![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW236_CAGATC_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 7827595 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 37 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
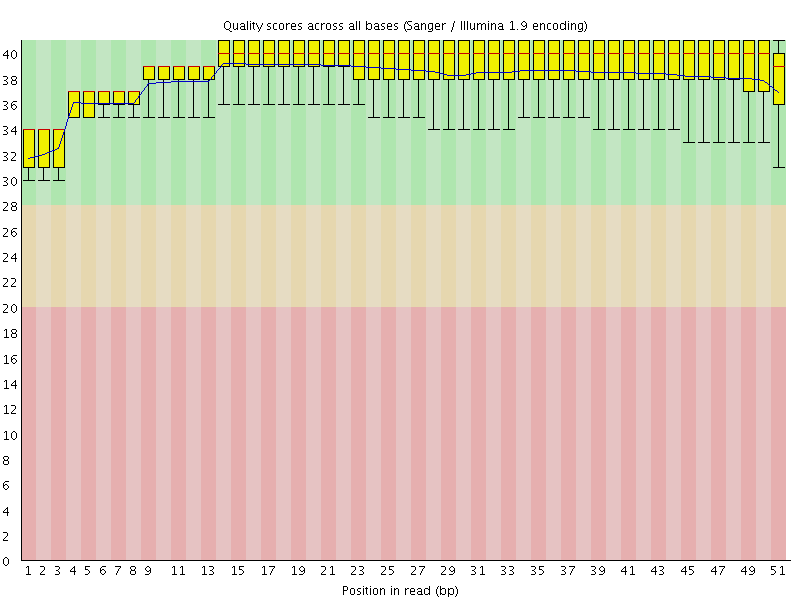
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
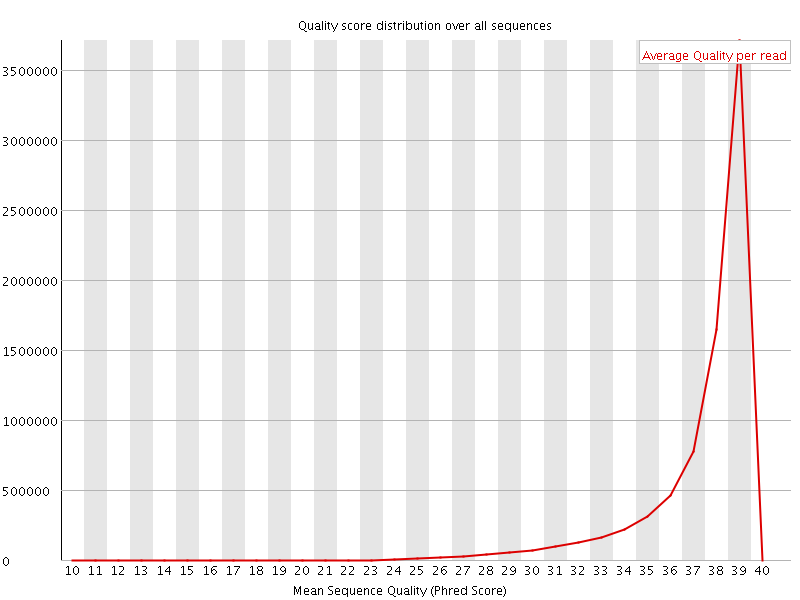
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
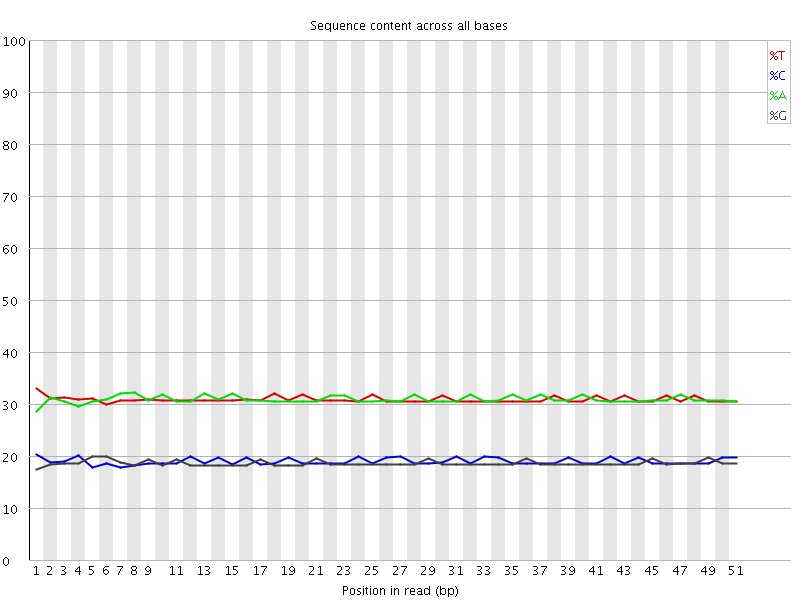
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
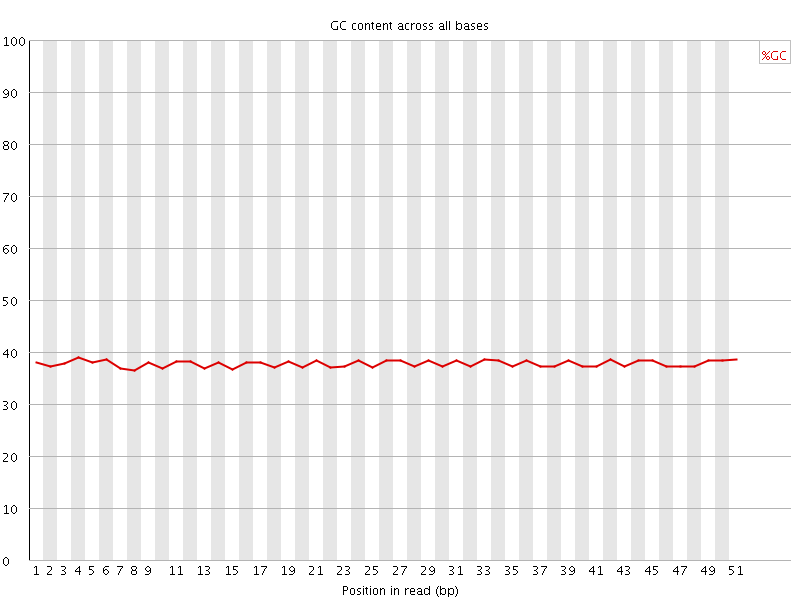
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
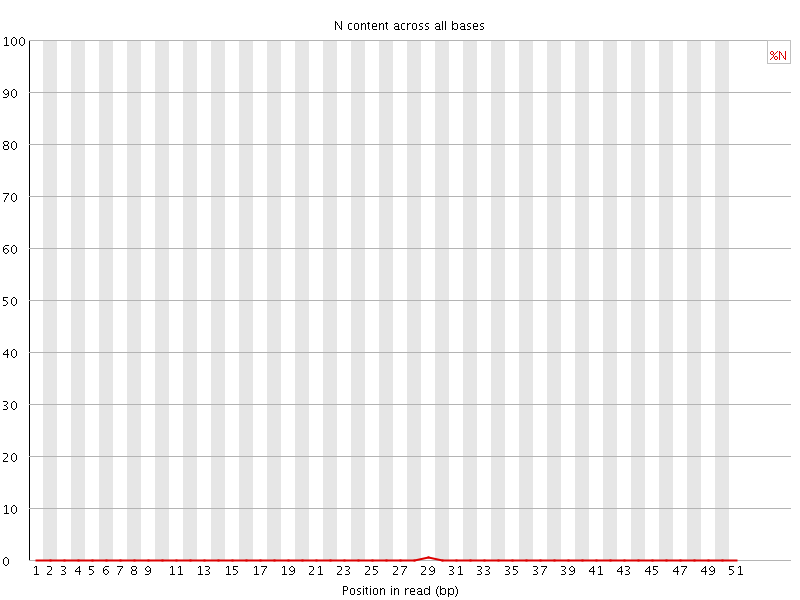
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
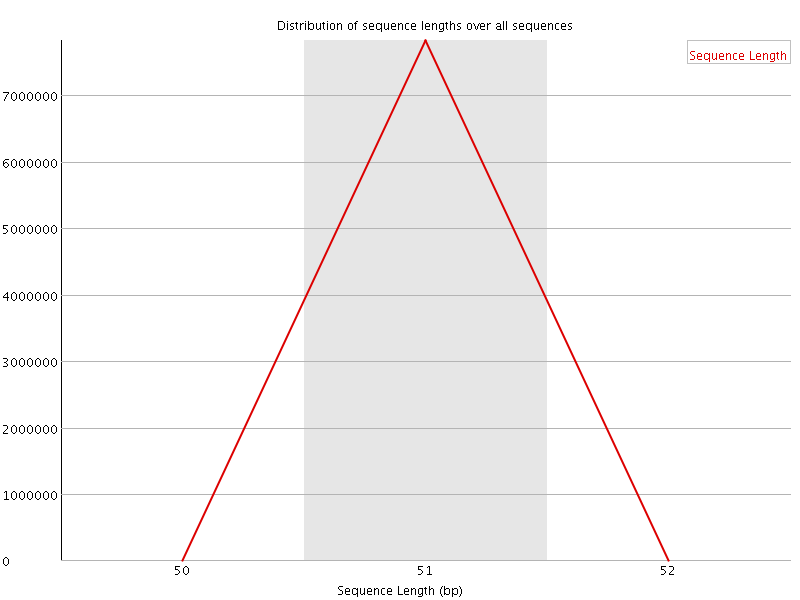
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCAGATCATCTCGTATGCC | 79796 | 1.0194191191547342 | TruSeq Adapter, Index 7 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
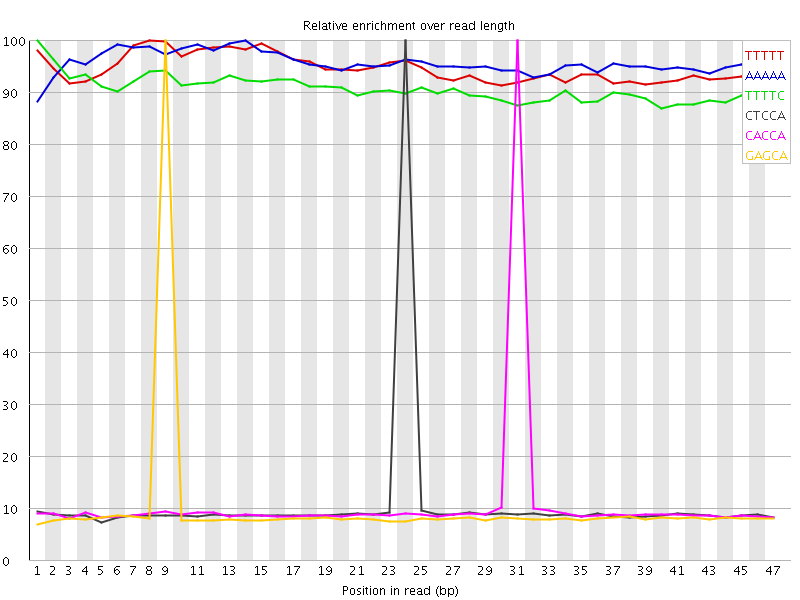
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 3732395 | 3.5301256 | 3.7293928 | 8 |
| AAAAA | 3740550 | 3.5296223 | 3.6846182 | 14 |
| TTTTC | 1964510 | 3.009996 | 3.3194056 | 1 |
| CTCCA | 516130 | 2.0743742 | 19.277742 | 24 |
| CACCA | 515445 | 2.070658 | 19.102032 | 31 |
| GAGCA | 489185 | 2.046052 | 20.387703 | 9 |
| GGAAG | 475395 | 2.0288823 | 20.931301 | 5 |
| CGGAA | 483065 | 2.0204546 | 20.650858 | 4 |
| GAAGA | 781625 | 2.0171156 | 13.395442 | 6 |
| CTGAA | 791200 | 2.00199 | 13.053639 | 19 |
| TCCAG | 479250 | 1.9653906 | 19.581068 | 25 |
| CCAGA | 451605 | 1.8511579 | 19.076925 | 33 |
| TCACC | 419980 | 1.6879385 | 18.657377 | 30 |
| TCTCG | 411235 | 1.6872475 | 19.146738 | 41 |
| TCGGA | 401765 | 1.681193 | 20.24686 | 3 |
| TCTGA | 627665 | 1.5889328 | 12.664655 | 18 |
| GCACA | 377295 | 1.5465564 | 19.506186 | 11 |
| AGAGC | 365770 | 1.5298595 | 19.855473 | 8 |
| CACAC | 376465 | 1.5123439 | 19.064856 | 12 |
| TCATC | 595290 | 1.4768876 | 11.840654 | 38 |
| AGCAC | 355950 | 1.4590621 | 19.427273 | 10 |
| ATCGG | 340140 | 1.4233221 | 19.679028 | 2 |
| CCAGT | 346895 | 1.4226062 | 18.939634 | 26 |
| TGAAC | 554170 | 1.402228 | 12.484772 | 20 |
| GTCAC | 331850 | 1.3609073 | 18.81278 | 29 |
| AAGAG | 524255 | 1.352929 | 12.669598 | 7 |
| CTCGT | 328925 | 1.3495395 | 18.678255 | 42 |
| ACCAG | 326900 | 1.339984 | 18.636148 | 32 |
| CACGT | 321680 | 1.3192003 | 19.224487 | 14 |
| GTCTG | 312690 | 1.3090656 | 19.549387 | 17 |
| CGTCT | 317835 | 1.3040384 | 19.11601 | 16 |
| ACTCC | 321310 | 1.2913744 | 18.655773 | 23 |
| ATGCC | 301785 | 1.2376117 | 18.467907 | 47 |
| ACACG | 301450 | 1.235663 | 19.149149 | 13 |
| AACTC | 497145 | 1.2328207 | 11.988774 | 22 |
| GATCG | 291210 | 1.2185736 | 19.203474 | 1 |
| GAACT | 477330 | 1.2077981 | 12.190076 | 21 |
| CATCT | 486500 | 1.2069844 | 11.611704 | 39 |
| ACGTC | 294125 | 1.2061981 | 19.112902 | 15 |
| CAGTC | 291830 | 1.1967864 | 18.753603 | 27 |
| GATCA | 467445 | 1.182786 | 11.799403 | 36 |
| ATCTC | 471730 | 1.1703408 | 11.721028 | 40 |
| CAGAT | 457625 | 1.1579381 | 11.772887 | 34 |
| AGTCA | 423620 | 1.0718946 | 11.902296 | 28 |
| AGATC | 415650 | 1.0517279 | 11.608994 | 35 |
| ATCAT | 619640 | 0.9485216 | 7.356214 | 37 |
| TCGTA | 328095 | 0.8305719 | 11.487225 | 43 |
| TATGC | 314955 | 0.79730797 | 11.40843 | 46 |
| GTATG | 301870 | 0.77975166 | 11.606732 | 45 |
| CGTAT | 288935 | 0.7314384 | 11.3671055 | 44 |