![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW228_GAGTGG_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 9431558 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
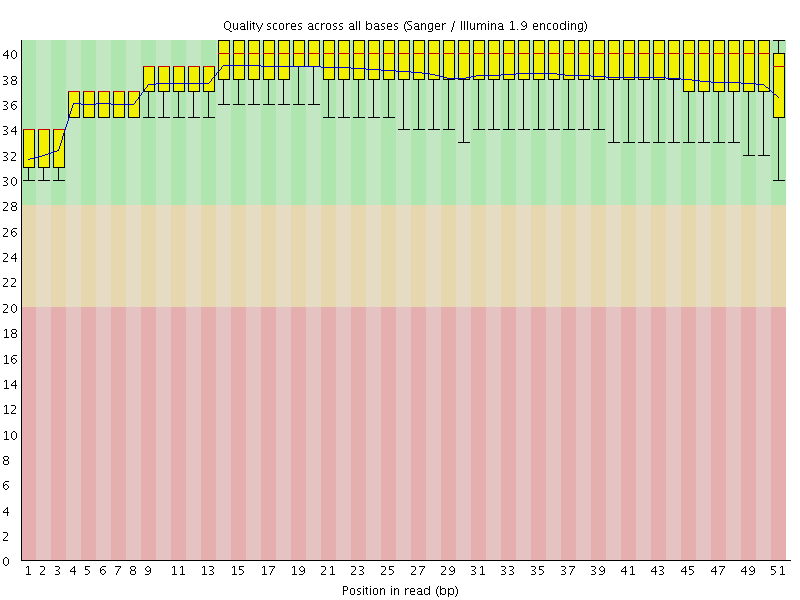
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
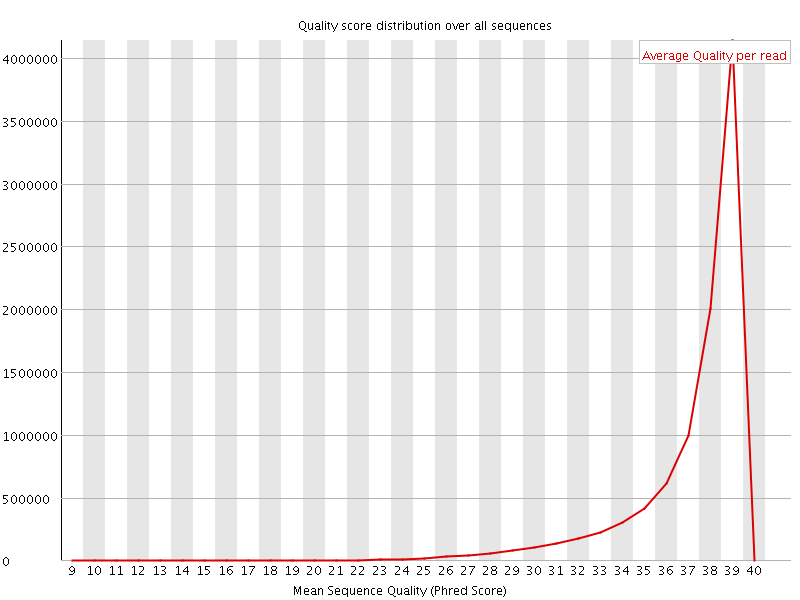
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
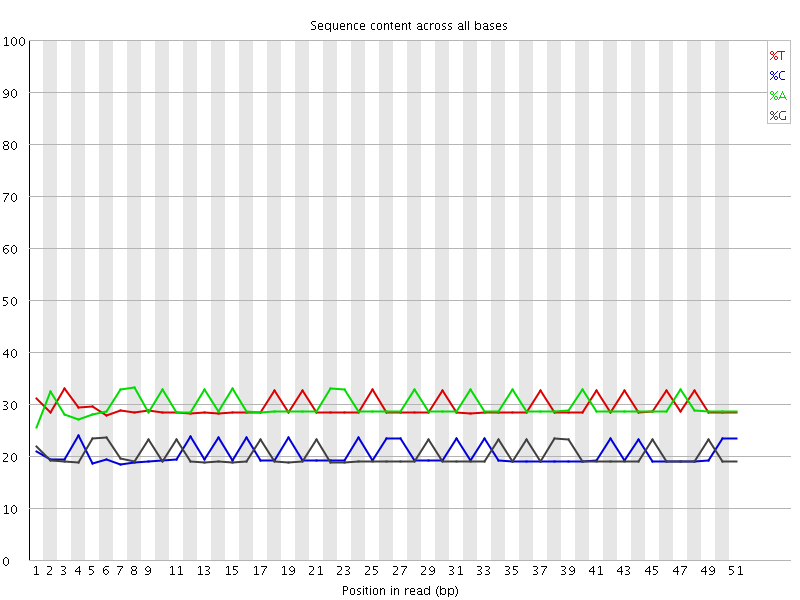
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
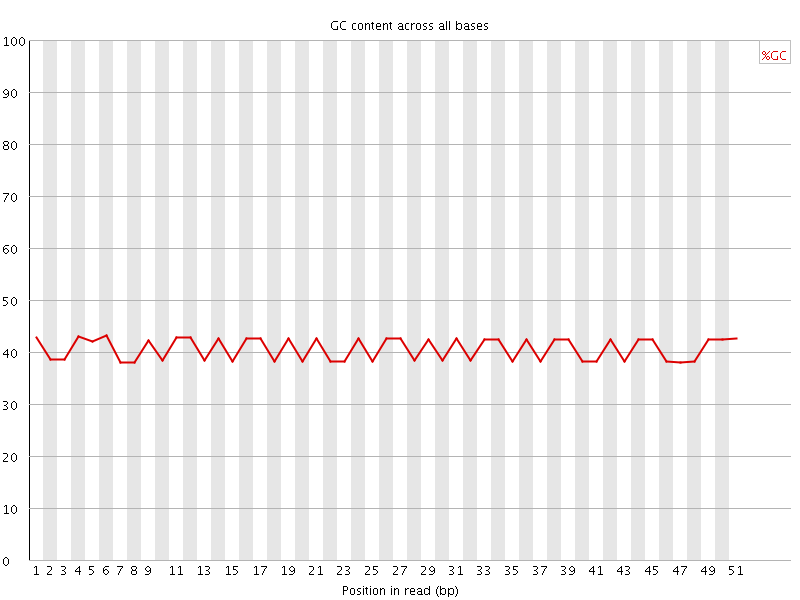
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
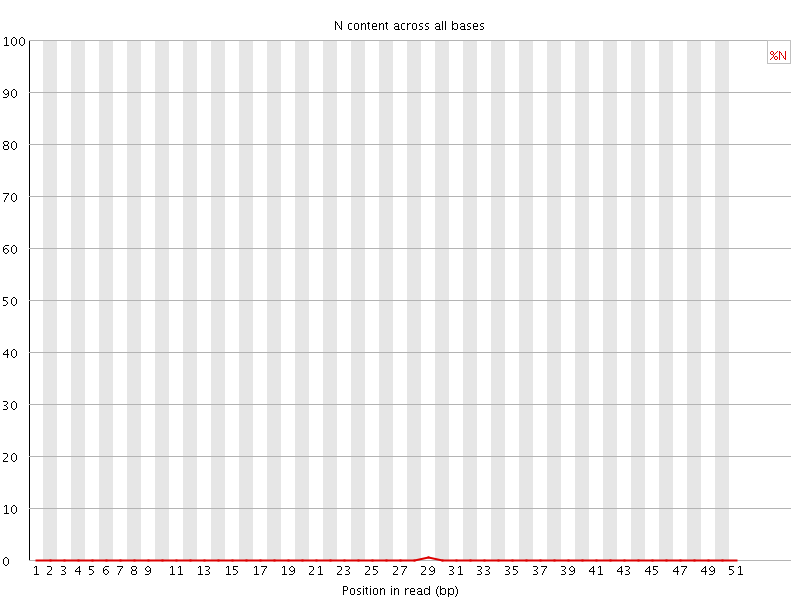
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
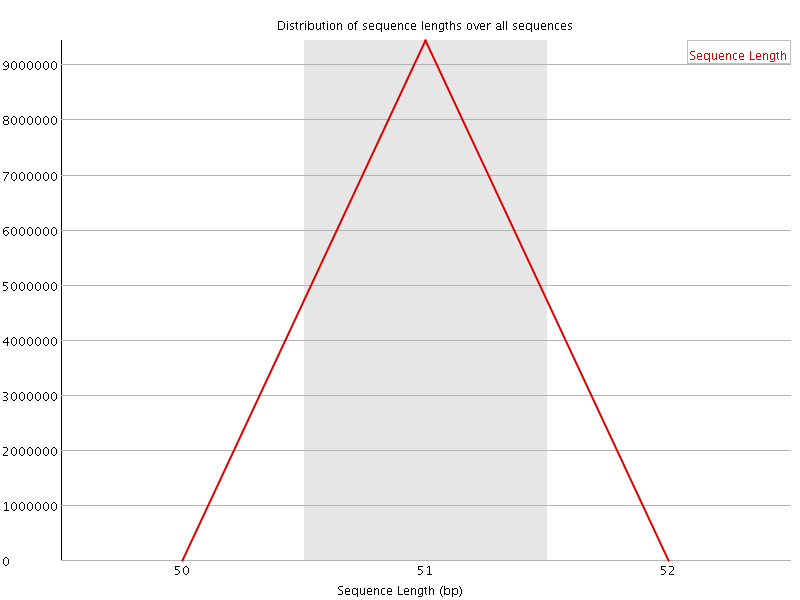
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
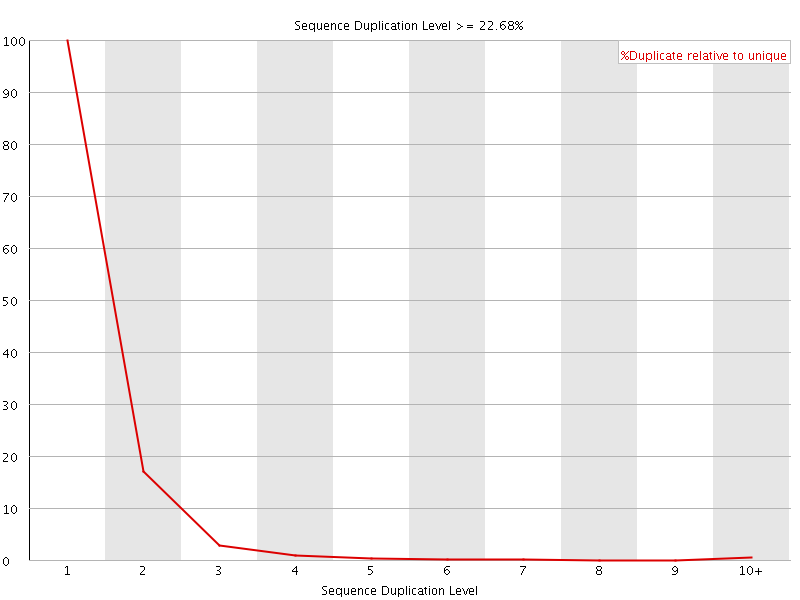
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGAGTGGATCTCGTATGCC | 359815 | 3.8150112632504616 | TruSeq Adapter, Index 7 (97% over 36bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
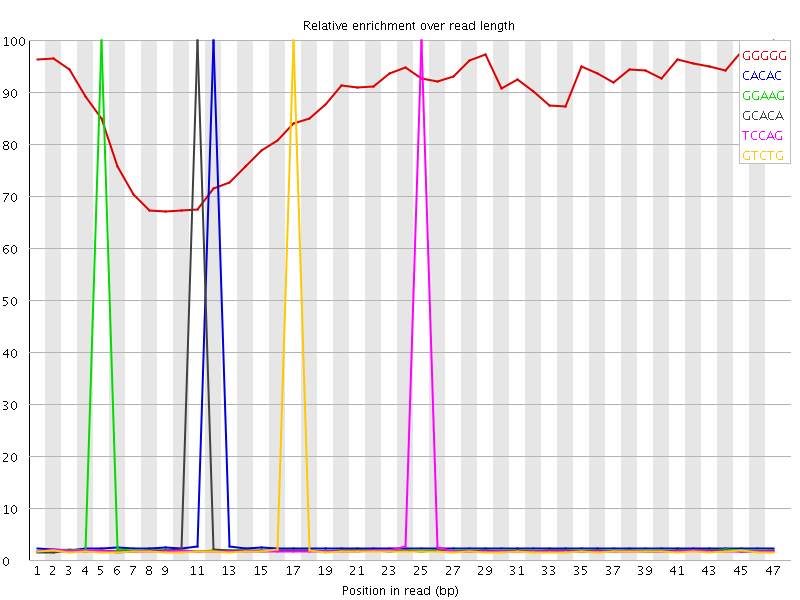
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGGGG | 461200 | 3.0390239 | 3.4538465 | 47 |
| CACAC | 892570 | 2.6461737 | 58.748768 | 12 |
| GGAAG | 811865 | 2.4899714 | 60.64439 | 5 |
| GCACA | 797075 | 2.3899388 | 59.10931 | 11 |
| TCCAG | 775215 | 2.3418343 | 58.724724 | 25 |
| GTCTG | 757755 | 2.3324952 | 60.27331 | 17 |
| CTCCA | 779265 | 2.3275964 | 58.11251 | 24 |
| GAGCA | 759565 | 2.3033712 | 59.564423 | 9 |
| GAAGA | 1083165 | 2.2664077 | 41.920383 | 6 |
| ATGCC | 747130 | 2.2569926 | 57.253735 | 47 |
| GTCAC | 744000 | 2.2475376 | 58.601475 | 29 |
| AGAGC | 733650 | 2.2247846 | 59.466896 | 8 |
| CCAGT | 735665 | 2.2223582 | 58.53915 | 26 |
| GTGGA | 717490 | 2.2170367 | 58.018517 | 36 |
| AGTGG | 708930 | 2.190586 | 58.391346 | 35 |
| CAGTC | 722860 | 2.1836758 | 58.434235 | 27 |
| GAGTG | 697810 | 2.1562254 | 58.588646 | 34 |
| AGCAC | 710170 | 2.1293638 | 58.96429 | 10 |
| AAGAG | 983000 | 2.0568233 | 41.56079 | 7 |
| CTGAA | 973080 | 2.0282753 | 41.280537 | 19 |
| ACGTC | 671315 | 2.0279644 | 59.05383 | 15 |
| GATCG | 662535 | 2.0242043 | 58.877407 | 1 |
| CACGT | 668555 | 2.0196269 | 59.0599 | 14 |
| ACACG | 666705 | 1.999039 | 58.670746 | 13 |
| TCTCG | 652595 | 1.9862056 | 57.94613 | 41 |
| ACTCC | 663380 | 1.9814581 | 57.936935 | 23 |
| TCACG | 655445 | 1.980023 | 58.11322 | 30 |
| CGGAA | 650255 | 1.9718902 | 59.87058 | 4 |
| GGATC | 642675 | 1.9635268 | 57.624836 | 38 |
| CGTCT | 644680 | 1.9621159 | 59.323288 | 16 |
| ATCGG | 639100 | 1.9526047 | 59.30699 | 2 |
| CACGA | 640540 | 1.9205863 | 57.45097 | 31 |
| CTCGT | 617920 | 1.8806704 | 57.475765 | 42 |
| ACGAG | 618920 | 1.8768673 | 58.008766 | 32 |
| TCTGA | 890605 | 1.8702943 | 41.37595 | 18 |
| TGAAC | 887270 | 1.8494142 | 41.031944 | 20 |
| TCGGA | 604975 | 1.8483446 | 60.121822 | 3 |
| CGAGT | 599170 | 1.8306091 | 57.668747 | 33 |
| TGGAT | 839140 | 1.7822587 | 40.128838 | 37 |
| ATCTC | 848800 | 1.7624576 | 39.71787 | 40 |
| GTATG | 814435 | 1.7297876 | 40.37557 | 45 |
| AGTCA | 822560 | 1.7145333 | 40.58623 | 28 |
| TATGC | 797365 | 1.6744878 | 39.75883 | 46 |
| GATCT | 780380 | 1.6388187 | 39.973484 | 39 |
| GAACT | 778020 | 1.6216949 | 40.819035 | 21 |
| AACTC | 783865 | 1.6155044 | 40.30456 | 22 |
| TCGTA | 652235 | 1.3697108 | 39.579494 | 43 |
| CGTAT | 650755 | 1.3666029 | 39.545067 | 44 |