![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW238_GATCAG_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 5522722 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
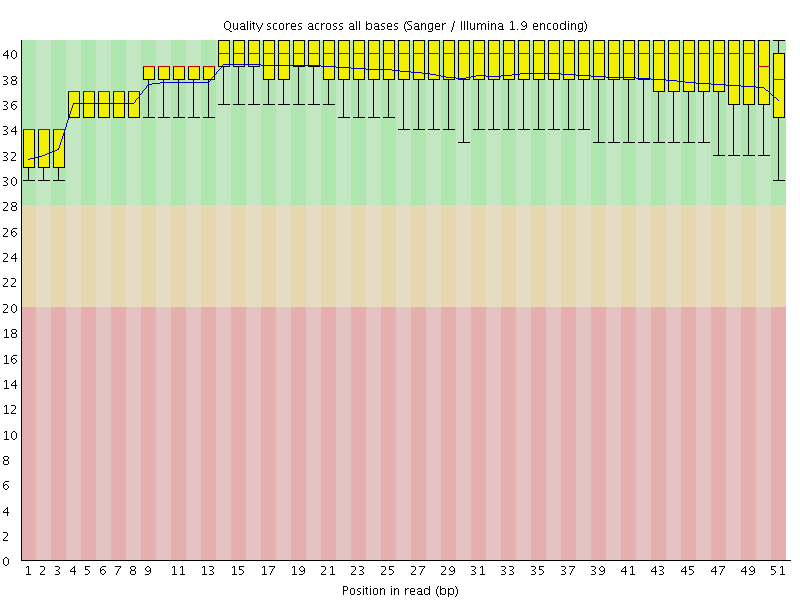
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
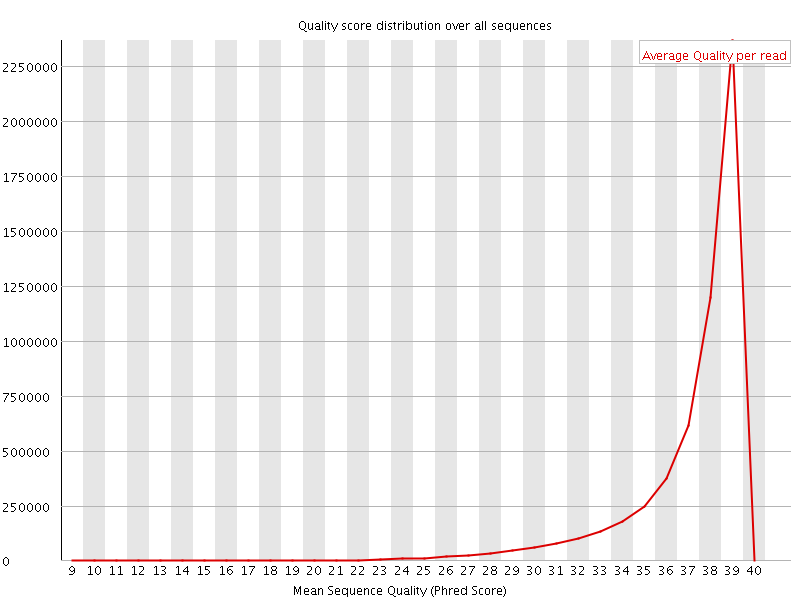
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
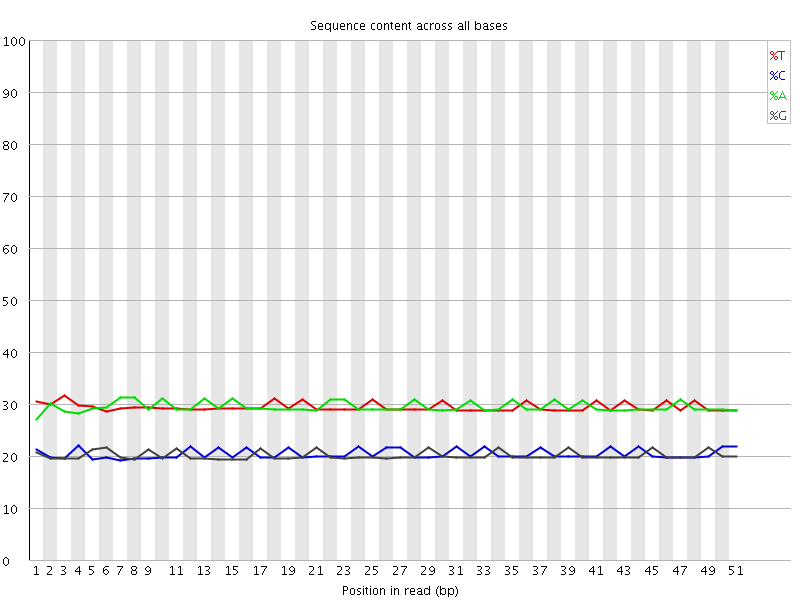
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
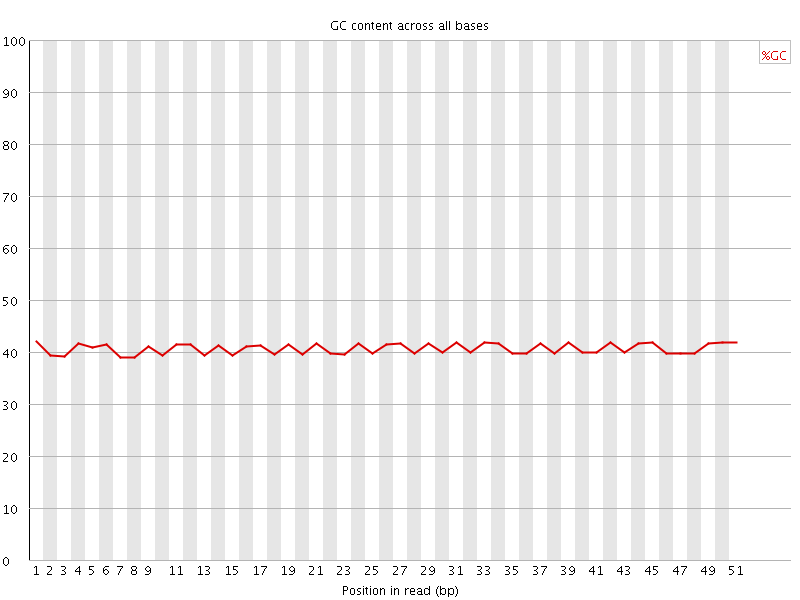
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
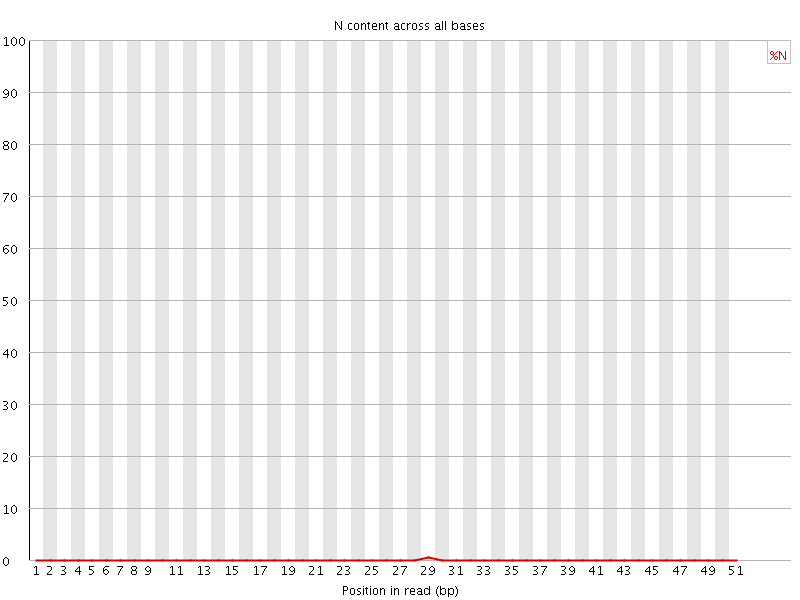
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
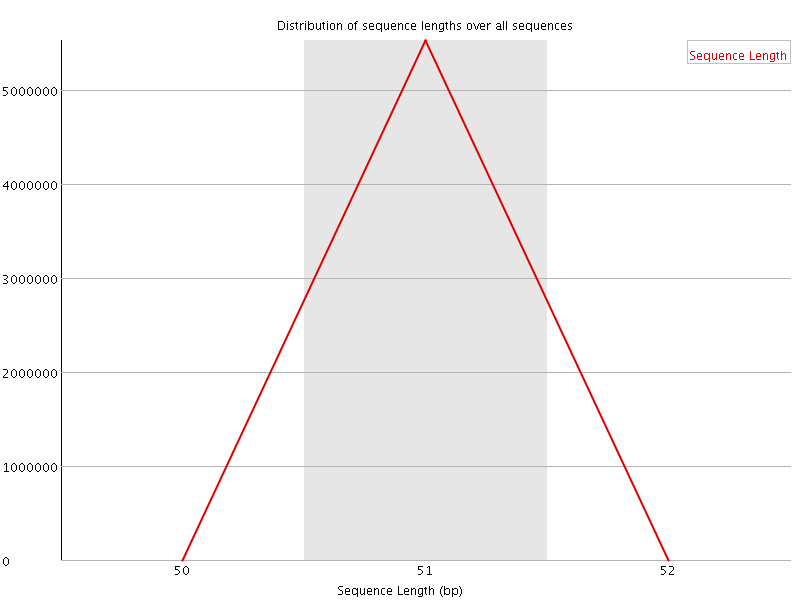
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
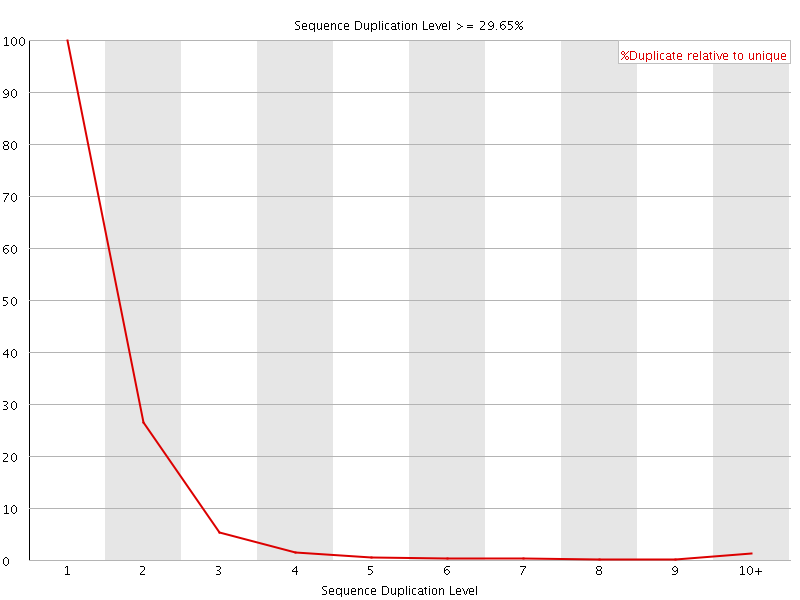
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGATCAGATCTCGTATGCC | 92120 | 1.6680180534164133 | TruSeq Adapter, Index 9 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
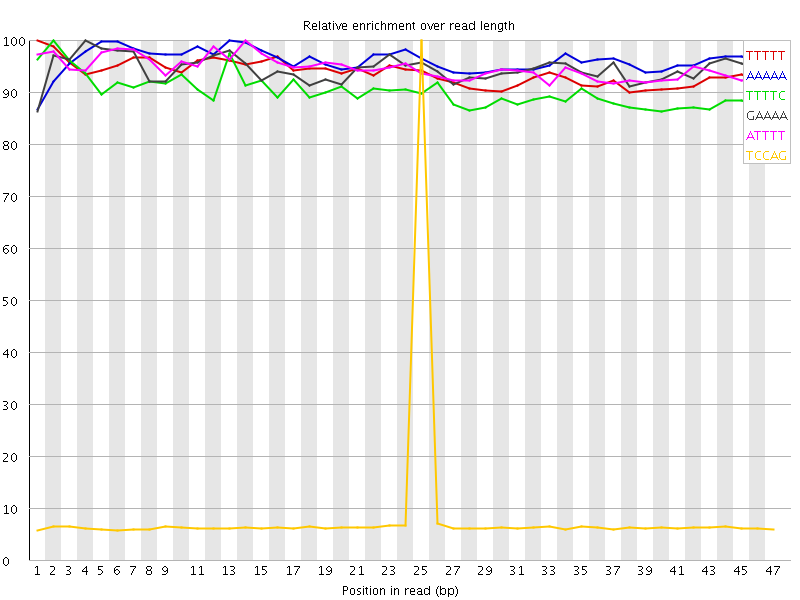
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 2434925 | 4.159459 | 4.435069 | 1 |
| AAAAA | 2428080 | 4.115898 | 4.2806735 | 13 |
| TTTTC | 1275780 | 3.1316178 | 3.472819 | 2 |
| GAAAA | 1250850 | 3.09856 | 3.2789853 | 4 |
| ATTTT | 1766430 | 3.0128522 | 3.181923 | 14 |
| TCCAG | 441130 | 2.2668066 | 27.171902 | 25 |
| GGAAG | 382260 | 2.0221803 | 28.320024 | 5 |
| CTGAA | 563665 | 2.0125992 | 19.568947 | 19 |
| CGGAA | 379505 | 1.9771382 | 27.930597 | 4 |
| GAAGA | 525965 | 1.90399 | 19.77291 | 6 |
| GAGCA | 362465 | 1.8883631 | 27.61502 | 9 |
| CTCCA | 365585 | 1.8500978 | 26.440855 | 24 |
| TCTCG | 328740 | 1.6918833 | 26.384567 | 41 |
| CCAGT | 312200 | 1.6042823 | 26.456787 | 26 |
| ATCGG | 302980 | 1.5808964 | 26.977785 | 2 |
| TCTGA | 439635 | 1.5721663 | 19.20881 | 18 |
| TCAGA | 437825 | 1.5632801 | 18.422213 | 36 |
| GCACA | 301335 | 1.5460645 | 26.894785 | 11 |
| AGCAC | 299010 | 1.5341353 | 26.899601 | 10 |
| GATCG | 293445 | 1.5311445 | 26.771372 | 1 |
| AGAGC | 290975 | 1.515916 | 27.260307 | 8 |
| TCGGA | 289930 | 1.5128038 | 27.492273 | 3 |
| ATGCC | 293490 | 1.5081385 | 26.129807 | 47 |
| CACAC | 296010 | 1.4956945 | 26.392902 | 12 |
| CGATC | 291065 | 1.4956774 | 25.783178 | 33 |
| GTCTG | 282795 | 1.4778526 | 27.148018 | 17 |
| TGAAC | 402095 | 1.4357042 | 19.007 | 20 |
| GTCAC | 275395 | 1.4151549 | 26.30267 | 29 |
| CACGA | 272300 | 1.397094 | 25.90282 | 31 |
| TCACG | 267355 | 1.3738401 | 26.19323 | 30 |
| ATCAG | 384645 | 1.373398 | 18.169521 | 35 |
| CGTCT | 264660 | 1.3620913 | 26.705053 | 16 |
| AAGAG | 374935 | 1.3572625 | 19.217926 | 7 |
| GAACT | 372510 | 1.3300691 | 18.84099 | 21 |
| CAGAT | 364825 | 1.3026294 | 18.113504 | 37 |
| GATCA | 364525 | 1.3015581 | 18.16151 | 34 |
| CACGT | 253200 | 1.3011029 | 26.6178 | 14 |
| CTCGT | 252590 | 1.2999719 | 25.821653 | 42 |
| ACTCC | 254150 | 1.2861645 | 25.998783 | 23 |
| ACGTC | 249585 | 1.2825266 | 26.609596 | 15 |
| CAGTC | 246405 | 1.2661858 | 26.12655 | 27 |
| AACTC | 355585 | 1.2503691 | 18.463417 | 22 |
| ATCTC | 354335 | 1.2478971 | 18.109608 | 40 |
| ACACG | 240090 | 1.2318336 | 26.486427 | 13 |
| AGATC | 321905 | 1.1493809 | 18.154306 | 38 |
| GATCT | 317815 | 1.1365292 | 18.182467 | 39 |
| AGTCA | 313045 | 1.1177458 | 18.33194 | 28 |
| ACGAT | 307770 | 1.098911 | 18.119574 | 32 |
| TATGC | 262685 | 0.93938047 | 18.013083 | 46 |
| GTATG | 244480 | 0.8877508 | 18.255466 | 45 |
| TCGTA | 244640 | 0.8748502 | 18.011305 | 43 |
| CGTAT | 235050 | 0.8405557 | 17.97974 | 44 |