![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW244_ATGTCA_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 7106634 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
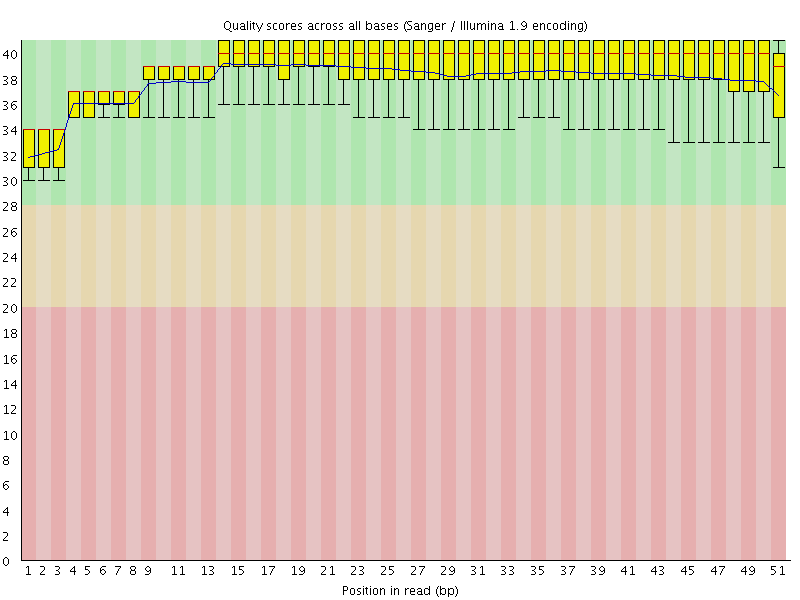
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
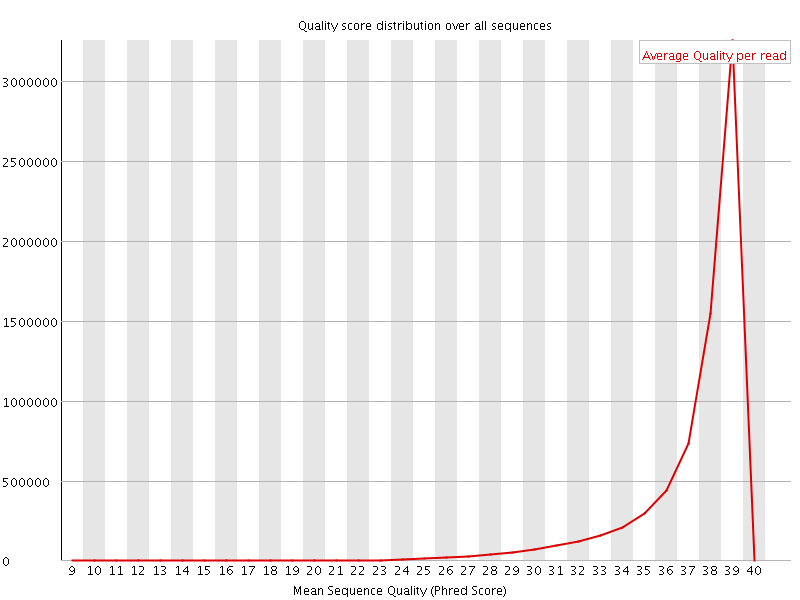
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
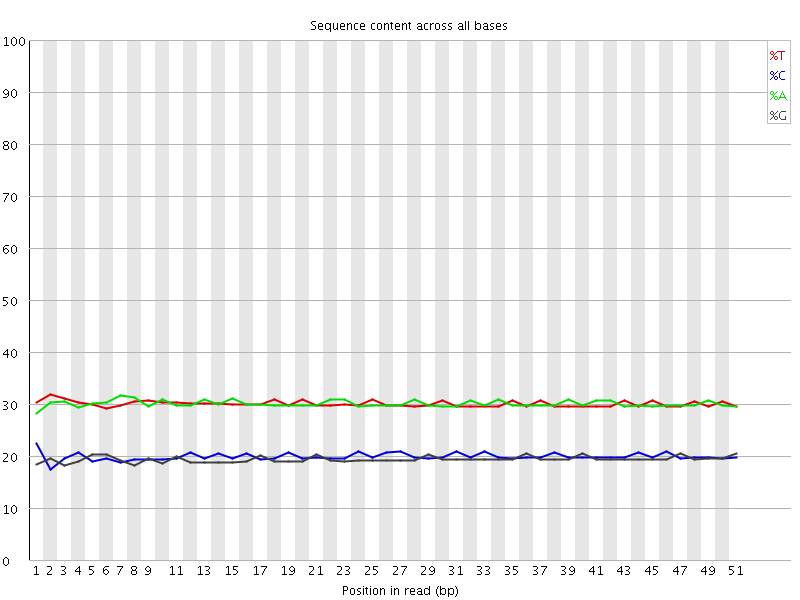
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
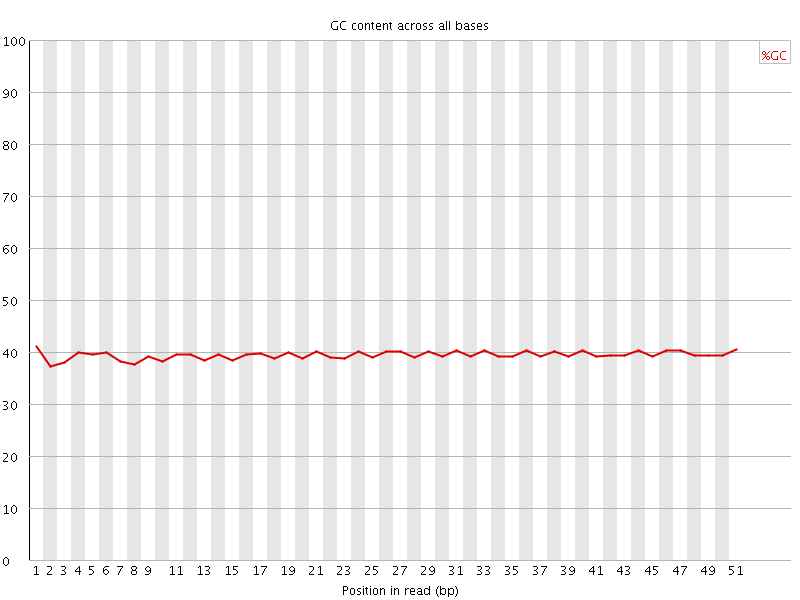
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
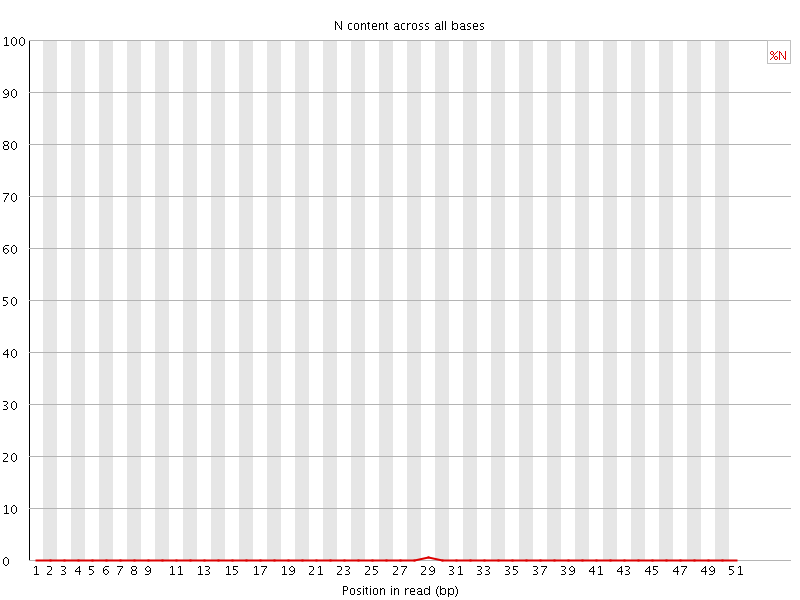
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
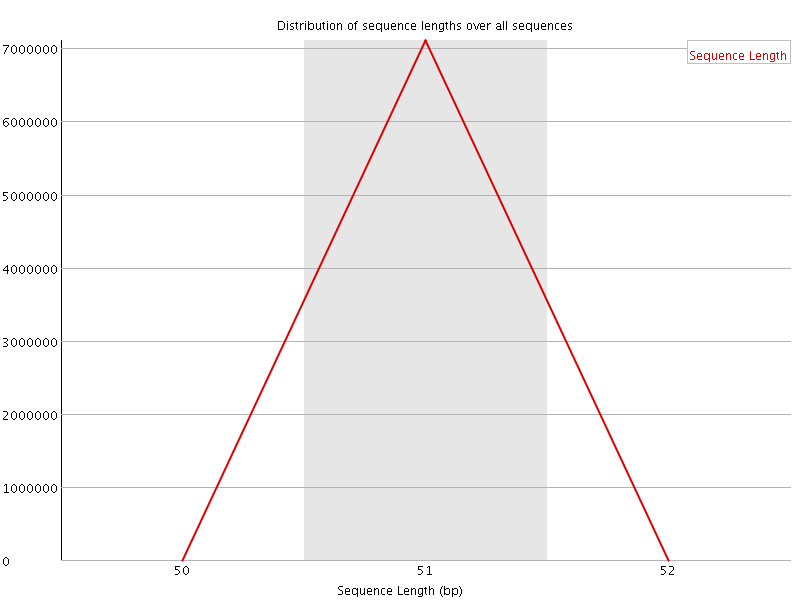
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
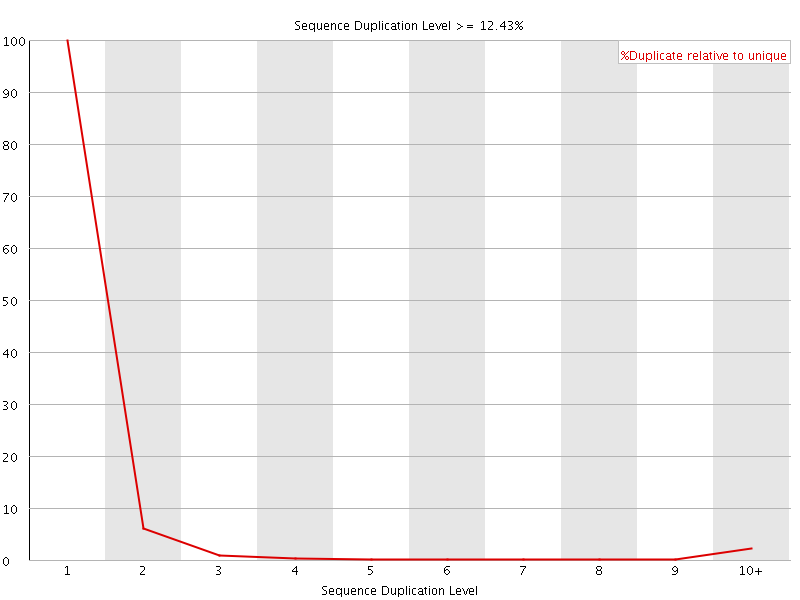
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACATGTCAGAATCTCGTATG | 67134 | 0.9446666312068414 | TruSeq Adapter, Index 1 (97% over 35bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
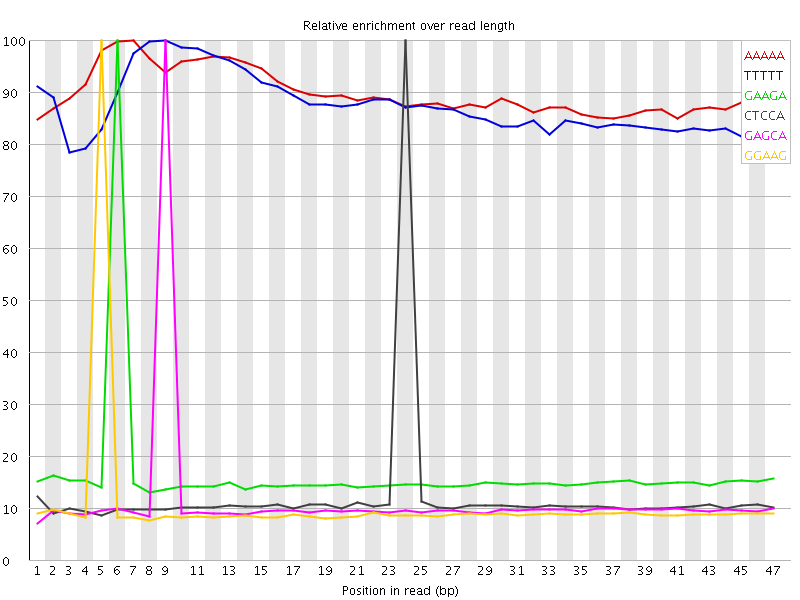
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 2663490 | 3.1673365 | 3.5250106 | 7 |
| TTTTT | 2635345 | 3.1360211 | 3.5838673 | 9 |
| GAAGA | 745145 | 2.1242838 | 12.809804 | 6 |
| CTCCA | 510765 | 2.0802696 | 16.925629 | 24 |
| GAGCA | 478510 | 2.056181 | 17.93211 | 9 |
| GGAAG | 451420 | 1.9925851 | 18.543642 | 5 |
| TCCAG | 450780 | 1.8859444 | 17.088581 | 25 |
| CTGAA | 674345 | 1.8717501 | 12.149798 | 19 |
| CAGAA | 653805 | 1.8144886 | 11.603837 | 38 |
| CGGAA | 417310 | 1.7932017 | 17.926035 | 4 |
| TCTCG | 388520 | 1.6256883 | 16.652134 | 43 |
| TCGGA | 365425 | 1.570465 | 17.72771 | 3 |
| AGAGC | 362375 | 1.5571431 | 17.444841 | 8 |
| TCAGA | 556405 | 1.5443892 | 11.352004 | 37 |
| AGCAC | 367430 | 1.5370189 | 17.009962 | 10 |
| CACAC | 377085 | 1.5355998 | 16.562958 | 12 |
| TCTGA | 547830 | 1.5207969 | 11.821993 | 18 |
| GCACA | 353895 | 1.4803997 | 16.949072 | 11 |
| AAGAG | 505475 | 1.4410248 | 12.09298 | 7 |
| TCACA | 507690 | 1.3718244 | 11.10549 | 30 |
| TGAAC | 487980 | 1.3544648 | 11.658073 | 20 |
| CCAGT | 321740 | 1.346075 | 16.53603 | 26 |
| ATCGG | 312230 | 1.3418521 | 17.201036 | 2 |
| CTCGT | 319450 | 1.336678 | 16.191017 | 44 |
| CGTCT | 311435 | 1.3031408 | 16.79577 | 16 |
| CACGT | 308445 | 1.2904524 | 16.759865 | 14 |
| GTCAC | 307090 | 1.2847832 | 16.436275 | 29 |
| TGTCA | 459295 | 1.2750202 | 11.1368885 | 35 |
| GAATC | 458965 | 1.2739291 | 11.126242 | 40 |
| GTCAG | 295540 | 1.2701243 | 16.568132 | 36 |
| GTCTG | 292945 | 1.259145 | 17.213402 | 17 |
| ACTCC | 307810 | 1.2536641 | 16.136429 | 23 |
| AACTC | 454720 | 1.2286946 | 11.188406 | 22 |
| CACAT | 447690 | 1.2096989 | 10.824349 | 31 |
| ACACG | 288750 | 1.2078876 | 16.629889 | 13 |
| ATCTC | 446750 | 1.2073249 | 10.8512945 | 42 |
| GAACT | 425745 | 1.1817218 | 11.471669 | 21 |
| ACGTC | 282130 | 1.1803572 | 16.70113 | 15 |
| GATCG | 272795 | 1.1723745 | 16.879593 | 1 |
| AGAAT | 635170 | 1.1696532 | 7.7113585 | 39 |
| CAGTC | 277660 | 1.1616559 | 16.298798 | 27 |
| AGTCA | 379165 | 1.0524317 | 11.115634 | 28 |
| CATGT | 378240 | 1.0500085 | 10.991539 | 33 |
| ACATG | 376290 | 1.0444517 | 11.005223 | 32 |
| ATGTC | 375080 | 1.0412363 | 10.91233 | 34 |
| AATCT | 568180 | 1.0187016 | 7.3309913 | 41 |
| TCGTA | 336425 | 0.93392855 | 10.811794 | 45 |
| CGTAT | 286785 | 0.79612607 | 10.650888 | 46 |
| GTATG | 276015 | 0.78708893 | 10.831784 | 47 |