![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW243_AGTTCC_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 6921054 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 38 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
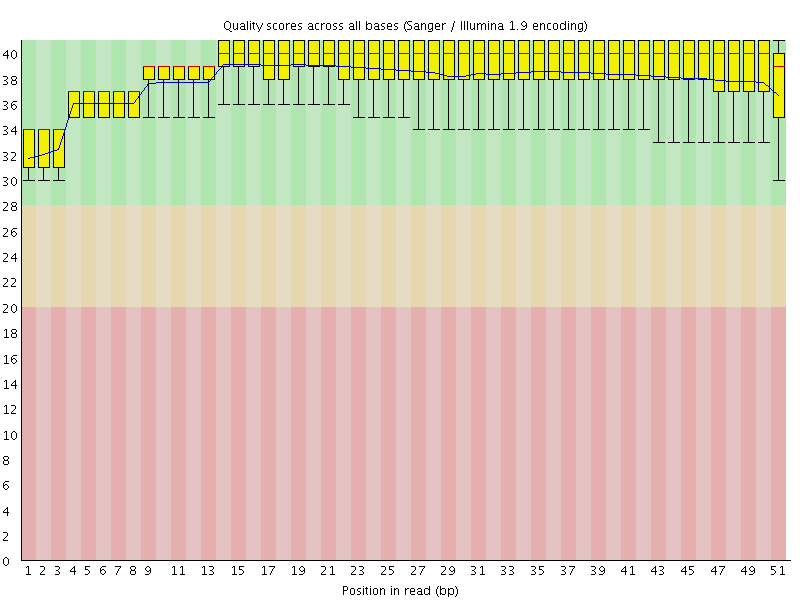
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
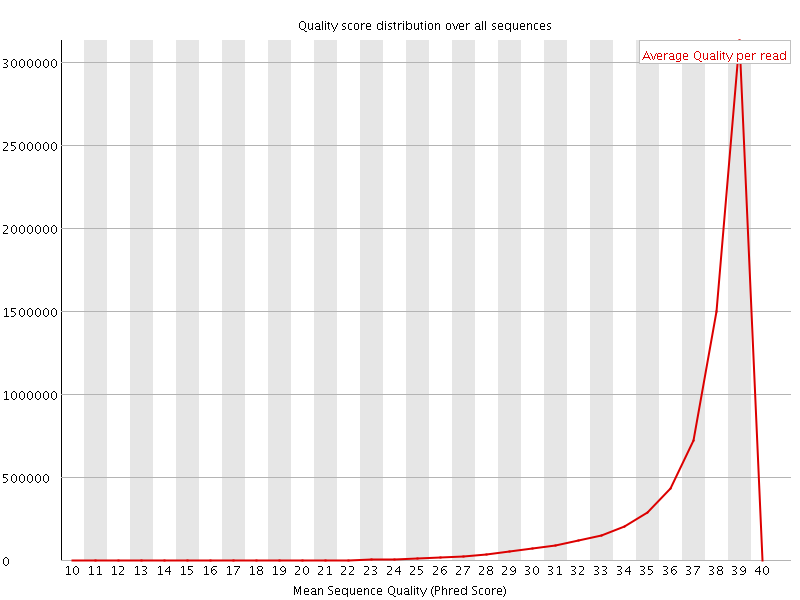
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
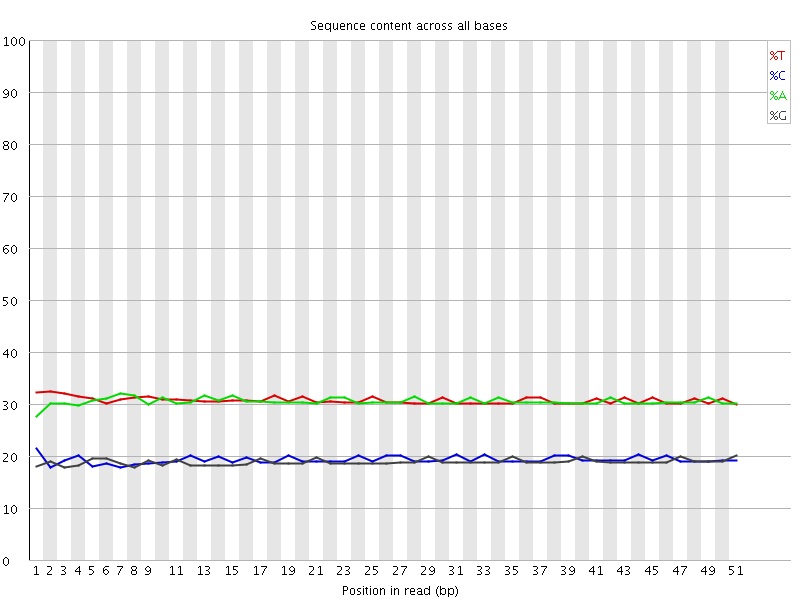
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
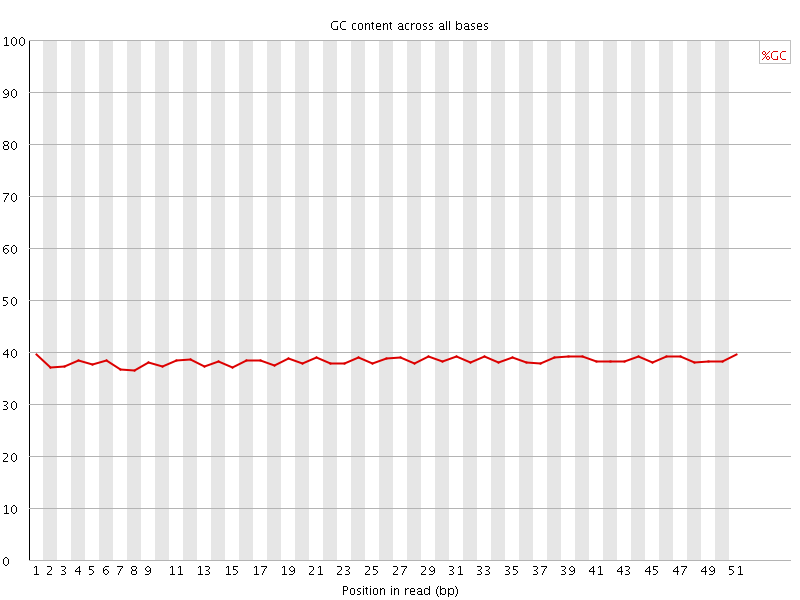
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
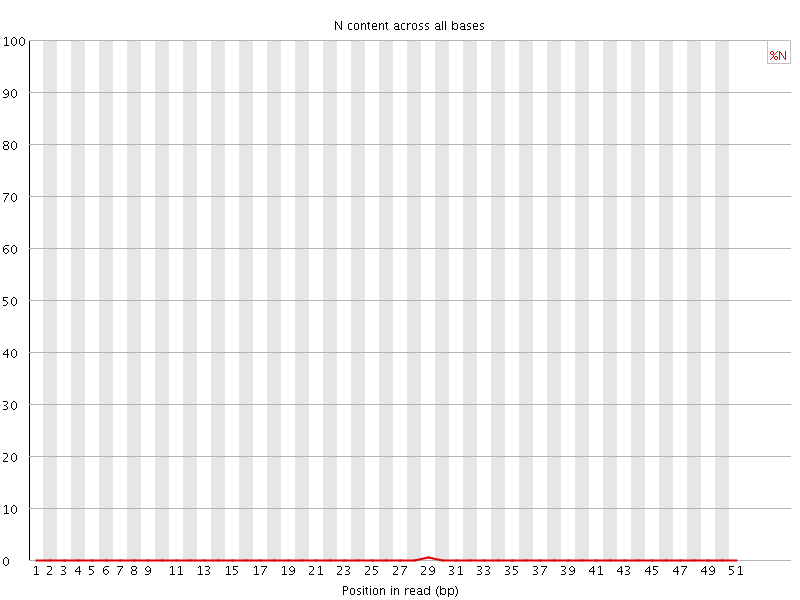
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
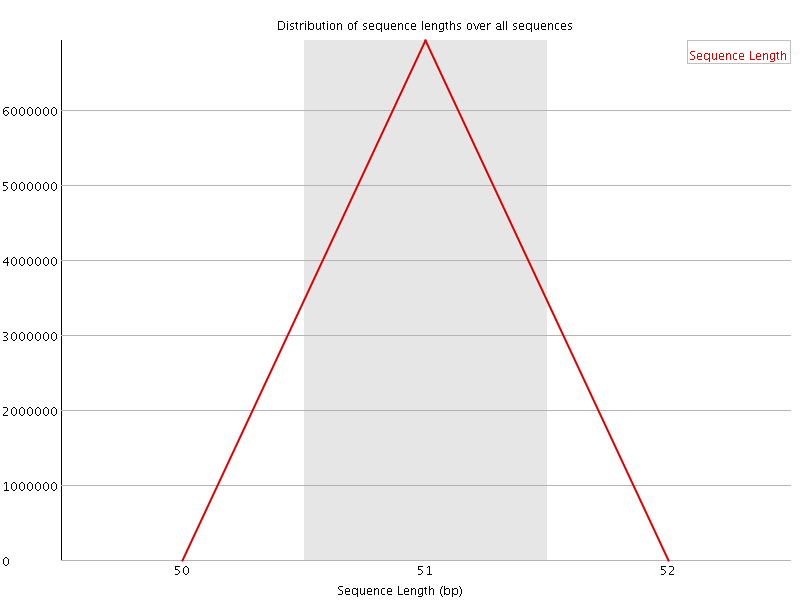
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACAGTTCCGTATCTCGTATG | 67700 | 0.9781747115396008 | TruSeq Adapter, Index 8 (97% over 37bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
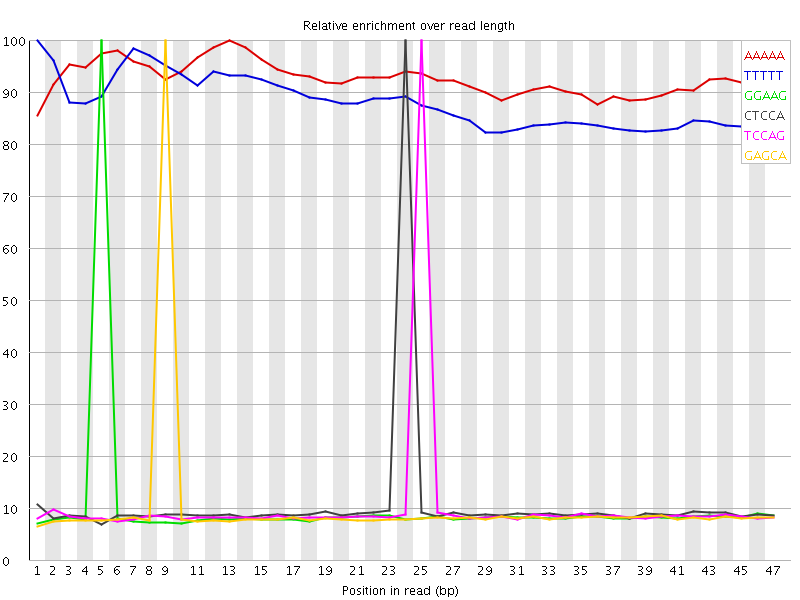
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 3235220 | 3.672909 | 3.9640453 | 13 |
| TTTTT | 3254130 | 3.5534687 | 4.0342226 | 1 |
| GGAAG | 423985 | 2.0173655 | 19.87458 | 5 |
| CTCCA | 445845 | 1.9775242 | 18.236933 | 24 |
| TCCAG | 427720 | 1.9370327 | 18.548485 | 25 |
| GAGCA | 413435 | 1.9266456 | 19.190561 | 9 |
| CTGAA | 668350 | 1.9168063 | 12.575057 | 19 |
| GAAGA | 642455 | 1.8959818 | 12.852141 | 6 |
| CGGAA | 394510 | 1.8384533 | 19.397318 | 4 |
| TTCCG | 397695 | 1.7871037 | 17.546288 | 36 |
| CACAG | 344385 | 1.5718074 | 17.839298 | 31 |
| TCTCG | 339205 | 1.5242699 | 17.476612 | 43 |
| CACAC | 339210 | 1.516297 | 18.08587 | 12 |
| TCTGA | 525485 | 1.4953983 | 12.04026 | 18 |
| AGAGC | 320865 | 1.4952607 | 18.787931 | 8 |
| GCACA | 325350 | 1.4849299 | 18.438398 | 11 |
| AGCAC | 323435 | 1.4761895 | 18.53698 | 10 |
| TCGGA | 318495 | 1.4727176 | 18.839931 | 3 |
| CAGTT | 497885 | 1.4168555 | 11.482588 | 33 |
| CCAGT | 312275 | 1.4142121 | 17.932198 | 26 |
| GTTCC | 313745 | 1.4098616 | 17.231281 | 35 |
| AAGAG | 453835 | 1.3393357 | 12.315163 | 7 |
| ATCGG | 285145 | 1.3185076 | 18.39157 | 2 |
| TGAAC | 457065 | 1.3108475 | 11.947646 | 20 |
| TCACA | 466490 | 1.3103192 | 11.329895 | 30 |
| GTCTG | 279510 | 1.2824382 | 18.320013 | 17 |
| ACTCC | 287010 | 1.2730191 | 17.645649 | 23 |
| GTCAC | 275670 | 1.2484376 | 17.584557 | 29 |
| TCCGT | 271170 | 1.2185441 | 16.9781 | 37 |
| CTCGT | 270830 | 1.2170162 | 17.106274 | 44 |
| CACGT | 265160 | 1.2008406 | 17.995707 | 14 |
| CGTCT | 267175 | 1.200592 | 17.859661 | 16 |
| AACTC | 425050 | 1.193919 | 11.546726 | 22 |
| ACAGT | 416205 | 1.1936625 | 11.385424 | 32 |
| GAACT | 403955 | 1.1585298 | 11.732816 | 21 |
| CAGTC | 253940 | 1.1500282 | 17.641176 | 27 |
| GATCG | 248660 | 1.1498013 | 18.069016 | 1 |
| ACACG | 251215 | 1.1465703 | 18.159792 | 13 |
| ATCTC | 410550 | 1.1442556 | 10.986891 | 42 |
| AGTTC | 398955 | 1.1353254 | 11.176195 | 34 |
| ACGTC | 243660 | 1.1034728 | 17.942753 | 15 |
| AGTCA | 350475 | 1.005151 | 11.396804 | 28 |
| CCGTA | 214210 | 0.9701015 | 16.73512 | 38 |
| CGTAT | 328055 | 0.9335621 | 10.719224 | 46 |
| TCGTA | 279950 | 0.7966673 | 10.832664 | 45 |
| GTATC | 272125 | 0.7743992 | 10.769484 | 40 |
| GTATG | 260990 | 0.7583328 | 10.975018 | 47 |
| TATCT | 382850 | 0.67050934 | 6.826267 | 41 |