![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW237_ACTTGA_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 5155376 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
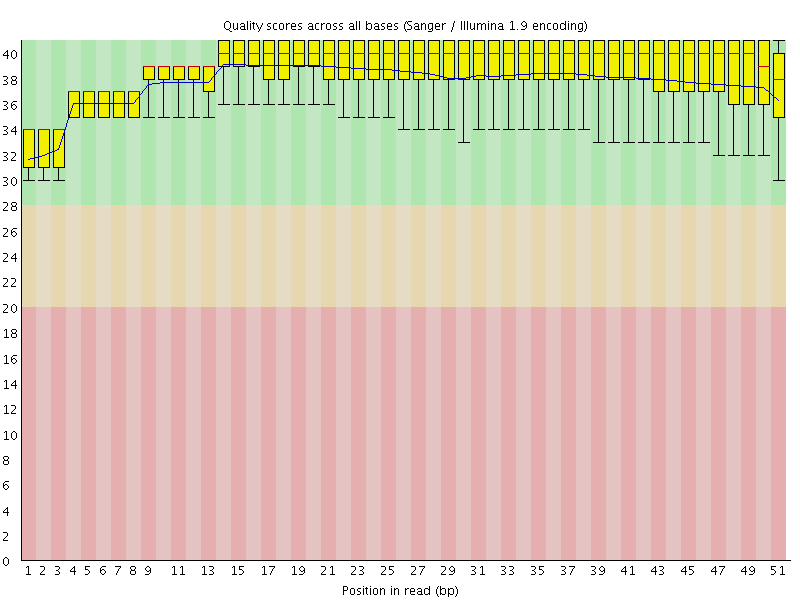
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
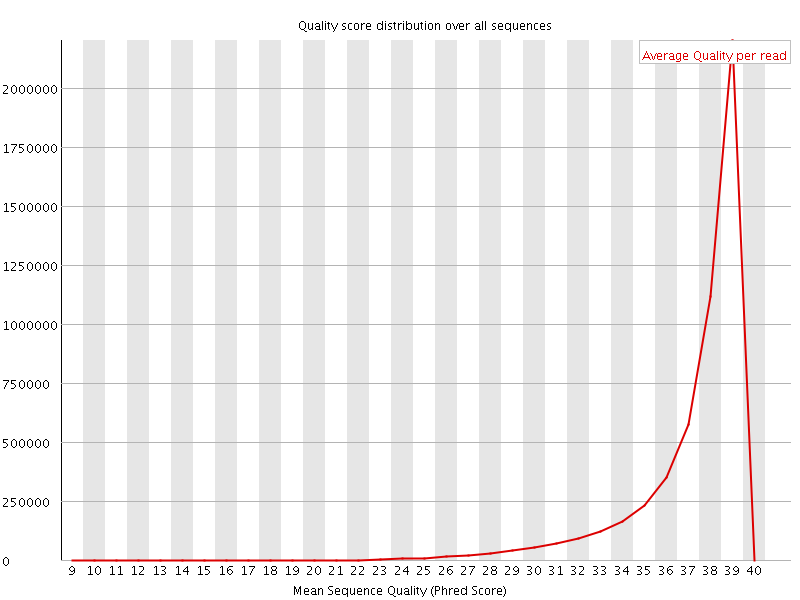
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
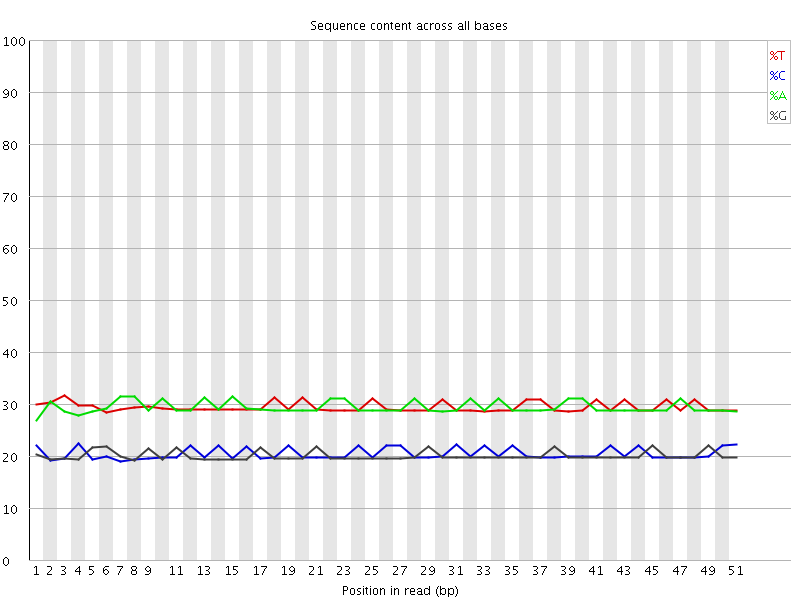
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
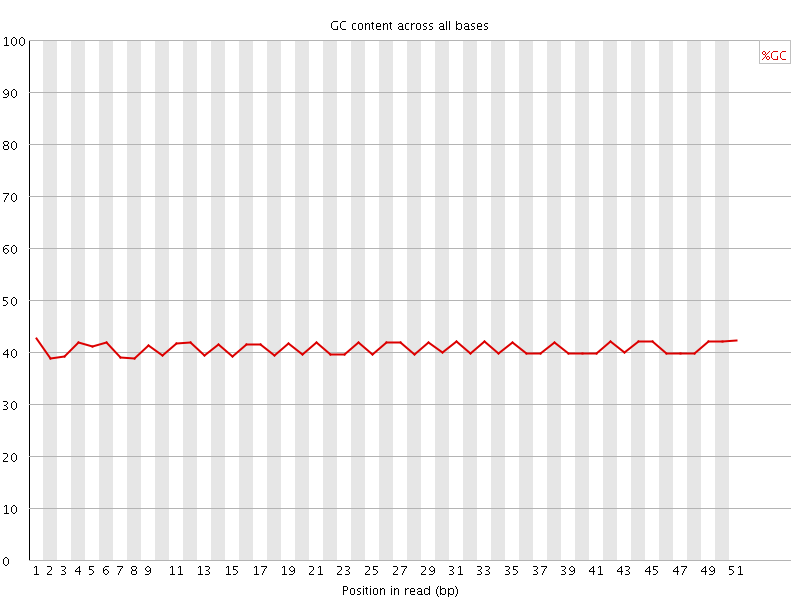
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
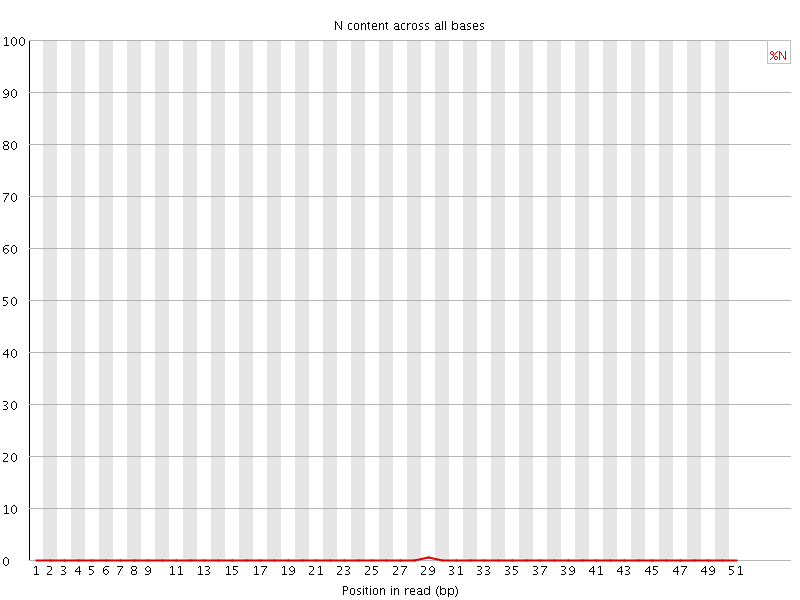
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
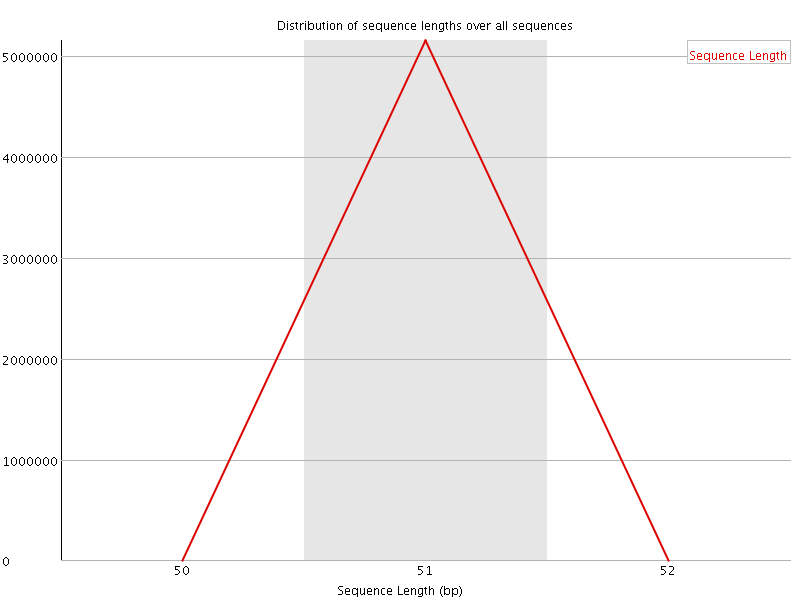
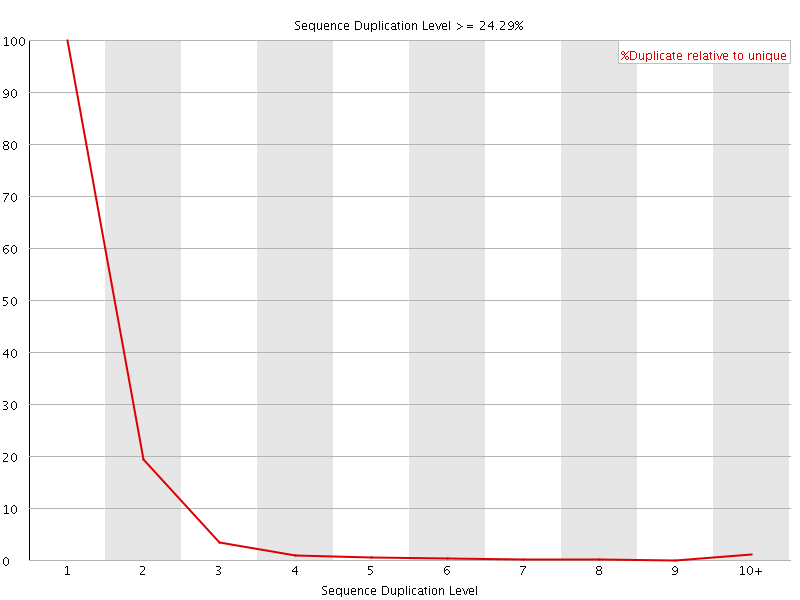
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACACTTGAATCTCGTATGCC | 102566 | 1.989496013481849 | TruSeq Adapter, Index 8 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
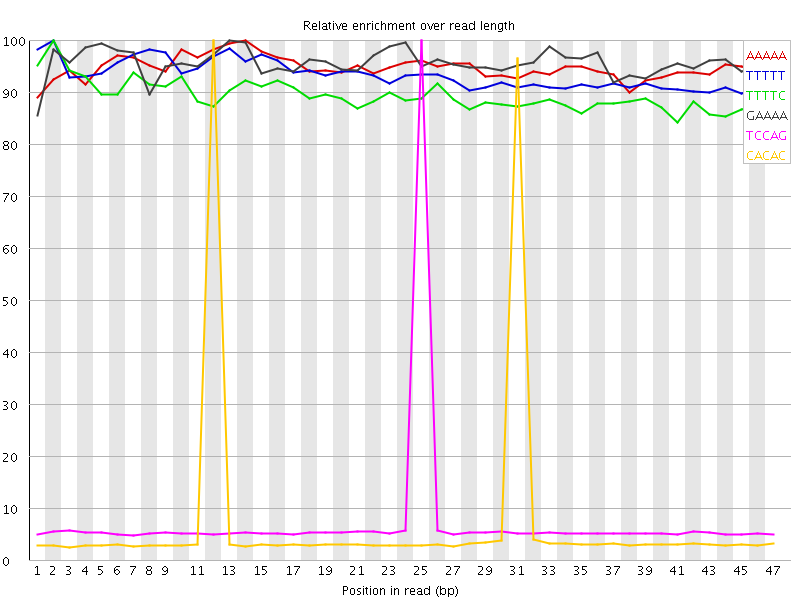
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 2234230 | 4.0900135 | 4.313735 | 14 |
| TTTTT | 2238060 | 4.0882597 | 4.3787003 | 2 |
| TTTTC | 1170410 | 3.0635164 | 3.4271693 | 2 |
| GAAAA | 1145200 | 3.063002 | 3.199358 | 13 |
| TCCAG | 422185 | 2.3154826 | 31.351715 | 25 |
| CACAC | 410125 | 2.2059786 | 30.614288 | 12 |
| TTGAA | 801460 | 2.141784 | 16.006021 | 36 |
| GGAAG | 370265 | 2.1140566 | 32.909847 | 5 |
| CGGAA | 368545 | 2.0627894 | 32.43377 | 4 |
| CTGAA | 537125 | 2.056765 | 22.584204 | 19 |
| GAAGA | 510520 | 1.9950219 | 23.056715 | 6 |
| GAGCA | 354670 | 1.9851294 | 32.13942 | 9 |
| CTCCA | 357425 | 1.9216931 | 30.512817 | 24 |
| TCTCG | 325555 | 1.7847489 | 30.490337 | 41 |
| CCAGT | 308820 | 1.6937298 | 30.693424 | 26 |
| ATCGG | 299080 | 1.6732687 | 31.43869 | 2 |
| GATCG | 291760 | 1.6323154 | 31.145061 | 1 |
| TCTGA | 426055 | 1.6307559 | 22.211624 | 18 |
| GCACA | 296495 | 1.6268296 | 31.323448 | 11 |
| AGCAC | 296130 | 1.624827 | 31.26677 | 10 |
| TCGGA | 288130 | 1.6120067 | 31.864271 | 3 |
| AGAGC | 287685 | 1.6102065 | 31.799076 | 8 |
| ATGCC | 288150 | 1.5803647 | 30.115099 | 47 |
| GTCTG | 281380 | 1.5735682 | 31.606916 | 17 |
| TGAAC | 394935 | 1.5122895 | 22.075216 | 20 |
| GTCAC | 274665 | 1.5064061 | 30.35533 | 29 |
| CGTCT | 268360 | 1.4711958 | 30.887577 | 16 |
| TGAAT | 543395 | 1.4521434 | 15.304957 | 37 |
| AAGAG | 369975 | 1.4457967 | 22.516327 | 7 |
| CACGT | 256545 | 1.4070264 | 30.957226 | 14 |
| CTCGT | 254790 | 1.3968028 | 29.93394 | 42 |
| GAACT | 362940 | 1.389774 | 21.936745 | 21 |
| CACTT | 370095 | 1.388663 | 21.018858 | 33 |
| ACGTC | 252515 | 1.3849239 | 30.921415 | 15 |
| TCACA | 368195 | 1.3821259 | 21.019173 | 30 |
| CTTGA | 359250 | 1.375055 | 21.282894 | 35 |
| ACTCC | 253810 | 1.3646078 | 30.055489 | 23 |
| CAGTC | 247110 | 1.3552802 | 30.31504 | 27 |
| ACACG | 242880 | 1.3326511 | 30.91896 | 13 |
| ATCTC | 350240 | 1.3141634 | 20.97548 | 40 |
| AACTC | 346165 | 1.2994299 | 21.40717 | 22 |
| GAATC | 334545 | 1.2810435 | 21.114752 | 38 |
| ACTTG | 325750 | 1.2468313 | 21.2621 | 34 |
| AGTCA | 310950 | 1.1906933 | 21.417953 | 28 |
| AATCT | 437930 | 1.1472523 | 14.774854 | 39 |
| ACACT | 292635 | 1.0984896 | 20.737953 | 32 |
| TATGC | 261100 | 0.99937886 | 20.848316 | 46 |
| GTATG | 247125 | 0.96489394 | 21.198875 | 45 |
| TCGTA | 251150 | 0.96129453 | 20.889746 | 43 |
| CGTAT | 238145 | 0.91151696 | 20.832184 | 44 |