![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Undetermined_Undetermined_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 4736347 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
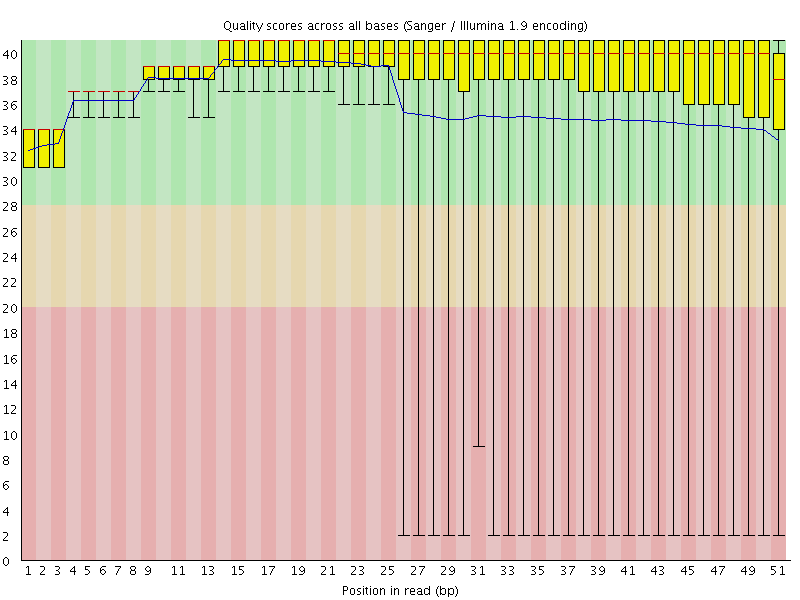
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
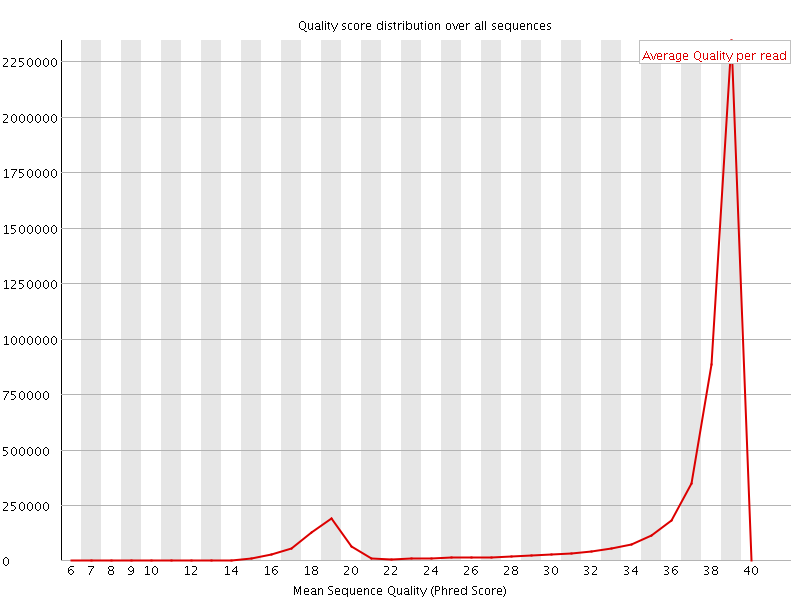
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
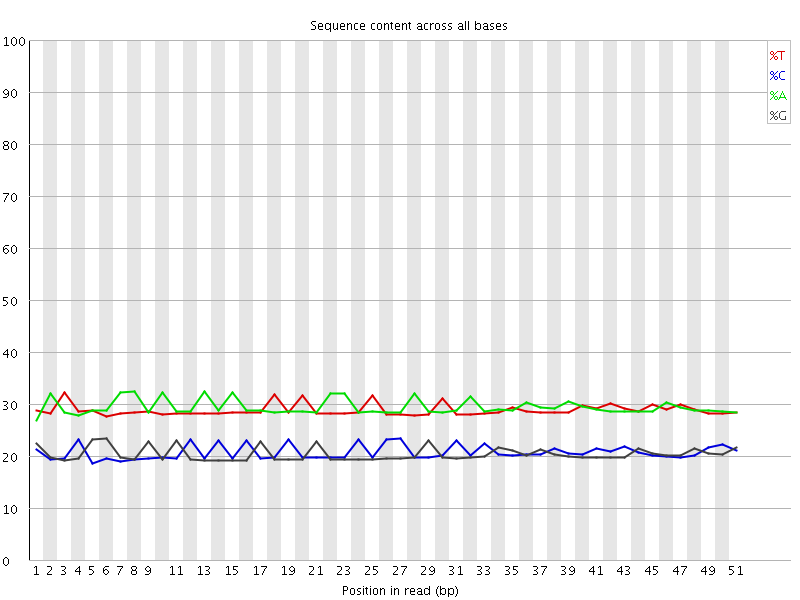
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
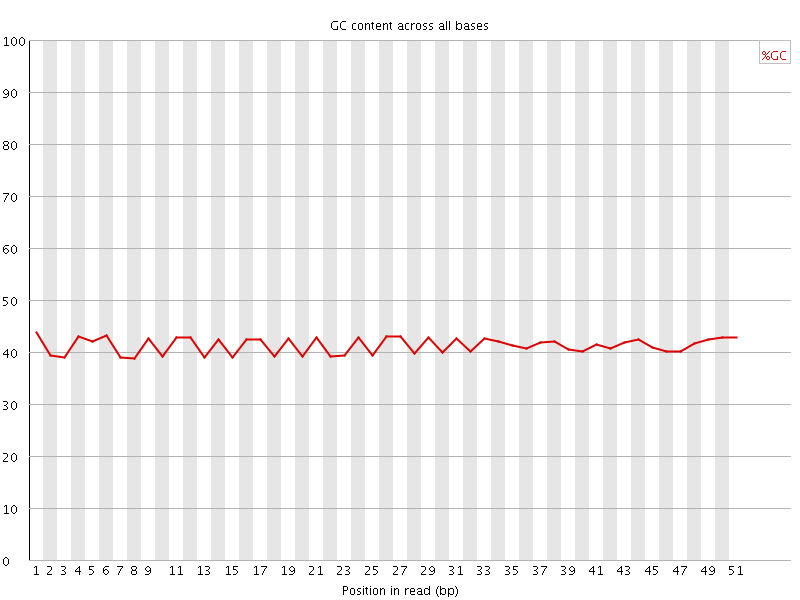
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
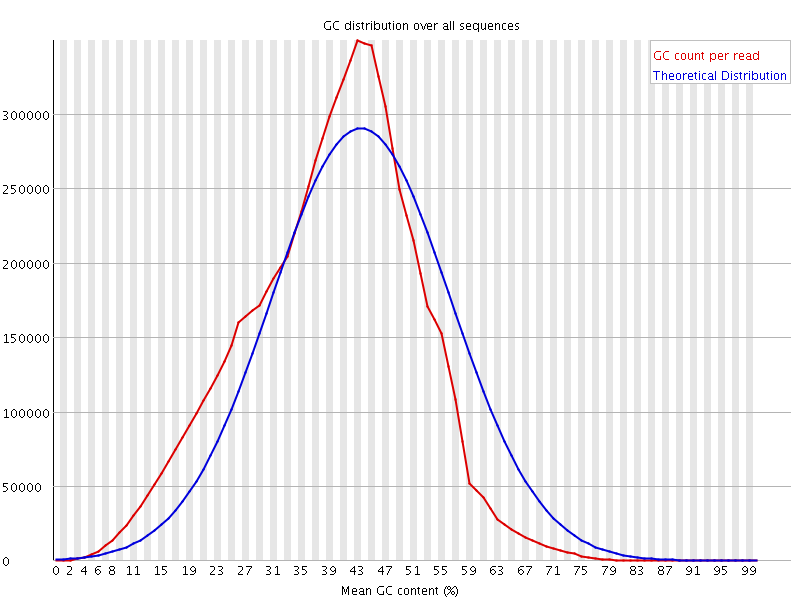
![[WARN]](Icons/warning.png) Per base N content
Per base N content
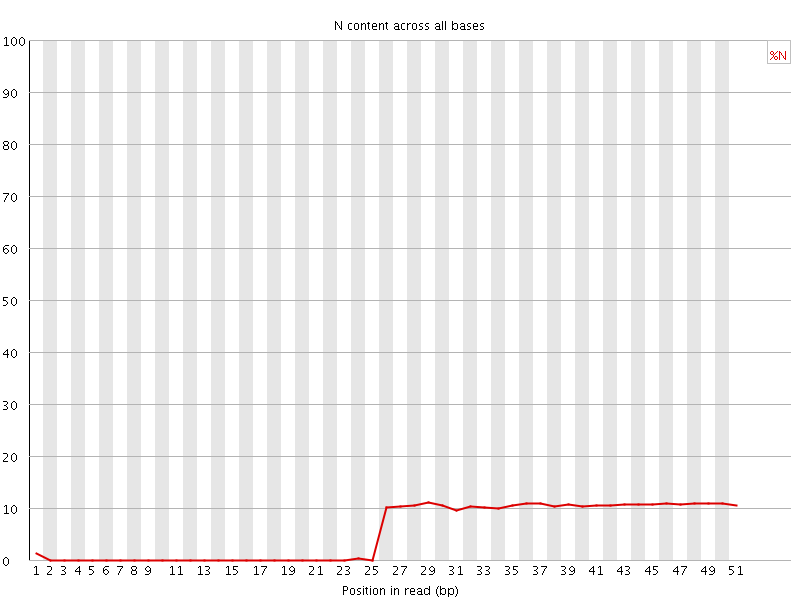
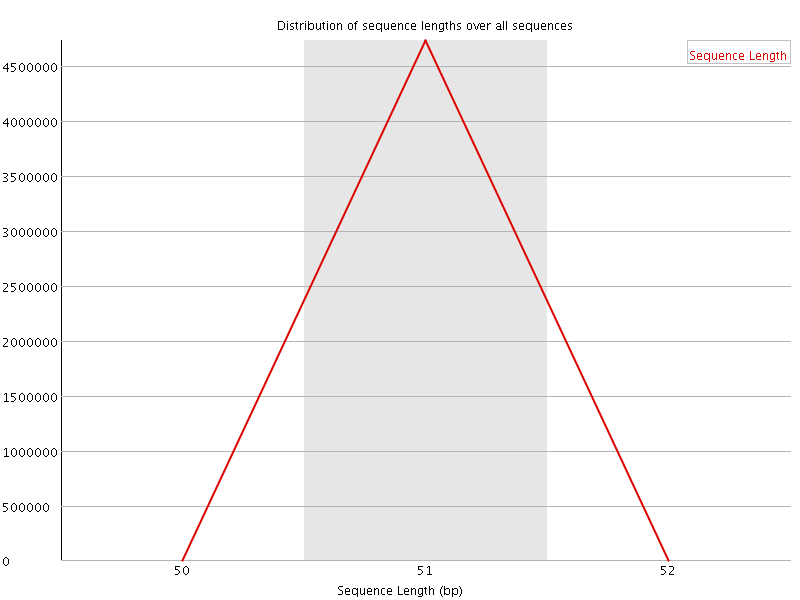
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
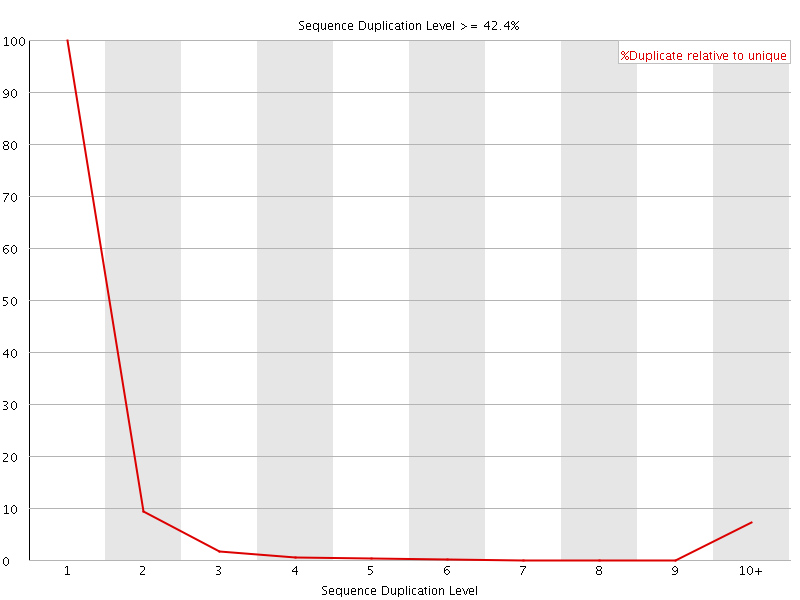
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGGAGCATCTCGTATGCCG | 21908 | 0.46255056903558794 | TruSeq Adapter, Index 2 (97% over 36bp) |
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGTAGCATCTCGTATGCCG | 19803 | 0.4181070348097384 | TruSeq Adapter, Index 3 (97% over 37bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
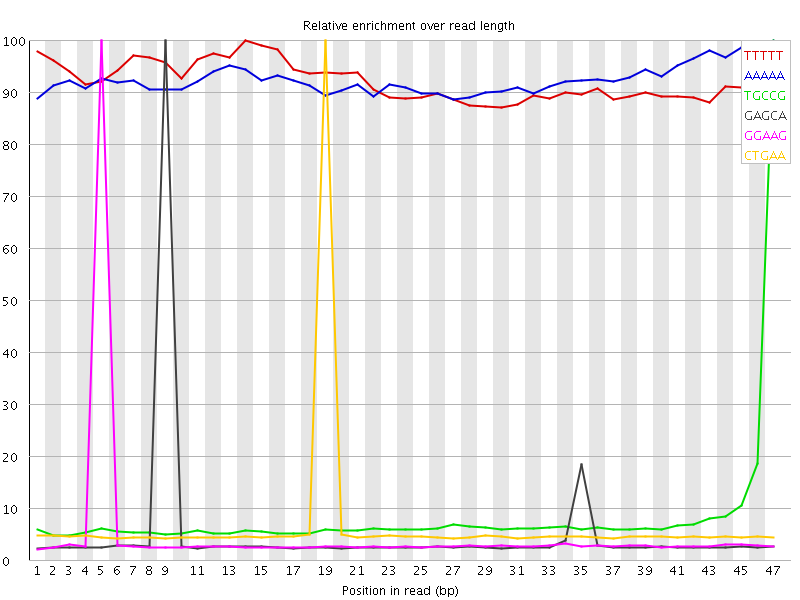
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 1495280 | 3.4931877 | 3.7861347 | 14 |
| AAAAA | 1574145 | 3.332356 | 3.6048977 | 47 |
| TGCCG | 318910 | 2.8634665 | 34.714283 | 47 |
| GAGCA | 396975 | 2.4656546 | 47.422752 | 9 |
| GGAAG | 388030 | 2.4463644 | 48.85872 | 5 |
| CTGAA | 514685 | 2.2661366 | 33.363853 | 19 |
| CGGAA | 364650 | 2.2648807 | 47.92089 | 4 |
| CTCCA | 354925 | 2.1821814 | 45.185955 | 24 |
| AGAGC | 351200 | 2.1813412 | 47.41317 | 8 |
| GAAGA | 487725 | 2.1372128 | 34.251865 | 6 |
| AGCAC | 345985 | 2.1170914 | 46.383522 | 10 |
| TCCAG | 333890 | 2.0837438 | 43.50223 | 25 |
| CACGT | 329905 | 2.0588741 | 46.5961 | 14 |
| CGTCT | 316975 | 2.017551 | 47.228207 | 16 |
| GCACA | 325035 | 1.9888979 | 46.077843 | 11 |
| AAGAG | 452220 | 1.9816296 | 34.06842 | 7 |
| CACAC | 328535 | 1.9805112 | 45.183525 | 12 |
| TCTGA | 437355 | 1.9639808 | 33.746445 | 18 |
| TCTCG | 308180 | 1.9615705 | 24.082115 | 40 |
| ATGCC | 312810 | 1.9521877 | 23.827917 | 46 |
| GTCTG | 299785 | 1.9368503 | 47.9013 | 17 |
| TCGGA | 304760 | 1.9305702 | 48.535454 | 3 |
| ATCGG | 301990 | 1.9130231 | 48.454506 | 2 |
| CCAGT | 288580 | 1.8009728 | 41.92416 | 26 |
| TGAAC | 403615 | 1.7771002 | 32.783783 | 20 |
| CTCGT | 278760 | 1.7743119 | 23.75286 | 41 |
| ACGTC | 284245 | 1.7739187 | 46.258858 | 15 |
| CAGTC | 279415 | 1.7437756 | 40.29722 | 27 |
| ACTCC | 276660 | 1.7009852 | 45.55068 | 23 |
| ACACG | 274885 | 1.6820287 | 45.51559 | 13 |
| GATCG | 263345 | 1.6682174 | 48.349434 | 1 |
| ATCTC | 368780 | 1.6314876 | 16.996443 | 39 |
| GTCAC | 255665 | 1.5955565 | 34.402885 | 29 |
| GCATC | 253425 | 1.5815772 | 18.832151 | 37 |
| GCCGT | 171000 | 1.5353949 | 5.905203 | 47 |
| TCACG | 241475 | 1.5069995 | 23.317936 | 30 |
| GAACT | 333965 | 1.470434 | 32.417736 | 21 |
| AACTC | 338100 | 1.4665707 | 32.50812 | 22 |
| AGTCA | 328610 | 1.446856 | 27.402578 | 28 |
| GTATG | 313965 | 1.4311036 | 17.180798 | 44 |
| GGAGC | 156830 | 1.4014618 | 12.037699 | 34 |
| TATGC | 308420 | 1.3849869 | 17.029274 | 45 |
| CATCT | 306500 | 1.35596 | 14.151691 | 38 |
| AGCAT | 297055 | 1.3079208 | 12.954967 | 36 |
| CGTAT | 289550 | 1.3002496 | 16.767775 | 43 |
| TCGTA | 277445 | 1.245891 | 16.753363 | 42 |
| CGGAG | 132680 | 1.185653 | 12.102124 | 33 |
| GTAGC | 175040 | 1.10883 | 10.525656 | 34 |
| CACGG | 121525 | 1.0698701 | 18.608381 | 31 |
| ACGGA | 155495 | 0.96579623 | 9.076279 | 32 |
| TAGCA | 203075 | 0.89413077 | 7.387507 | 35 |
| CGTAG | 83485 | 0.52885437 | 7.4229946 | 33 |
| ACGTA | 114685 | 0.50495327 | 5.3768034 | 32 |