![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW227_CGTACG_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 5742321 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
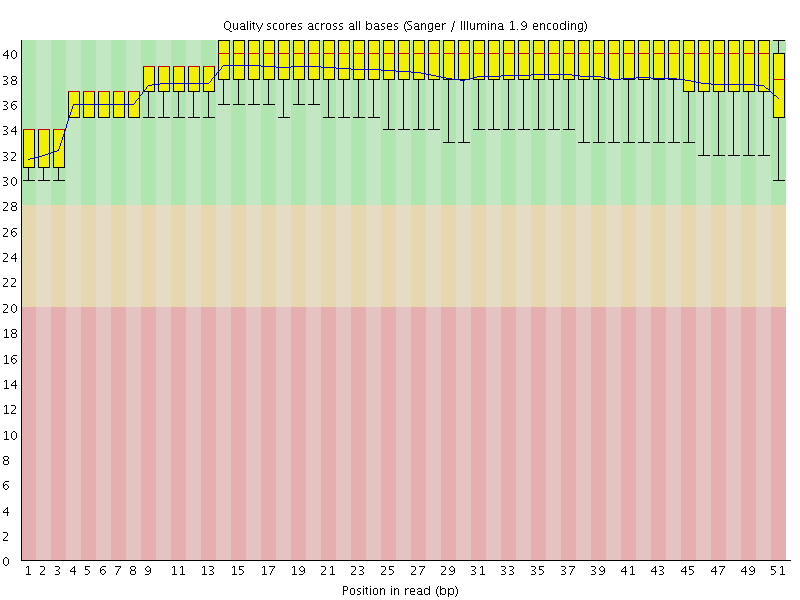
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
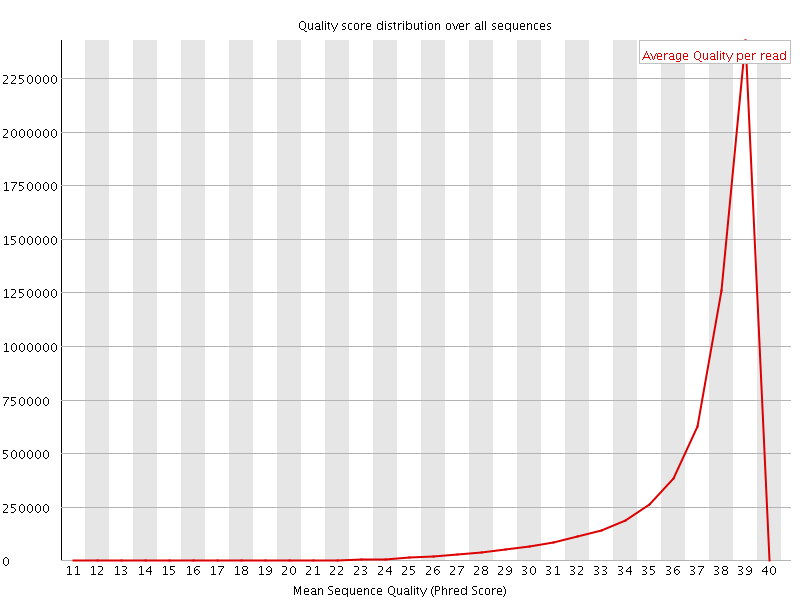
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
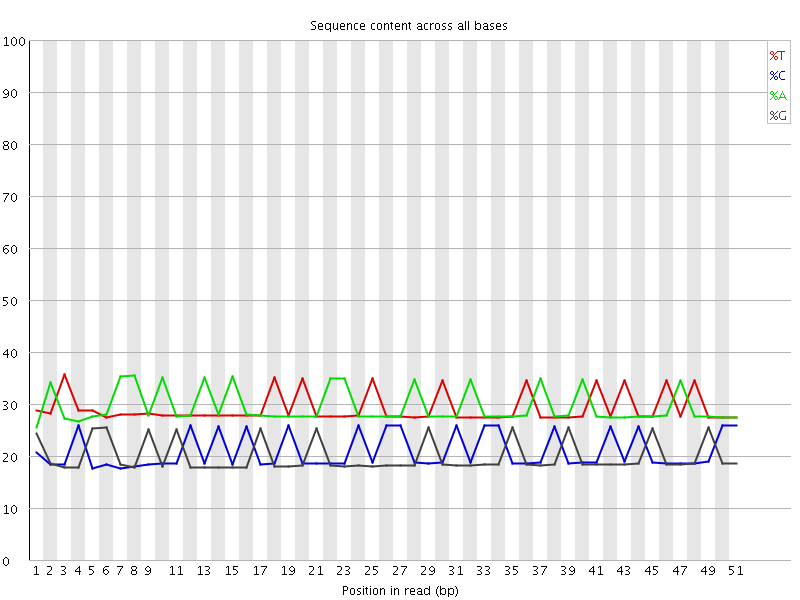
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
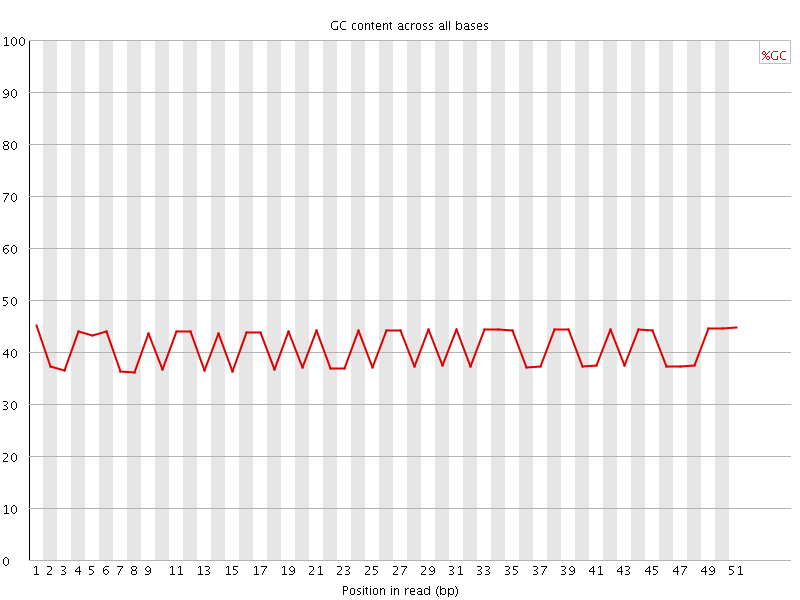
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
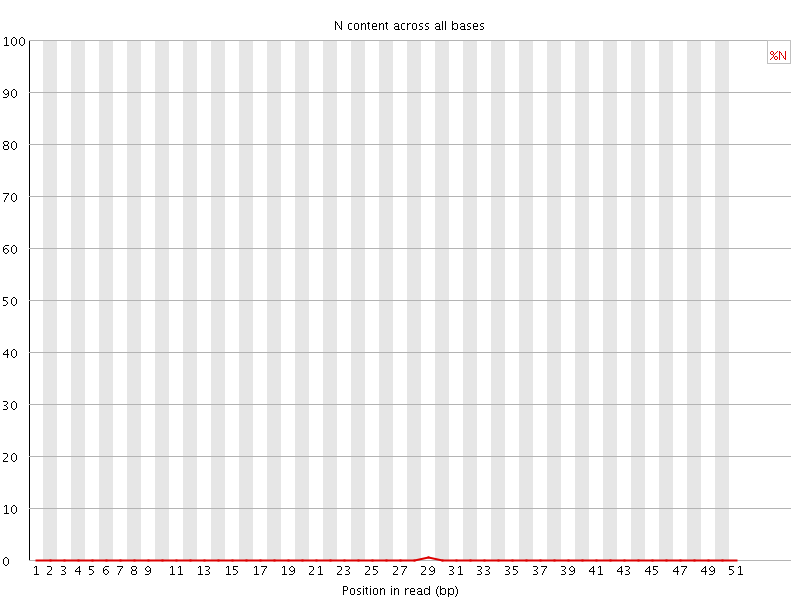
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
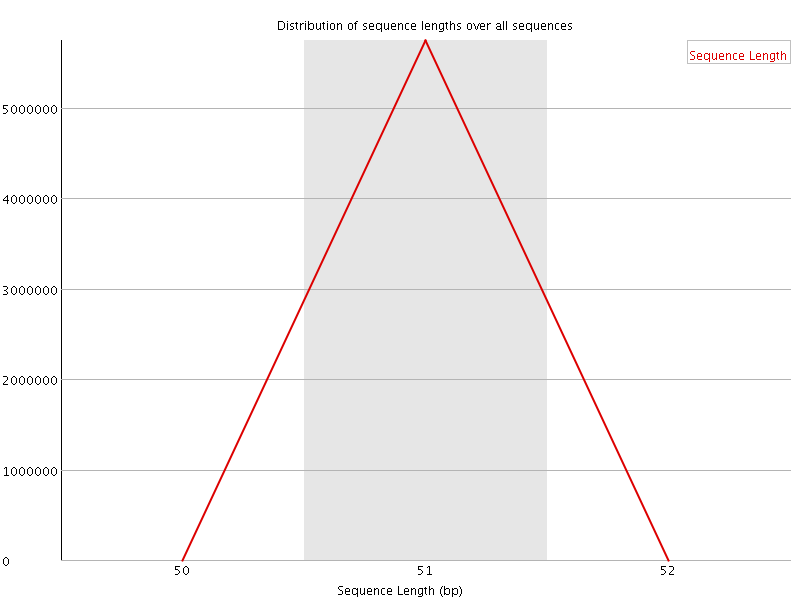
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
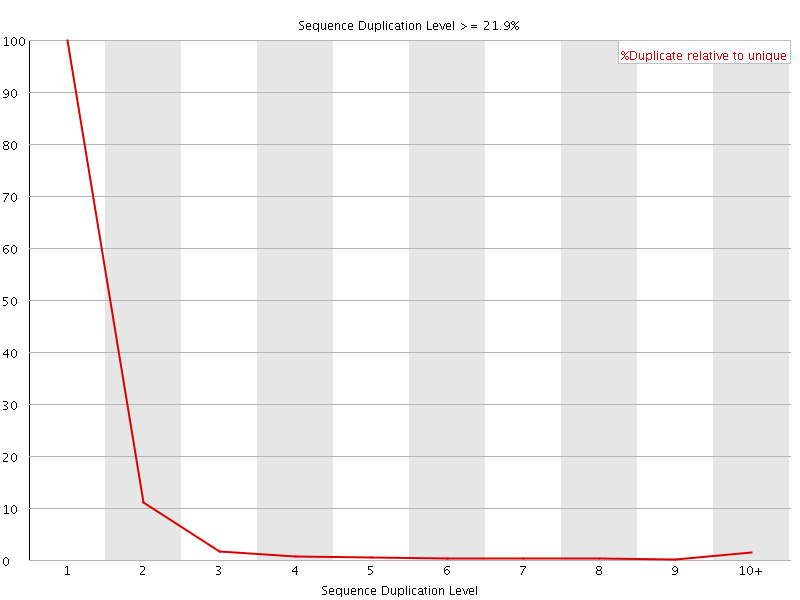
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCGTACGATCTCGTATGCC | 362672 | 6.315773708923622 | TruSeq Adapter, Index 12 (97% over 36bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
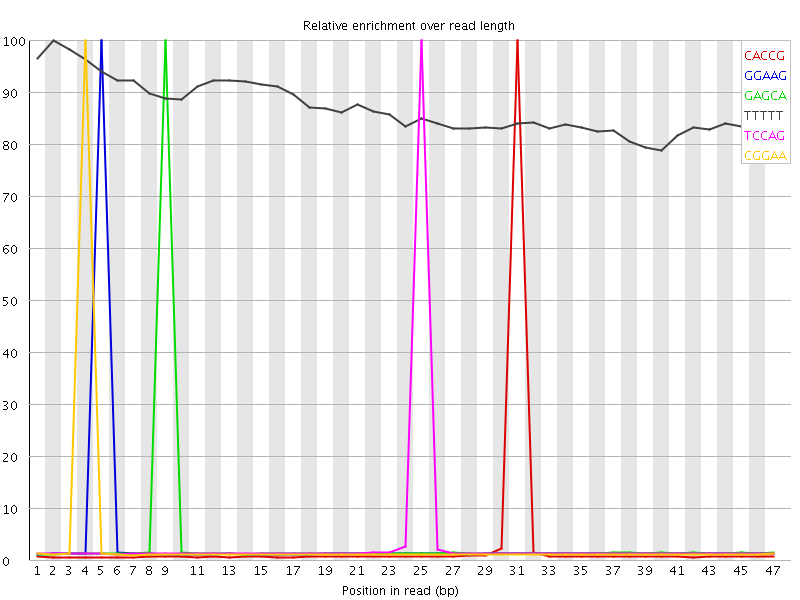
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CACCG | 571790 | 3.8331487 | 129.20018 | 31 |
| GGAAG | 702515 | 3.6898572 | 105.35108 | 5 |
| GAGCA | 714135 | 3.5719445 | 99.94574 | 9 |
| TTTTT | 2033450 | 3.4943783 | 4.0157247 | 2 |
| TCCAG | 708620 | 3.4123774 | 94.706955 | 25 |
| CGGAA | 680350 | 3.4029593 | 100.892006 | 4 |
| CTCCA | 738020 | 3.3844044 | 90.26742 | 24 |
| AAAAA | 2030165 | 3.303115 | 3.5614066 | 14 |
| TCGGA | 630915 | 3.1903923 | 101.71277 | 3 |
| GAAGA | 892130 | 3.1705034 | 71.831894 | 6 |
| AGAGC | 626265 | 3.1324382 | 99.57582 | 8 |
| TCTCG | 630915 | 3.0715904 | 92.984955 | 41 |
| AGCAC | 643225 | 3.0637813 | 95.20084 | 10 |
| ATCGG | 596940 | 3.018588 | 100.43052 | 2 |
| TCACC | 656985 | 3.012795 | 88.61489 | 30 |
| GTCTG | 587030 | 3.0011127 | 100.929436 | 17 |
| GCACA | 627870 | 2.9906425 | 94.950325 | 11 |
| CTGAA | 868350 | 2.9710786 | 68.431526 | 19 |
| CCAGT | 610975 | 2.9421654 | 93.894745 | 26 |
| CGTCT | 592655 | 2.8853226 | 96.20351 | 16 |
| CACAC | 634435 | 2.8777456 | 90.2959 | 12 |
| GATCG | 568075 | 2.8726246 | 99.4756 | 1 |
| CTCGT | 582450 | 2.8356402 | 92.226425 | 42 |
| GTCAC | 587715 | 2.8301563 | 93.92944 | 29 |
| CACGT | 586835 | 2.8259187 | 95.30911 | 14 |
| ATGCC | 580560 | 2.7957013 | 91.04223 | 47 |
| ACGTC | 570270 | 2.746149 | 95.18783 | 15 |
| CAGTC | 568450 | 2.7373848 | 93.66609 | 27 |
| ACACG | 571745 | 2.7233107 | 94.38445 | 13 |
| ACTCC | 592255 | 2.7159564 | 89.98906 | 23 |
| CGATC | 559165 | 2.6926727 | 90.967995 | 38 |
| ACCGT | 556775 | 2.6811638 | 92.42767 | 32 |
| TCTGA | 766535 | 2.651552 | 68.914276 | 18 |
| CCGTA | 535870 | 2.5804956 | 92.3307 | 33 |
| AAGAG | 725140 | 2.5770447 | 71.15019 | 7 |
| GTACG | 508090 | 2.5692945 | 97.016266 | 35 |
| TGAAC | 723045 | 2.4739144 | 67.836586 | 20 |
| CGTAC | 509840 | 2.455147 | 91.93108 | 34 |
| GAACT | 680250 | 2.3274903 | 67.61884 | 21 |
| AACTC | 696780 | 2.2703114 | 64.42245 | 22 |
| ACGAT | 657510 | 2.2496848 | 65.46277 | 37 |
| ATCTC | 678595 | 2.235369 | 62.767204 | 40 |
| GATCT | 640465 | 2.2154582 | 65.62526 | 39 |
| AGTCA | 638220 | 2.1836836 | 66.7741 | 28 |
| TCGTA | 604835 | 2.092209 | 65.447716 | 43 |
| GTATG | 566285 | 2.0569928 | 68.49853 | 45 |
| TACGA | 599830 | 2.0523314 | 65.40083 | 36 |
| TATGC | 577730 | 1.9984488 | 65.14119 | 46 |
| CGTAT | 573485 | 1.9837648 | 65.29386 | 44 |
| TGCCG | 210770 | 1.5000519 | 5.243103 | 47 |