![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW240_GGCTAC_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 4925928 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 38 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
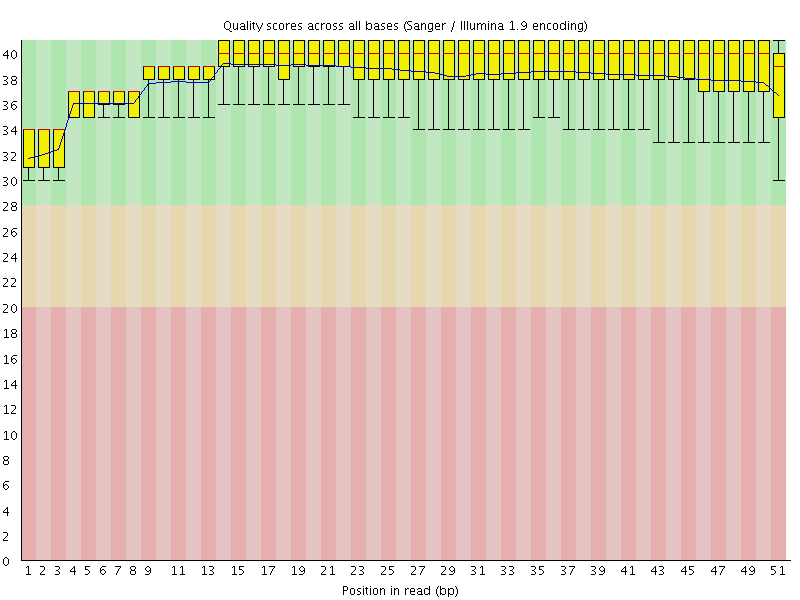
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
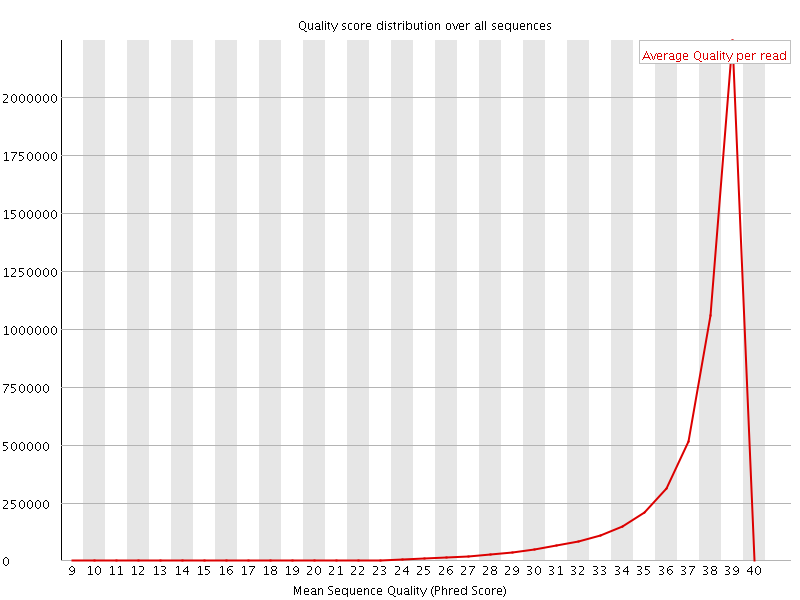
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
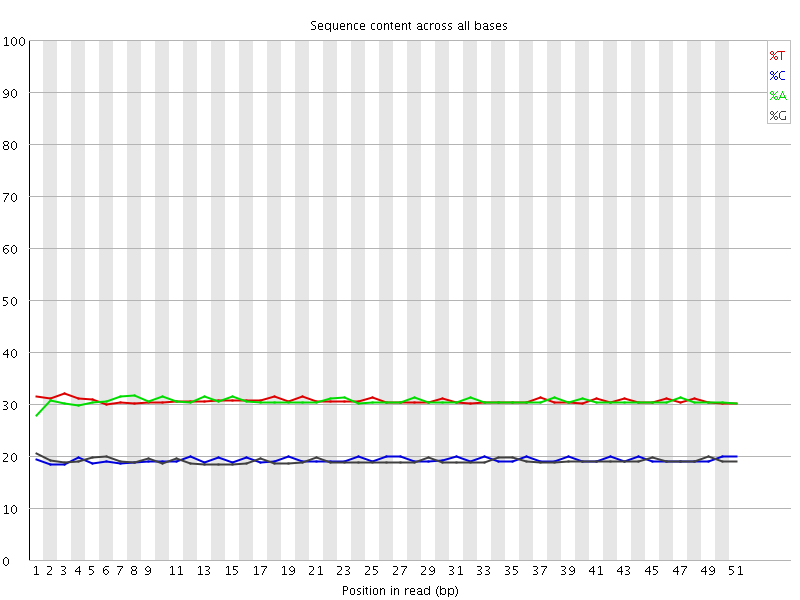
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
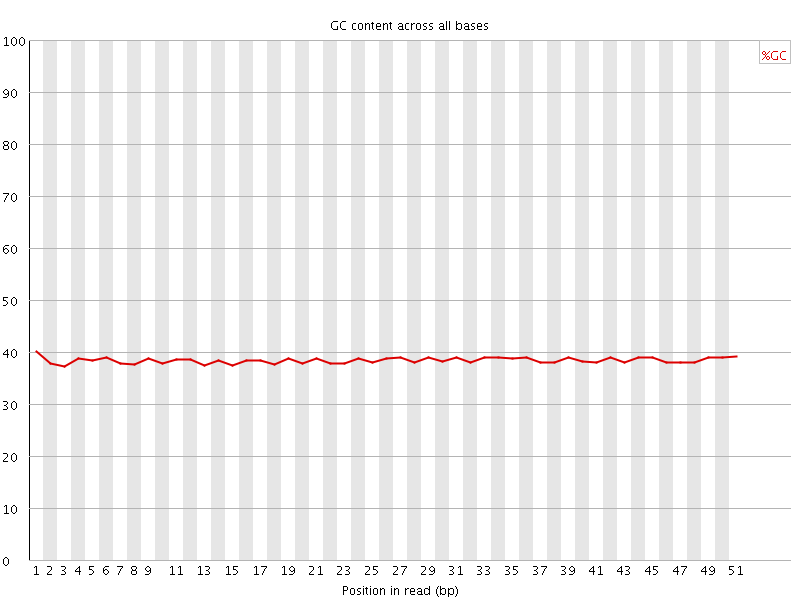
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
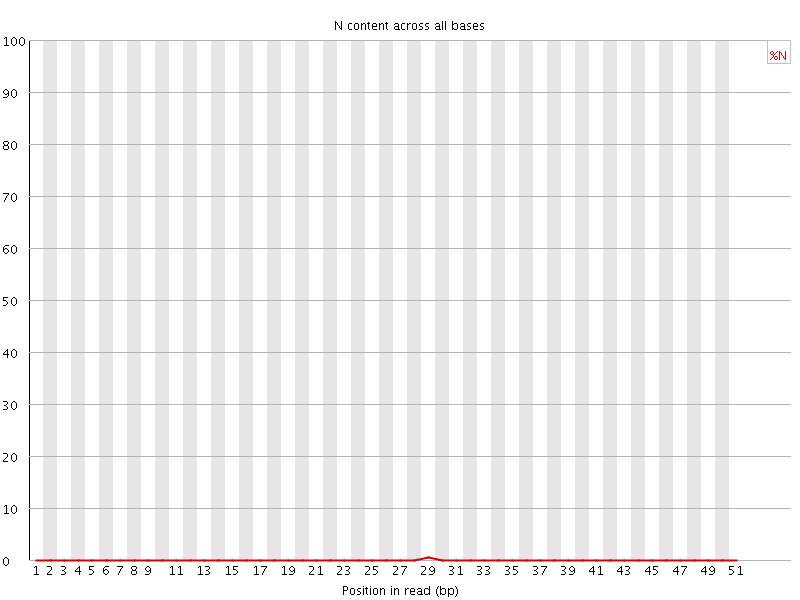
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
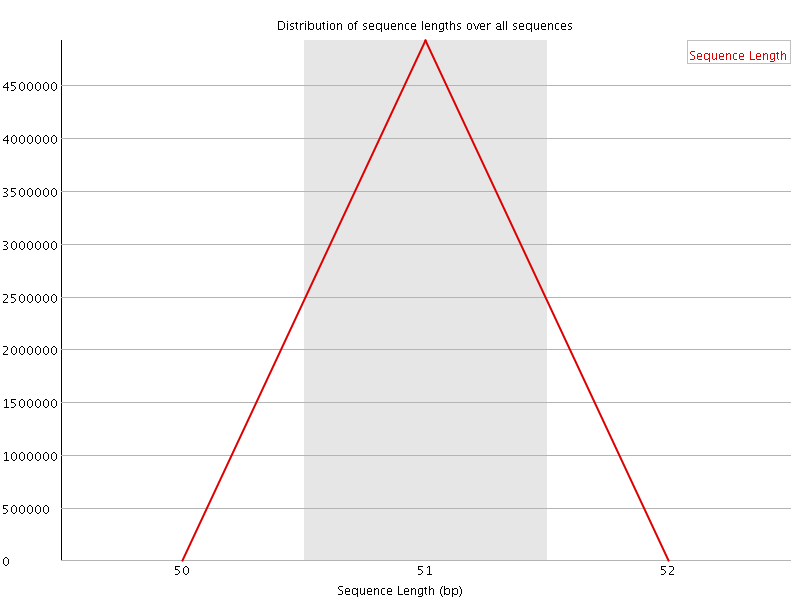
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
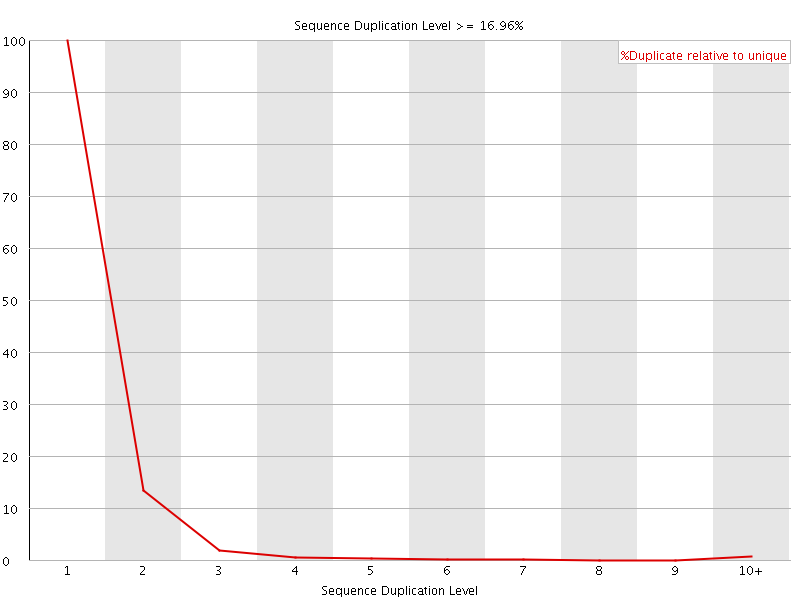
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGGCTACATCTCGTATGCC | 38320 | 0.7779244844829238 | TruSeq Adapter, Index 11 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
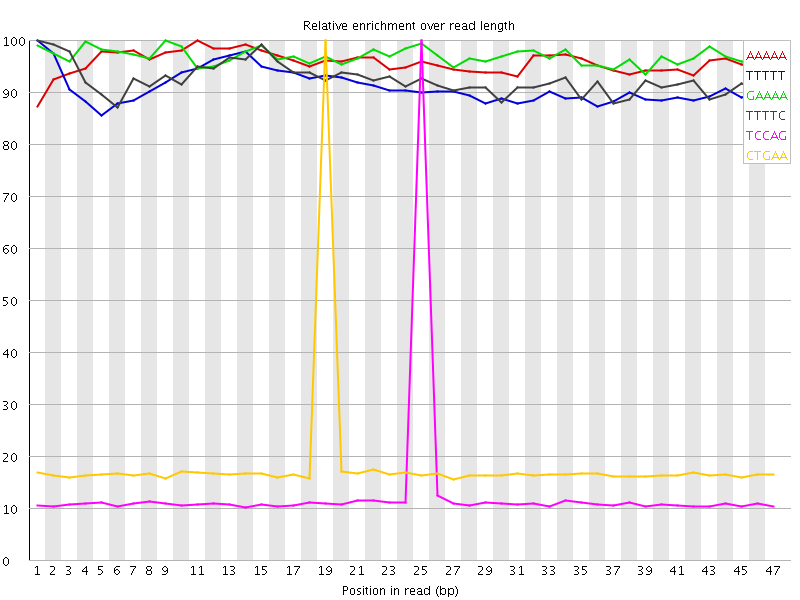
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 2383190 | 3.7987845 | 3.9665384 | 11 |
| TTTTT | 2395505 | 3.7648962 | 4.1347685 | 1 |
| GAAAA | 1194575 | 3.0409703 | 3.1347685 | 9 |
| TTTTC | 1214090 | 3.0283144 | 3.2769675 | 1 |
| TCCAG | 304260 | 1.9344337 | 15.059863 | 25 |
| CTGAA | 470895 | 1.8917553 | 10.303328 | 19 |
| GGAAG | 290705 | 1.8874564 | 15.575841 | 5 |
| CTCCA | 293395 | 1.8484895 | 14.739805 | 24 |
| GAGCA | 281090 | 1.8085276 | 15.296036 | 9 |
| CGGAA | 280780 | 1.8065331 | 15.385144 | 4 |
| GAAGA | 442675 | 1.7996855 | 10.241713 | 6 |
| ACGGC | 151685 | 1.544509 | 21.902369 | 32 |
| CGGCT | 150615 | 1.5292907 | 21.799501 | 33 |
| TCTGA | 365255 | 1.4632248 | 9.743522 | 18 |
| TCTCG | 225575 | 1.4301249 | 14.093766 | 41 |
| GCACA | 221600 | 1.4128785 | 14.774506 | 11 |
| CACAC | 218740 | 1.3820333 | 14.596414 | 12 |
| CCAGT | 216560 | 1.376852 | 14.42899 | 26 |
| CACGG | 135180 | 1.3764495 | 21.88615 | 31 |
| TCGGA | 214225 | 1.3744339 | 14.980257 | 3 |
| AGAGC | 211805 | 1.3627491 | 14.830779 | 8 |
| AGCAC | 212745 | 1.3564208 | 14.743071 | 10 |
| ATCGG | 200060 | 1.2835535 | 14.602174 | 2 |
| TGAAC | 317520 | 1.2755927 | 9.59655 | 20 |
| AAGAG | 303525 | 1.2339741 | 9.7616005 | 7 |
| GTCTG | 191750 | 1.2267697 | 14.581246 | 17 |
| ATGCC | 191585 | 1.2180651 | 13.7540865 | 47 |
| GTCAC | 190870 | 1.2135192 | 14.125583 | 29 |
| ACTCC | 184385 | 1.161689 | 14.179068 | 23 |
| AACTC | 285935 | 1.1383178 | 9.396582 | 22 |
| GAACT | 281040 | 1.1290392 | 9.457878 | 21 |
| CGTCT | 177855 | 1.1275845 | 14.32822 | 16 |
| CACGT | 176070 | 1.1194234 | 14.397 | 14 |
| CTCGT | 174685 | 1.1074868 | 13.688607 | 42 |
| CATCT | 278180 | 1.1043229 | 8.915043 | 39 |
| TCACG | 172340 | 1.0957085 | 13.984537 | 30 |
| GATCG | 169200 | 1.0855606 | 14.279407 | 1 |
| ATCTC | 270415 | 1.0734973 | 8.99228 | 40 |
| CAGTC | 166685 | 1.0597552 | 14.050466 | 27 |
| ACACG | 166175 | 1.0594994 | 14.317947 | 13 |
| CTACA | 256445 | 1.0209169 | 8.836674 | 36 |
| ACGTC | 159590 | 1.0146464 | 14.31499 | 15 |
| ACATC | 247005 | 0.9833361 | 8.750539 | 38 |
| AGTCA | 240045 | 0.96434754 | 9.15341 | 28 |
| GCTAC | 142685 | 0.90716714 | 13.386336 | 35 |
| GGCTA | 126660 | 0.81263053 | 13.549878 | 34 |
| TATGC | 186270 | 0.7462044 | 8.6291895 | 46 |
| TCGTA | 176220 | 0.70594364 | 8.601538 | 43 |
| GTATG | 174060 | 0.7036531 | 8.672681 | 45 |
| TACAT | 274095 | 0.6875482 | 5.6124277 | 37 |
| CGTAT | 160980 | 0.6448917 | 8.532929 | 44 |