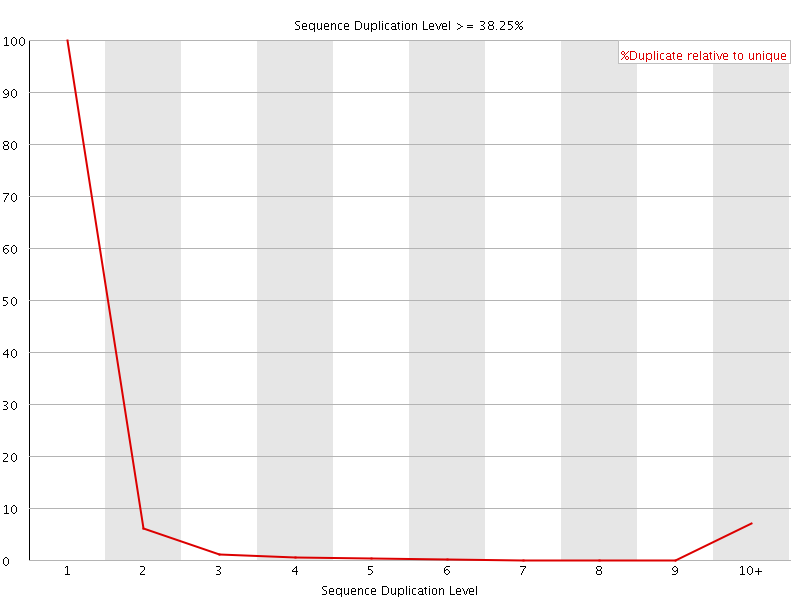
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
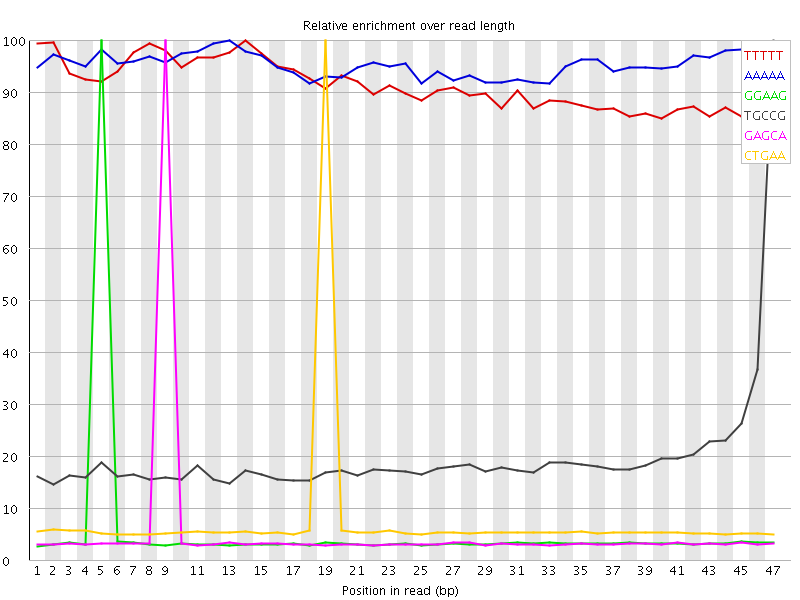
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 1425985 | 3.2550354 | 3.5512393 | 14 |
| AAAAA | 1478390 | 3.1751297 | 3.3267493 | 13 |
| GGAAG | 366230 | 2.3160172 | 41.9374 | 5 |
| TGCCG | 259030 | 2.2697458 | 11.555625 | 47 |
| GAGCA | 358270 | 2.2141237 | 40.46152 | 9 |
| CTGAA | 498790 | 2.1770296 | 28.558233 | 19 |
| CGGAA | 344700 | 2.1302605 | 40.879345 | 4 |
| GAAGA | 475725 | 2.0989654 | 29.653759 | 6 |
| CTCCA | 341805 | 2.0420706 | 36.13361 | 24 |
| AGAGC | 329515 | 2.0364165 | 40.41704 | 8 |
| AGCAC | 329785 | 1.9917092 | 39.19977 | 10 |
| TCCAG | 318905 | 1.9496202 | 35.853374 | 25 |
| AAGAG | 432175 | 1.9068168 | 29.356422 | 7 |
| TCTGA | 421365 | 1.8616531 | 28.628483 | 18 |
| CGTCT | 299585 | 1.8539689 | 39.276024 | 16 |
| GCACA | 305870 | 1.8472766 | 38.882744 | 11 |
| CACGT | 299305 | 1.8297961 | 39.010853 | 14 |
| CACAC | 308305 | 1.8196139 | 37.74233 | 12 |
| TCGGA | 290725 | 1.8187268 | 41.051865 | 3 |
| GTCTG | 280470 | 1.7760909 | 40.08498 | 17 |
| ATCGG | 281530 | 1.7612045 | 40.869045 | 2 |
| TCTCG | 281350 | 1.7411222 | 12.898943 | 41 |
| TGAAC | 384860 | 1.6797682 | 27.953001 | 20 |
| ACGTC | 270970 | 1.6565706 | 38.624493 | 15 |
| CCAGT | 267445 | 1.6350205 | 33.96934 | 26 |
| CAGTC | 260480 | 1.5924401 | 32.4603 | 27 |
| ATGCC | 259175 | 1.5844619 | 11.919875 | 47 |
| ACACG | 260675 | 1.5743251 | 38.339333 | 13 |
| CTCGT | 250955 | 1.5530242 | 11.593197 | 42 |
| GATCG | 244180 | 1.5275491 | 40.544674 | 1 |
| ACTCC | 253630 | 1.5152802 | 36.360966 | 23 |
| ATCTC | 338300 | 1.4606491 | 9.078599 | 40 |
| GTCAC | 231910 | 1.4177779 | 26.304777 | 29 |
| GAACT | 315200 | 1.3757287 | 27.518963 | 21 |
| TCACC | 228835 | 1.3671457 | 12.03871 | 30 |
| AACTC | 317805 | 1.3555357 | 26.526443 | 22 |
| AGTCA | 306130 | 1.3361416 | 22.094097 | 28 |
| GTATG | 264605 | 1.1962851 | 8.107712 | 45 |
| TCACG | 187190 | 1.144383 | 7.832561 | 30 |
| TATGC | 258800 | 1.1434168 | 7.9744887 | 46 |
| CGTAT | 254875 | 1.1260756 | 7.9781632 | 44 |
| TCGTA | 247840 | 1.094994 | 8.054189 | 43 |
| CACCG | 115855 | 0.98005825 | 5.256716 | 31 |
| CACCC | 97605 | 0.80688715 | 8.054262 | 31 |