![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW241_CTTGTA_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 9405925 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
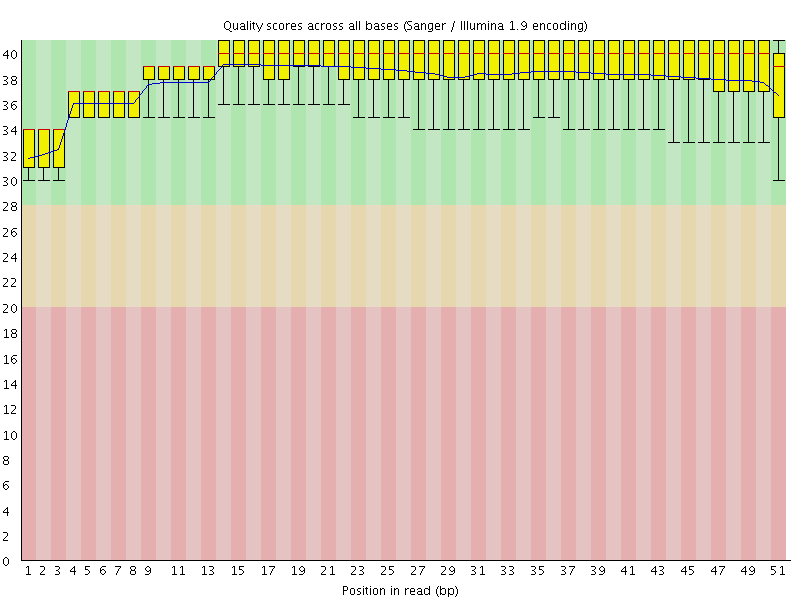
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
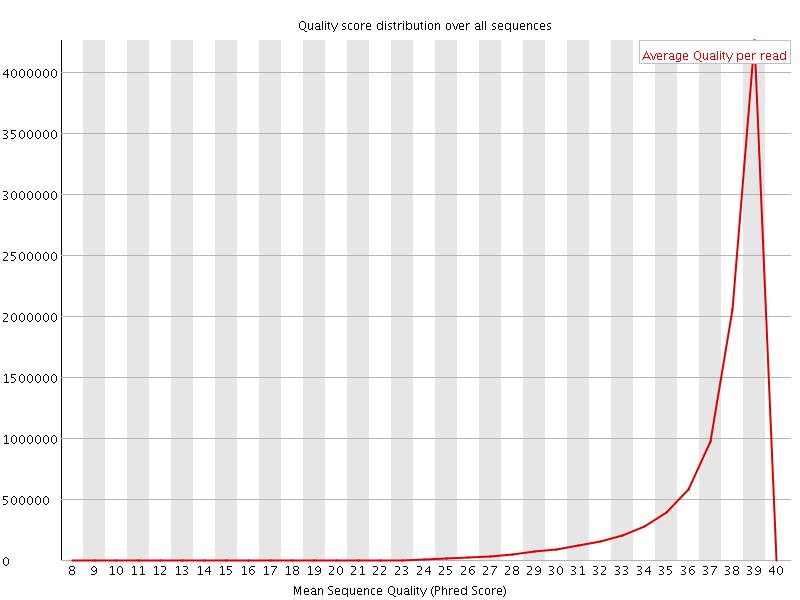
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
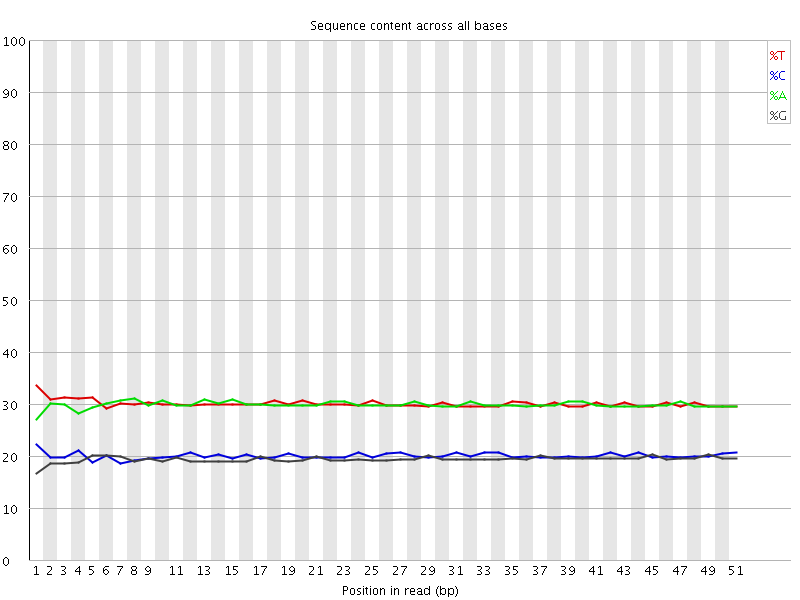
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
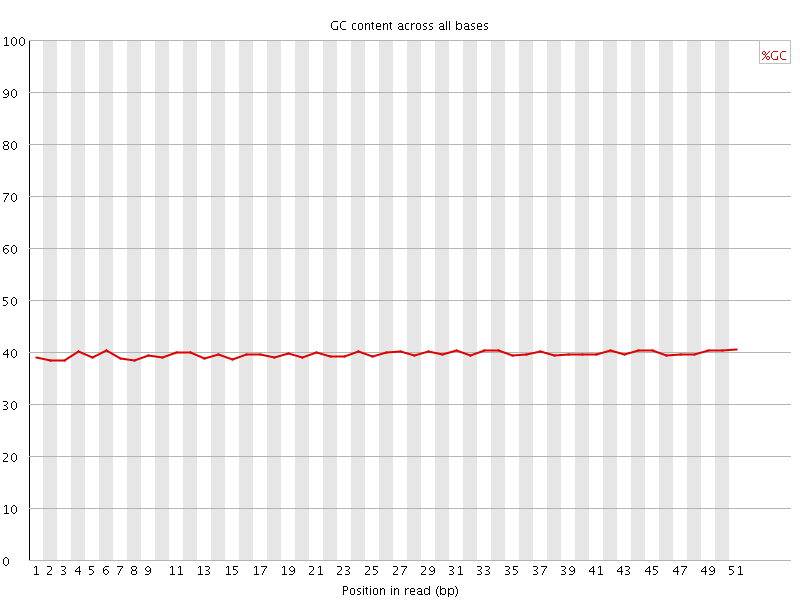
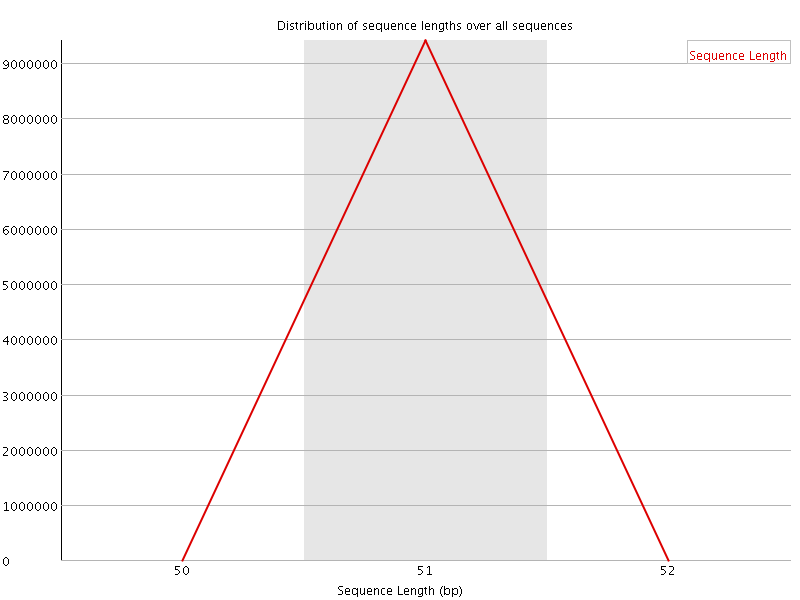
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
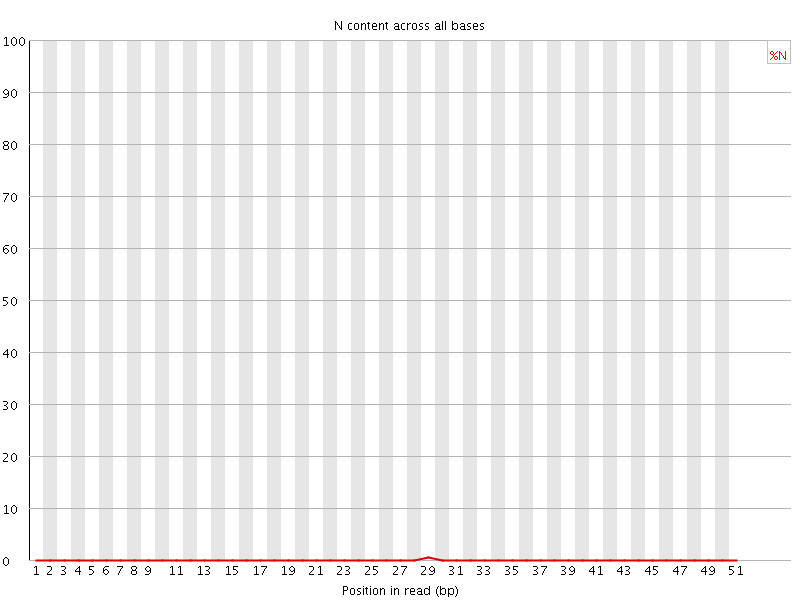
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
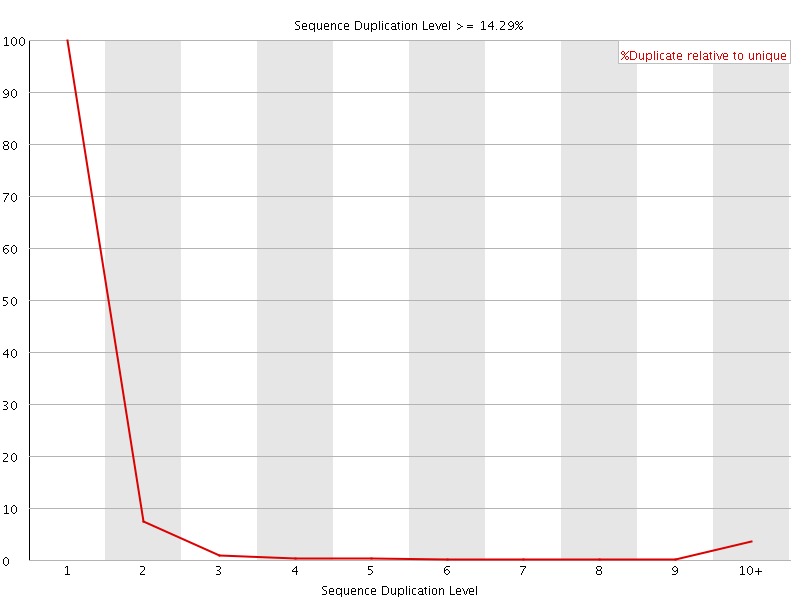
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCTTGTAATCTCGTATGCC | 67435 | 0.7169417149296853 | TruSeq Adapter, Index 12 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
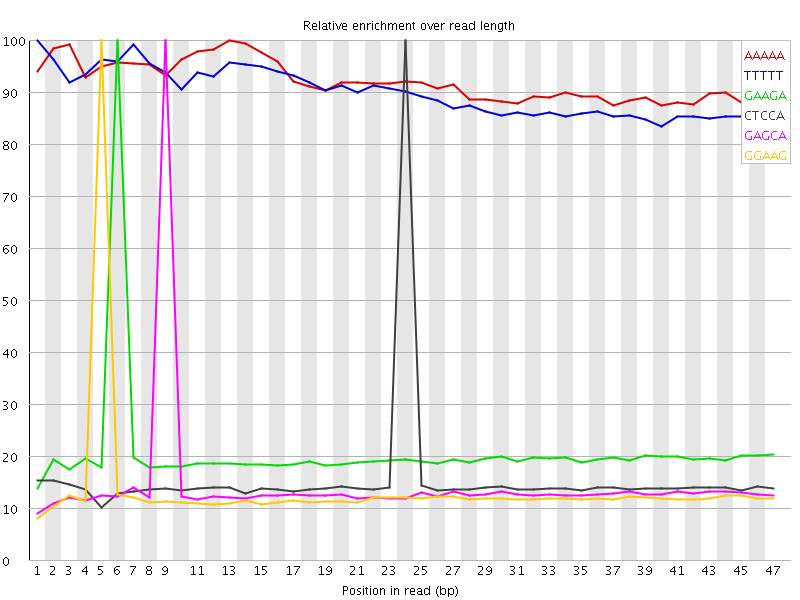
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 3475550 | 3.2358778 | 3.512485 | 13 |
| TTTTT | 3483730 | 3.1104057 | 3.4565105 | 1 |
| GAAGA | 937400 | 2.0577683 | 9.879349 | 6 |
| CTCCA | 653940 | 1.9793838 | 12.593816 | 24 |
| GAGCA | 602275 | 1.9640565 | 13.628137 | 9 |
| GGAAG | 568025 | 1.9146547 | 14.070584 | 5 |
| CTGAA | 863890 | 1.8193945 | 9.34436 | 19 |
| TCCAG | 575845 | 1.8016133 | 12.888811 | 25 |
| CGGAA | 527470 | 1.7201127 | 13.6181755 | 4 |
| TCTCG | 486470 | 1.5092906 | 12.358283 | 41 |
| TCACC | 496135 | 1.5017302 | 12.046932 | 30 |
| TCGGA | 463200 | 1.4979194 | 13.33371 | 3 |
| TCTGA | 696595 | 1.4548209 | 8.901785 | 18 |
| AGAGC | 442505 | 1.4430366 | 13.13737 | 8 |
| CACAC | 466655 | 1.4243845 | 12.327999 | 12 |
| AGCAC | 448300 | 1.4143728 | 12.75885 | 10 |
| AAGAG | 630300 | 1.3836262 | 9.224813 | 7 |
| GCACA | 437990 | 1.3818454 | 12.697372 | 11 |
| TGAAC | 620115 | 1.3059924 | 8.829419 | 20 |
| CCAGT | 404630 | 1.2659428 | 12.2501 | 26 |
| ATCGG | 384290 | 1.2427365 | 12.815912 | 2 |
| CTCGT | 394315 | 1.2233764 | 11.913616 | 42 |
| CACGT | 381095 | 1.1923102 | 12.398234 | 14 |
| CGTCT | 382000 | 1.1851686 | 12.267162 | 16 |
| GTCAC | 378000 | 1.182627 | 12.067558 | 29 |
| CACCT | 389680 | 1.1795062 | 11.67106 | 31 |
| AACTC | 574075 | 1.1696962 | 8.421771 | 22 |
| CCTTG | 374500 | 1.1618997 | 11.896942 | 33 |
| ACTCC | 381600 | 1.1550491 | 11.913745 | 23 |
| GTCTG | 358005 | 1.1480738 | 12.552884 | 17 |
| CTTGT | 551310 | 1.1417891 | 8.263968 | 34 |
| ATCTC | 564300 | 1.1401851 | 8.159203 | 40 |
| GAACT | 535020 | 1.1267782 | 8.651882 | 21 |
| ACACG | 350835 | 1.1068739 | 12.435158 | 13 |
| ATGCC | 351760 | 1.1005315 | 11.937047 | 47 |
| GATCG | 334850 | 1.0828549 | 12.387276 | 1 |
| ACGTC | 345300 | 1.0803204 | 12.2631445 | 15 |
| CAGTC | 340980 | 1.0668046 | 12.00393 | 27 |
| AGTCA | 473955 | 0.99817234 | 8.312861 | 28 |
| AATCT | 718405 | 0.9771181 | 5.5956855 | 39 |
| TTGTA | 649375 | 0.9053115 | 5.665728 | 35 |
| ACCTT | 431865 | 0.8725962 | 7.8115163 | 32 |
| TCGTA | 404470 | 0.8447253 | 8.086403 | 43 |
| TGTAA | 551400 | 0.77519035 | 5.5556626 | 36 |
| TATGC | 357815 | 0.7472877 | 7.9495163 | 46 |
| GTATG | 338065 | 0.72978234 | 8.188311 | 45 |
| CGTAT | 343175 | 0.7167123 | 7.9286346 | 44 |
| GTAAT | 468690 | 0.6589118 | 5.47432 | 37 |
| TAATC | 463775 | 0.6307904 | 5.300672 | 38 |