![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW235_GCCAAT_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 6441440 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
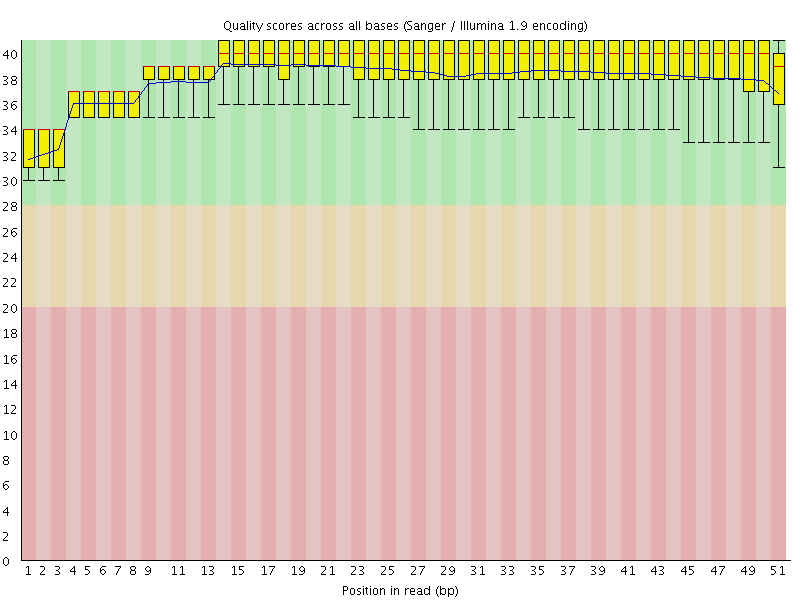
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
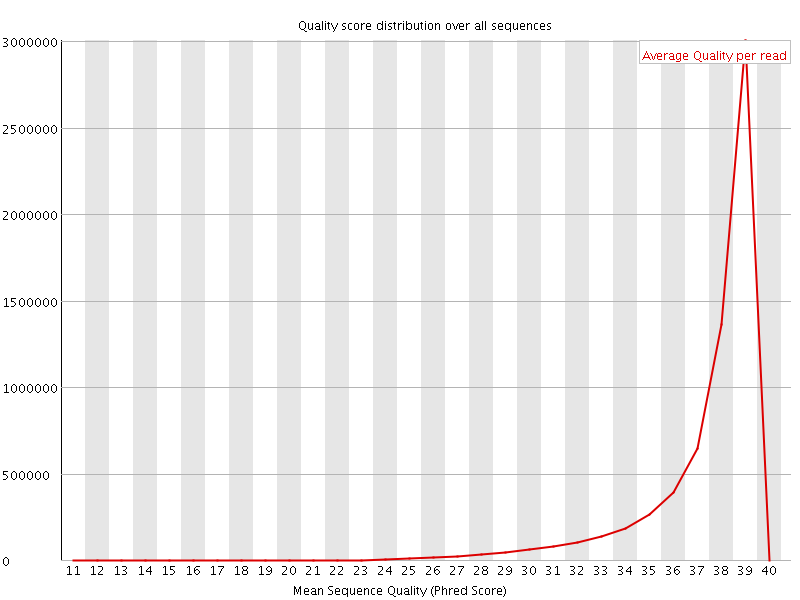
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
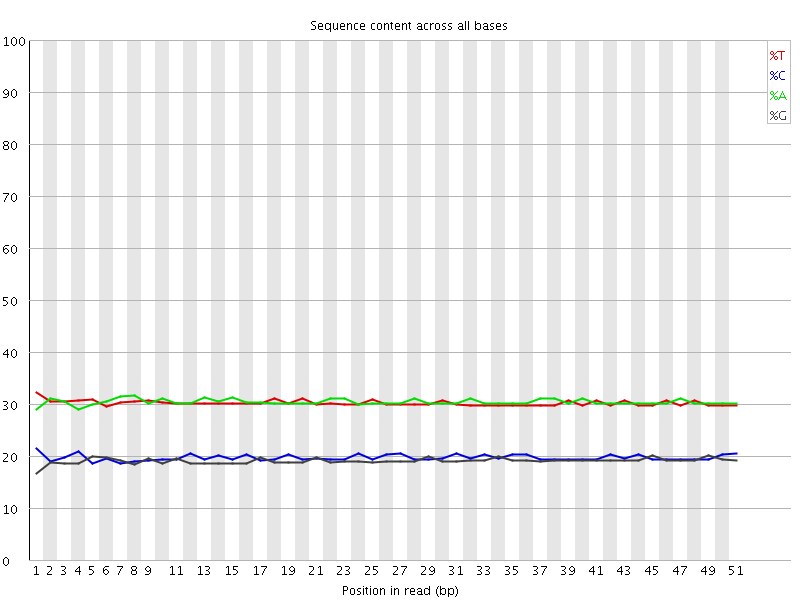
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
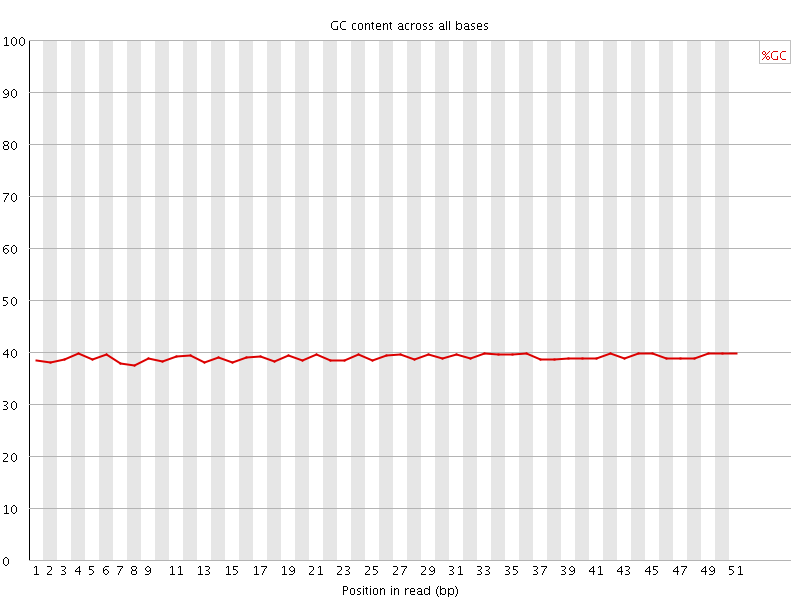
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
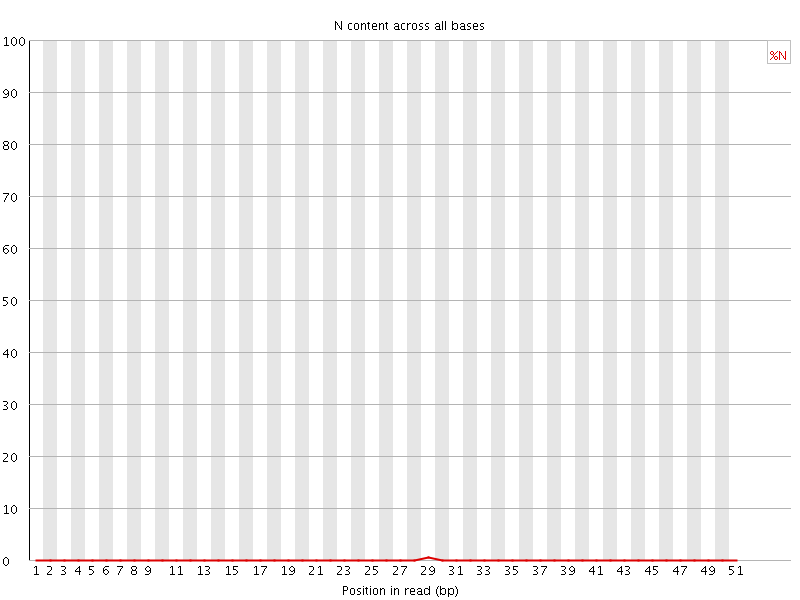
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
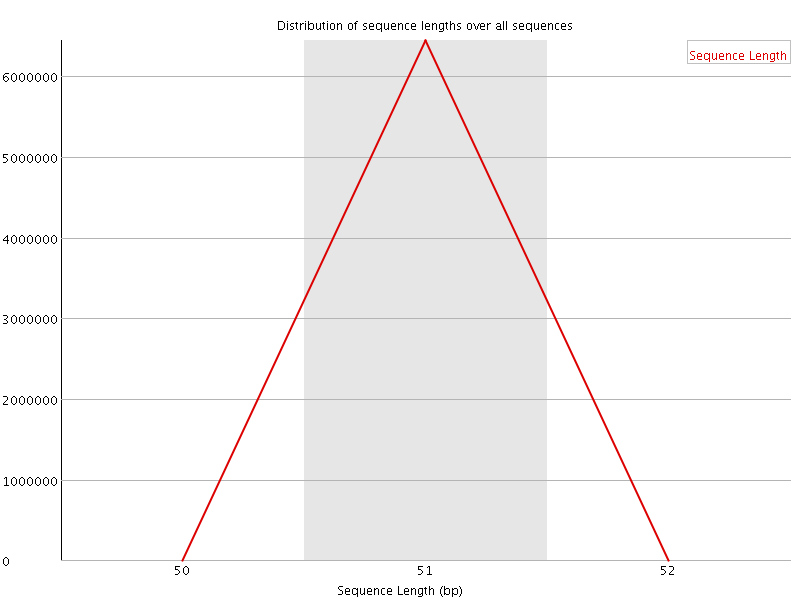
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
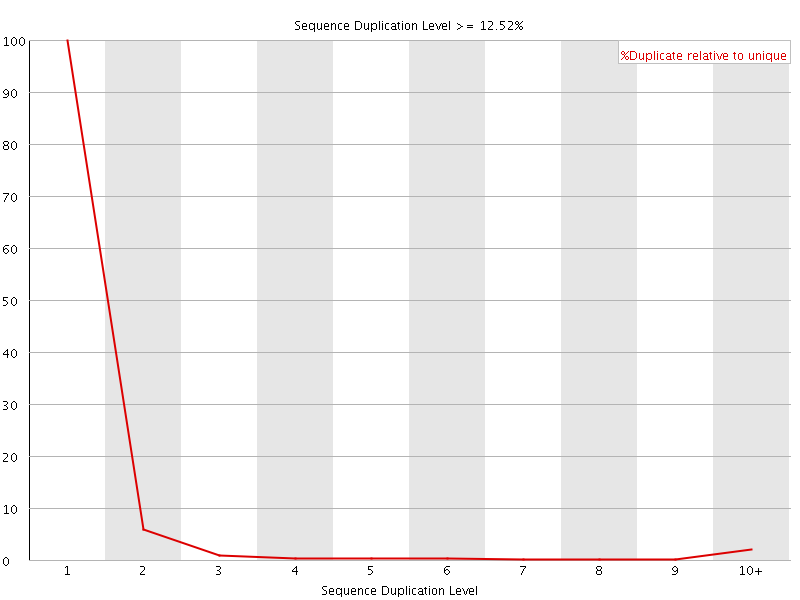
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGCCAATATCTCGTATGCC | 52881 | 0.8209499739188753 | TruSeq Adapter, Index 6 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
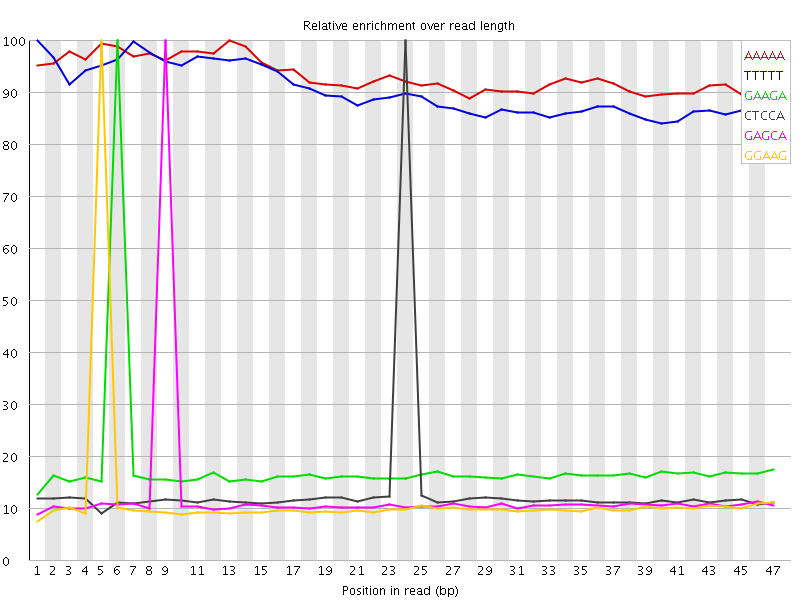
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 2564245 | 3.1877859 | 3.4192364 | 13 |
| TTTTT | 2474570 | 3.1852577 | 3.5305502 | 1 |
| GAAGA | 636555 | 1.9964632 | 11.102642 | 6 |
| CTCCA | 437110 | 1.9827775 | 14.741218 | 24 |
| GAGCA | 409280 | 1.9716179 | 15.828231 | 9 |
| GGAAG | 385200 | 1.9189315 | 16.39636 | 5 |
| CTGAA | 607970 | 1.8567761 | 10.561799 | 19 |
| TCCAG | 391850 | 1.838121 | 15.09952 | 25 |
| CGGAA | 373800 | 1.8007007 | 15.870077 | 4 |
| GCCAA | 367375 | 1.7113577 | 14.466535 | 34 |
| CGCCA | 231695 | 1.6577717 | 21.285835 | 33 |
| TCTCG | 334155 | 1.5784296 | 14.771915 | 41 |
| TCGGA | 320740 | 1.5558882 | 15.754291 | 3 |
| TCTGA | 483520 | 1.4870133 | 10.293684 | 18 |
| CACAC | 322520 | 1.4528369 | 14.424474 | 12 |
| AGCAC | 308850 | 1.4387285 | 14.923201 | 10 |
| GCACA | 307465 | 1.4322766 | 14.902421 | 11 |
| AGAGC | 292495 | 1.4090313 | 15.361129 | 8 |
| TGAAC | 449060 | 1.3714556 | 10.151749 | 20 |
| CCAAT | 450830 | 1.3314326 | 9.39691 | 35 |
| ATCGG | 271750 | 1.3182409 | 15.229272 | 2 |
| CCAGT | 280345 | 1.3150645 | 14.501037 | 26 |
| AAGAG | 419155 | 1.3146193 | 10.369976 | 7 |
| GTCAC | 277110 | 1.2998894 | 14.394661 | 29 |
| CTCGT | 271275 | 1.2814068 | 14.449513 | 42 |
| CACGT | 269880 | 1.2659745 | 14.715769 | 14 |
| CACGC | 174535 | 1.2487934 | 21.025694 | 31 |
| CGTCT | 258285 | 1.2200466 | 14.726603 | 16 |
| ACGCC | 167910 | 1.2013917 | 20.895567 | 32 |
| GTCTG | 245225 | 1.1978793 | 15.184763 | 17 |
| ACTCC | 263055 | 1.1932455 | 14.002327 | 23 |
| AACTC | 400490 | 1.1827639 | 9.565911 | 22 |
| ATGCC | 249260 | 1.1692485 | 14.261129 | 47 |
| ACACG | 249870 | 1.1639795 | 14.516354 | 13 |
| ATCTC | 386410 | 1.1491529 | 9.505202 | 40 |
| TCACG | 244365 | 1.1462867 | 14.295002 | 30 |
| GAACT | 375200 | 1.1458828 | 9.848403 | 21 |
| GATCG | 235575 | 1.1427584 | 14.808164 | 1 |
| ACGTC | 242315 | 1.1366704 | 14.578194 | 15 |
| CAGTC | 236300 | 1.1084547 | 14.298216 | 27 |
| AGTCA | 347220 | 1.0604303 | 9.583257 | 28 |
| CAATA | 470955 | 0.9055427 | 6.1696177 | 36 |
| TCGTA | 279610 | 0.8599102 | 9.404096 | 43 |
| ATATC | 419940 | 0.81309223 | 6.1843534 | 38 |
| TATGC | 252355 | 0.7760905 | 9.2747755 | 46 |
| GTATG | 240820 | 0.7658857 | 9.596387 | 45 |
| CGTAT | 240340 | 0.7391396 | 9.273057 | 44 |
| TATCT | 333935 | 0.65108484 | 6.08623 | 39 |