![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW226_GTTTCG_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 8424700 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
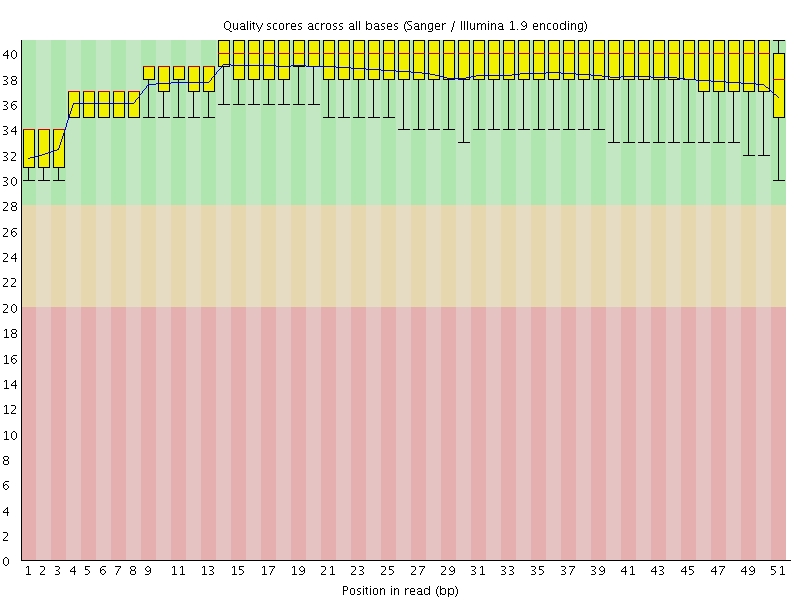
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
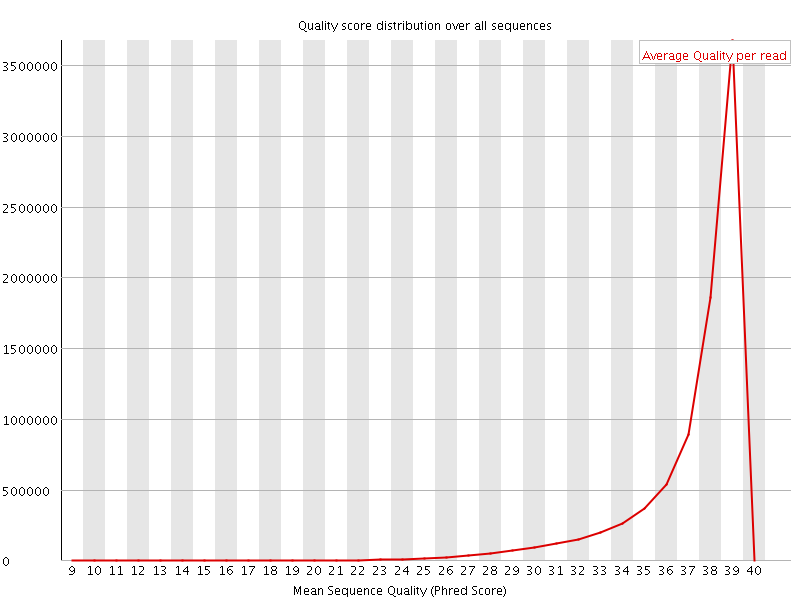
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
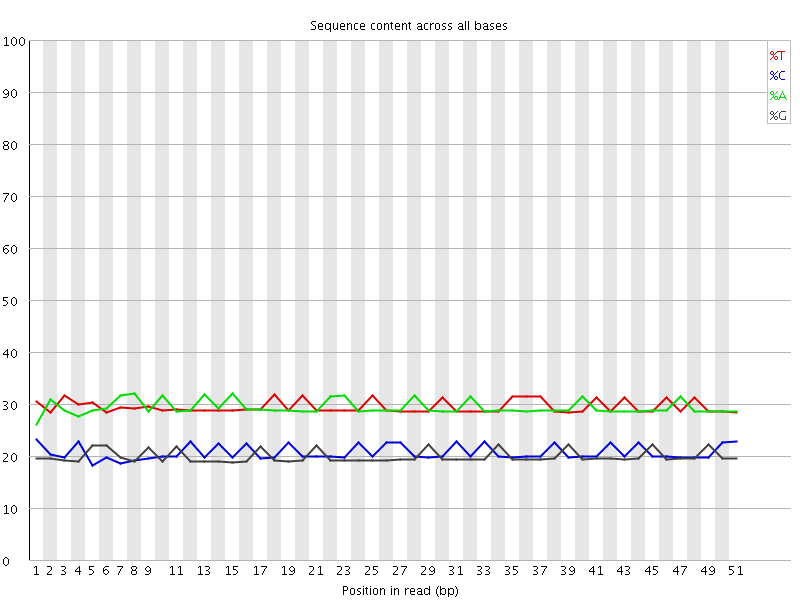
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
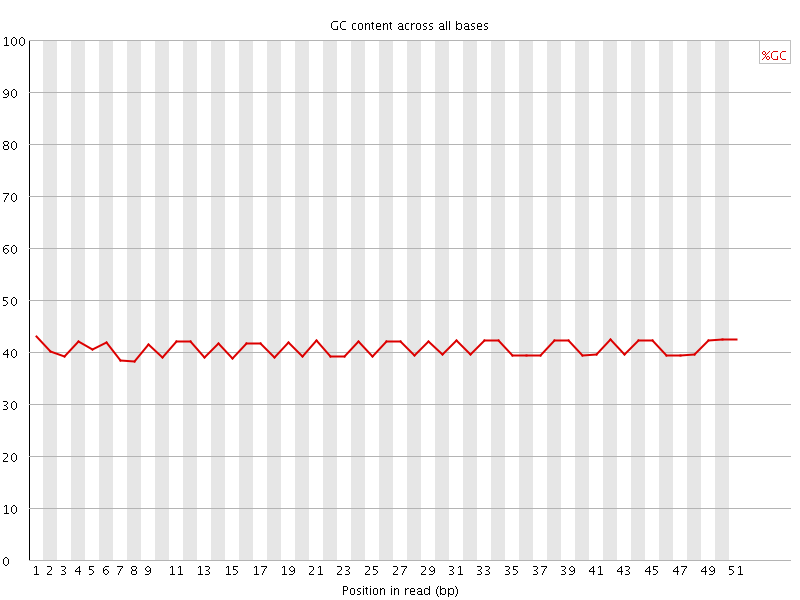
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
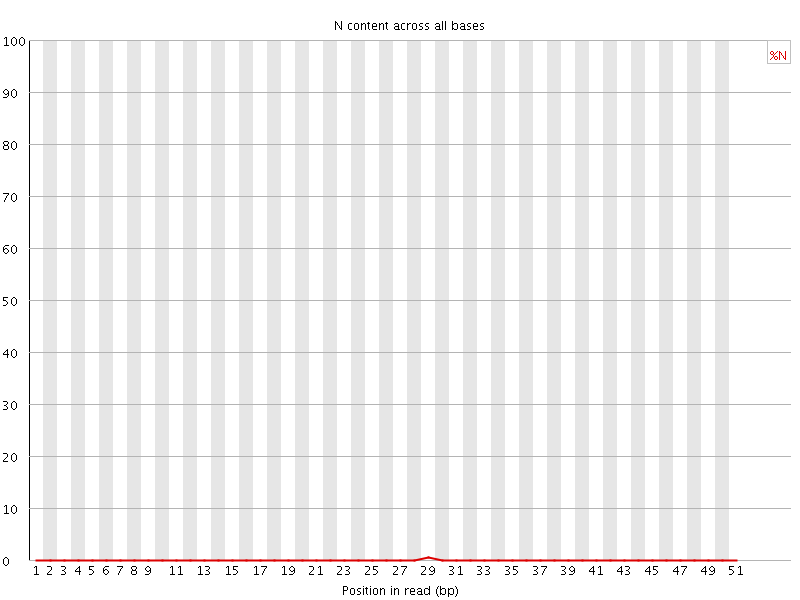
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
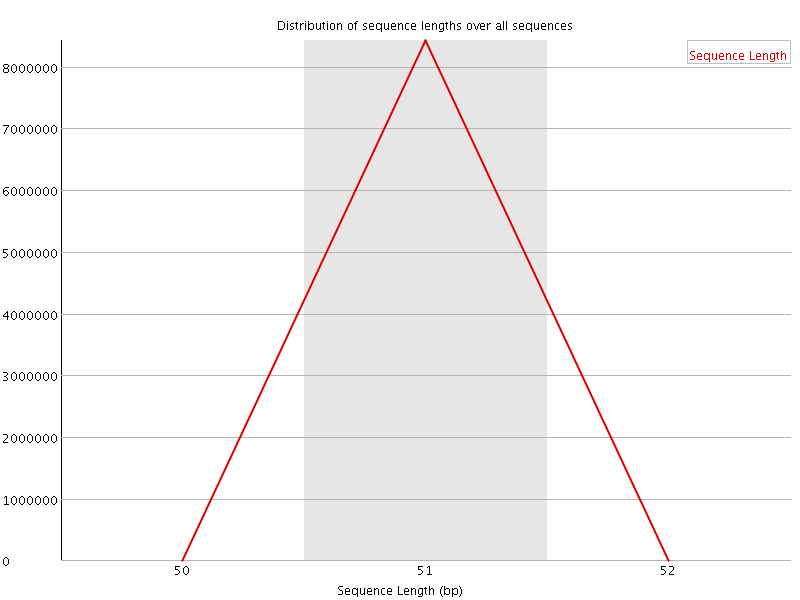
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGTTTCGATCTCGTATGCC | 212364 | 2.5207307085118758 | TruSeq Adapter, Index 9 (97% over 36bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
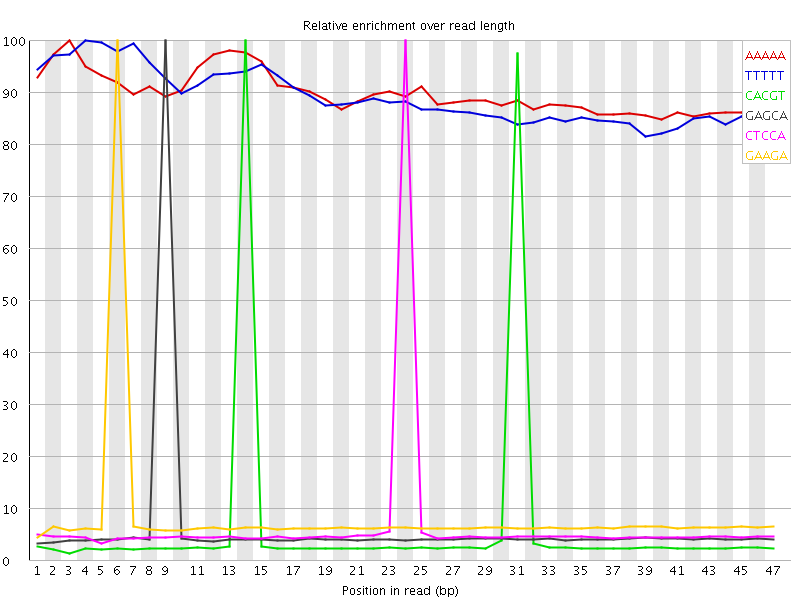
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 2865375 | 3.2407036 | 3.607802 | 3 |
| TTTTT | 2838300 | 3.1593072 | 3.5486457 | 4 |
| CACGT | 768160 | 2.5525568 | 38.730377 | 14 |
| GAGCA | 734050 | 2.541878 | 41.273544 | 9 |
| CTCCA | 784765 | 2.5104 | 37.81573 | 24 |
| GAAGA | 1013860 | 2.479878 | 29.900915 | 6 |
| GGAAG | 688735 | 2.4774315 | 43.03344 | 5 |
| TCCAG | 704485 | 2.3409681 | 38.890293 | 25 |
| CGGAA | 652730 | 2.260282 | 41.619072 | 4 |
| CTGAA | 933160 | 2.1902976 | 28.184824 | 19 |
| TCGGA | 596620 | 2.059406 | 41.218327 | 3 |
| AGCAC | 607685 | 2.025756 | 39.512295 | 10 |
| TTTCG | 865005 | 2.017418 | 26.873314 | 35 |
| TCTCG | 606035 | 2.0074124 | 37.993416 | 41 |
| AGAGC | 579575 | 2.00696 | 40.762985 | 8 |
| GCACA | 573960 | 1.9133314 | 39.27671 | 11 |
| CACAC | 593480 | 1.9045582 | 37.790356 | 12 |
| GTTTC | 816270 | 1.9037552 | 26.794865 | 34 |
| TTCGA | 798900 | 1.8691946 | 26.609123 | 36 |
| TCTGA | 782590 | 1.8310342 | 27.814215 | 18 |
| ATCGG | 526740 | 1.8181952 | 40.41521 | 2 |
| AAGAG | 730455 | 1.7866757 | 29.156086 | 7 |
| CCAGT | 537085 | 1.7847062 | 38.21435 | 26 |
| CTCGT | 529840 | 1.7550263 | 37.558247 | 42 |
| CGTCT | 524700 | 1.7380006 | 38.57608 | 16 |
| GTCTG | 496030 | 1.7067397 | 39.89711 | 17 |
| GTCAC | 510930 | 1.6977946 | 38.136765 | 29 |
| ACTCC | 523080 | 1.6732907 | 37.02111 | 23 |
| TGAAC | 711135 | 1.6691642 | 27.660748 | 20 |
| GATCG | 479600 | 1.6554778 | 39.8594 | 1 |
| ACACG | 492675 | 1.6423628 | 38.846252 | 13 |
| ATGCC | 492240 | 1.6356887 | 37.473988 | 47 |
| ACGTC | 491380 | 1.632831 | 38.65092 | 15 |
| CGTTT | 696310 | 1.6239771 | 26.807283 | 33 |
| TCACG | 485895 | 1.6146045 | 37.926205 | 30 |
| CAGTC | 481670 | 1.6005651 | 37.981194 | 27 |
| TCGAT | 682320 | 1.5964313 | 26.384981 | 37 |
| CGATC | 474080 | 1.5753438 | 36.682915 | 38 |
| AACTC | 680905 | 1.5385551 | 26.588703 | 22 |
| GAACT | 648905 | 1.523099 | 27.516346 | 21 |
| ATCTC | 667055 | 1.5024616 | 25.697777 | 40 |
| GATCT | 610695 | 1.4288495 | 26.536924 | 39 |
| AGTCA | 582305 | 1.3667766 | 27.020704 | 28 |
| ACGTT | 541695 | 1.2674094 | 26.626202 | 32 |
| TCGTA | 533915 | 1.2492065 | 26.44637 | 43 |
| TATGC | 487460 | 1.1405152 | 26.245848 | 46 |
| GTATG | 466945 | 1.1348757 | 27.276318 | 45 |
| CGTAT | 479270 | 1.121353 | 26.273157 | 44 |