![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW231_CGATGT_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 14552538 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 38 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
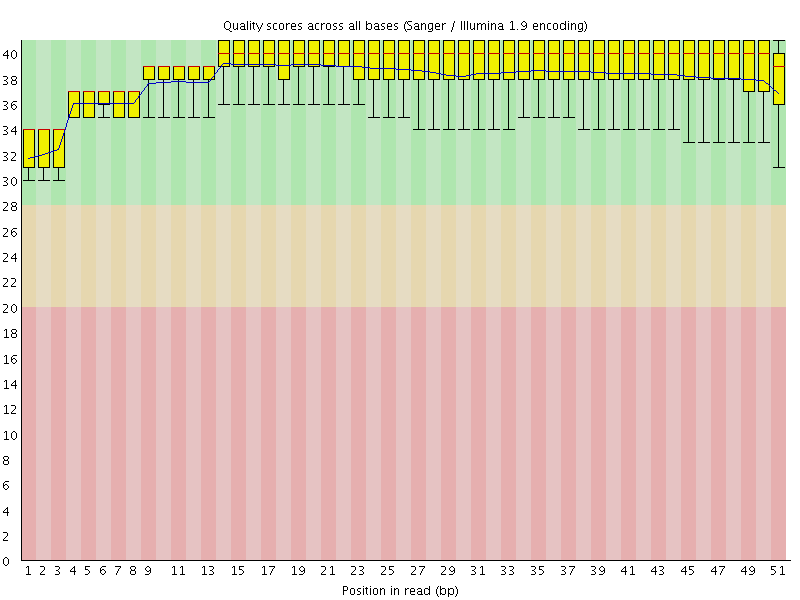
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
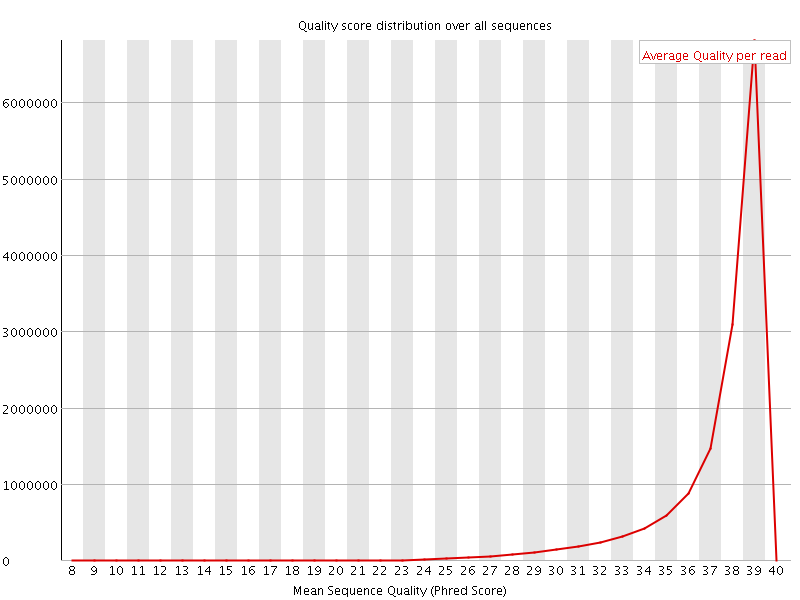
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
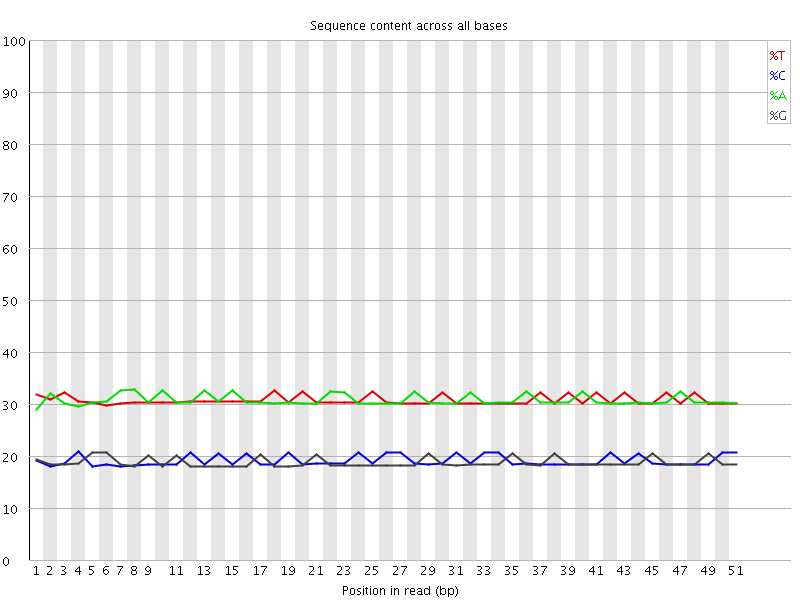
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
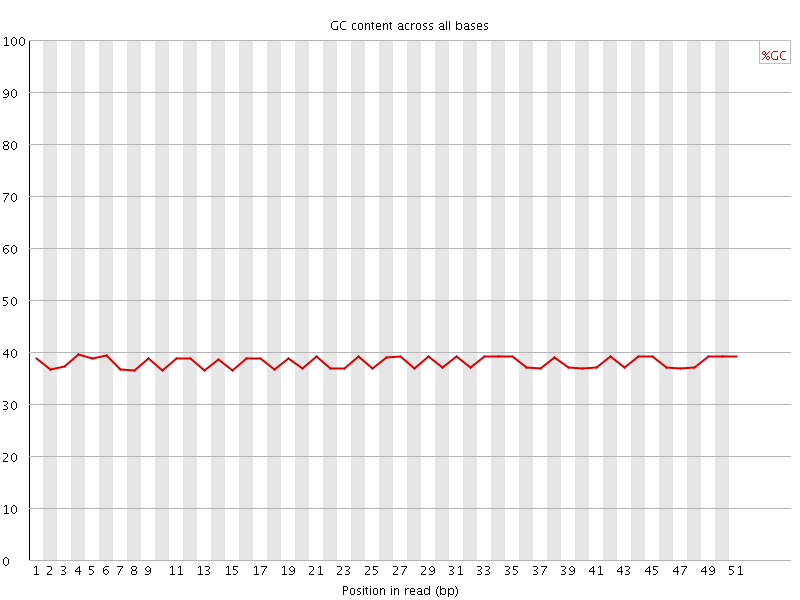
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
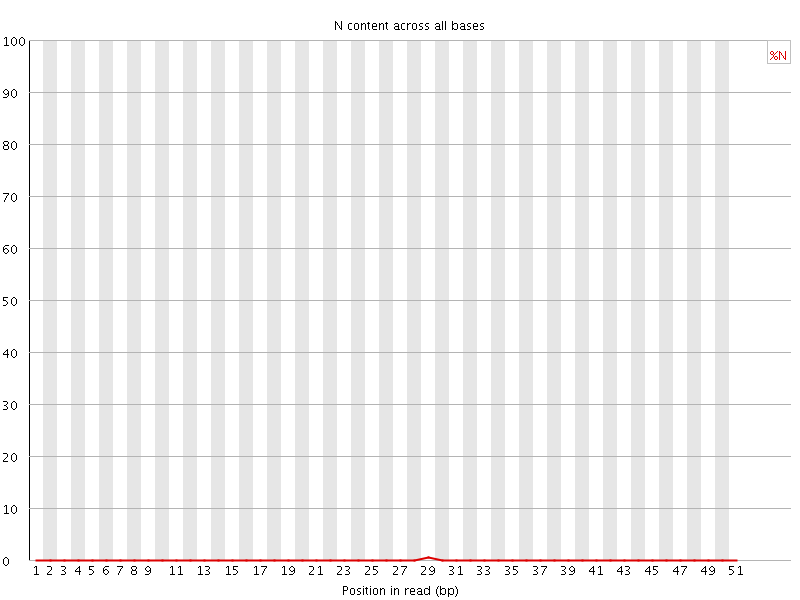
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
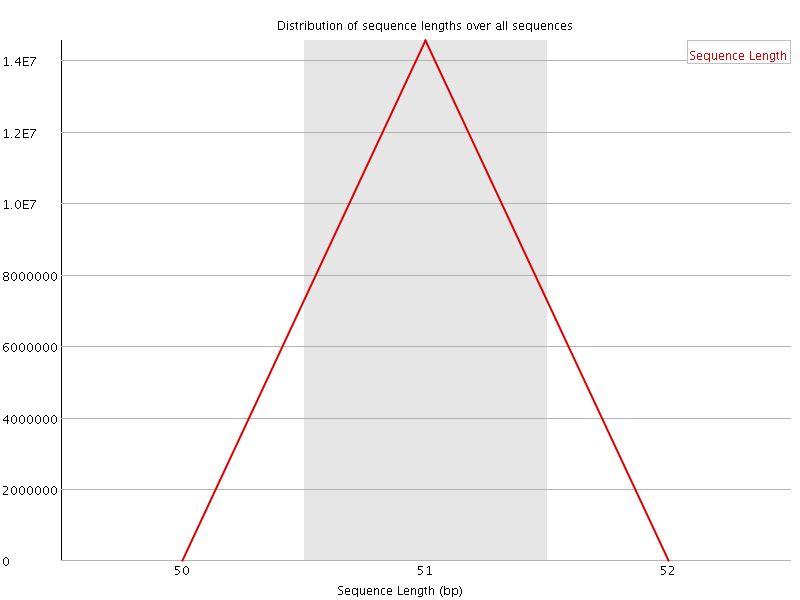
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCGATGTATCTCGTATGCC | 277259 | 1.905227802875347 | TruSeq Adapter, Index 2 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
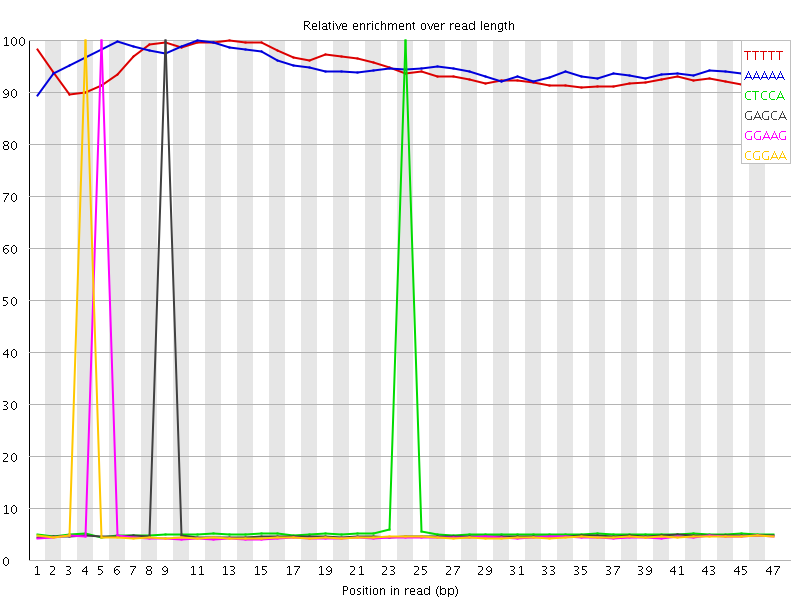
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 6757355 | 3.513844 | 3.721446 | 13 |
| AAAAA | 6760340 | 3.4912324 | 3.6729507 | 11 |
| CTCCA | 1104750 | 2.36695 | 33.1168 | 24 |
| GAGCA | 1059720 | 2.353274 | 34.764187 | 9 |
| GGAAG | 1029140 | 2.3282712 | 35.677624 | 5 |
| CGGAA | 1042025 | 2.3139796 | 35.18489 | 4 |
| TCCAG | 1033375 | 2.255593 | 33.637653 | 25 |
| CACCG | 640890 | 2.243671 | 52.255756 | 31 |
| GAAGA | 1601325 | 2.2140577 | 22.532602 | 6 |
| CTGAA | 1596625 | 2.1698704 | 21.947351 | 19 |
| TCGGA | 900650 | 2.0027952 | 34.82468 | 3 |
| TCACC | 924780 | 1.9813606 | 32.645023 | 30 |
| TCTCG | 903095 | 1.9739463 | 33.182537 | 41 |
| GCACA | 841725 | 1.8347375 | 33.74178 | 11 |
| AGAGC | 825375 | 1.8328744 | 34.314262 | 8 |
| CACAC | 848870 | 1.8162147 | 33.07981 | 12 |
| CGATG | 810720 | 1.8028159 | 33.39662 | 34 |
| AGCAC | 812840 | 1.771776 | 33.691116 | 10 |
| TCTGA | 1296585 | 1.7645378 | 21.602837 | 18 |
| ATCGG | 781025 | 1.7367824 | 34.11936 | 2 |
| CCAGT | 780325 | 1.7032497 | 32.97595 | 26 |
| CCGAT | 773970 | 1.6893783 | 32.76027 | 33 |
| GTCAC | 765720 | 1.6713709 | 32.984776 | 29 |
| CTCGT | 751810 | 1.6432741 | 32.681053 | 42 |
| CACGT | 737950 | 1.6107559 | 33.44758 | 14 |
| CGTCT | 736065 | 1.6088593 | 33.431316 | 16 |
| GTCTG | 721555 | 1.6067528 | 34.08084 | 17 |
| TGAAC | 1167575 | 1.586776 | 21.354053 | 20 |
| ACTCC | 740425 | 1.5863761 | 32.540127 | 23 |
| ACCGA | 727190 | 1.5850815 | 32.67882 | 32 |
| AAGAG | 1118975 | 1.5471407 | 21.789253 | 7 |
| GATCG | 687240 | 1.5282307 | 33.613342 | 1 |
| ACACG | 699630 | 1.5250081 | 33.38356 | 13 |
| ATGCC | 695745 | 1.5186331 | 32.400402 | 47 |
| ACGTC | 685880 | 1.4971004 | 33.327835 | 15 |
| CAGTC | 682890 | 1.490574 | 32.704662 | 27 |
| AACTC | 1066560 | 1.4227823 | 20.733335 | 22 |
| GAACT | 1026940 | 1.3956482 | 21.11217 | 21 |
| ATCTC | 1014560 | 1.355283 | 20.448244 | 40 |
| GATGT | 940420 | 1.3038555 | 21.012121 | 35 |
| AGTCA | 929090 | 1.2626666 | 20.731802 | 28 |
| TCGTA | 744950 | 1.0138112 | 20.282614 | 43 |
| TATGC | 721945 | 0.9825035 | 20.190134 | 46 |
| GTATG | 700470 | 0.9711743 | 20.587772 | 45 |
| GTATC | 713295 | 0.97073156 | 20.304108 | 38 |
| CGTAT | 672930 | 0.9157985 | 20.149813 | 44 |
| TGTAT | 1019065 | 0.8646872 | 12.955917 | 37 |
| ATGTA | 1000660 | 0.84789985 | 12.881766 | 36 |
| TATCT | 919665 | 0.76596534 | 12.608032 | 39 |