![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW225_GTGGCC_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 9287907 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
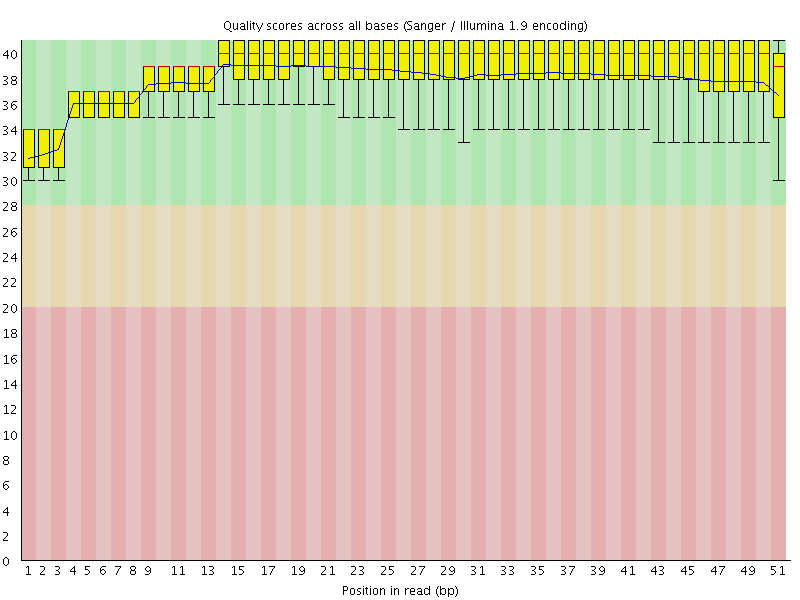
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
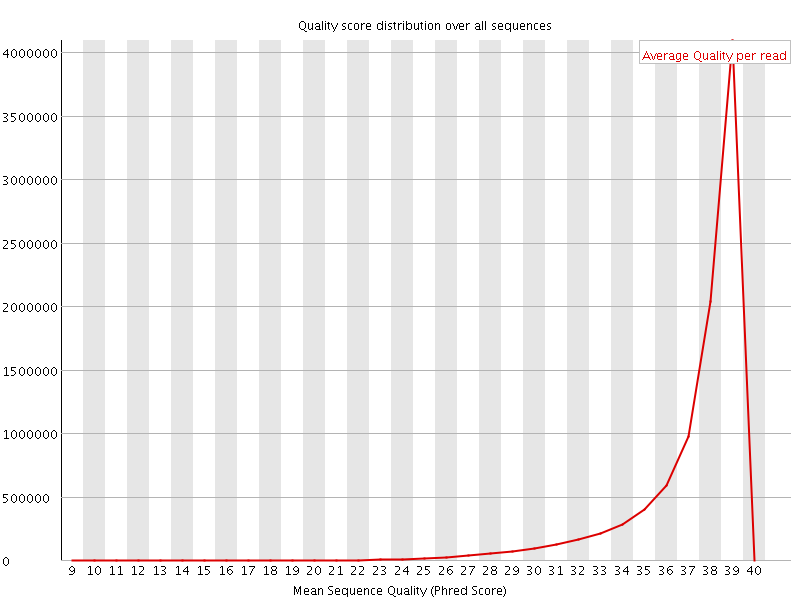
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
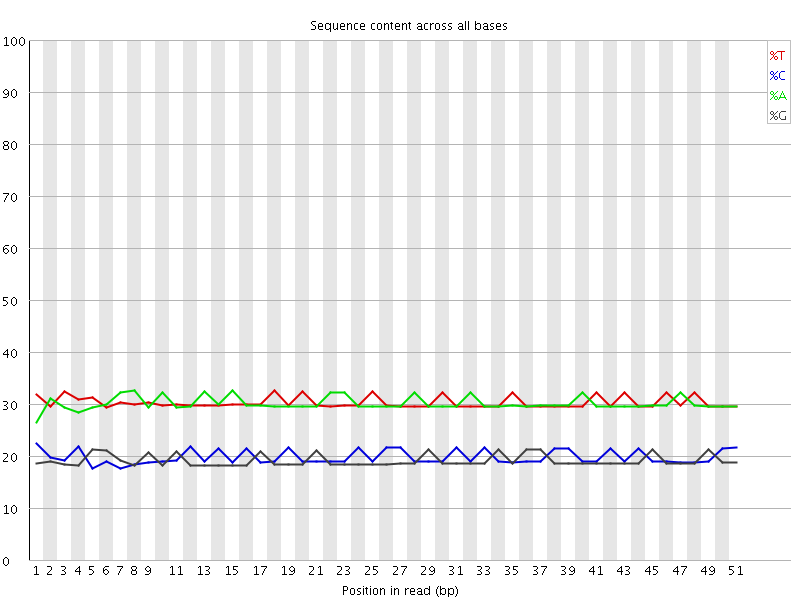
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
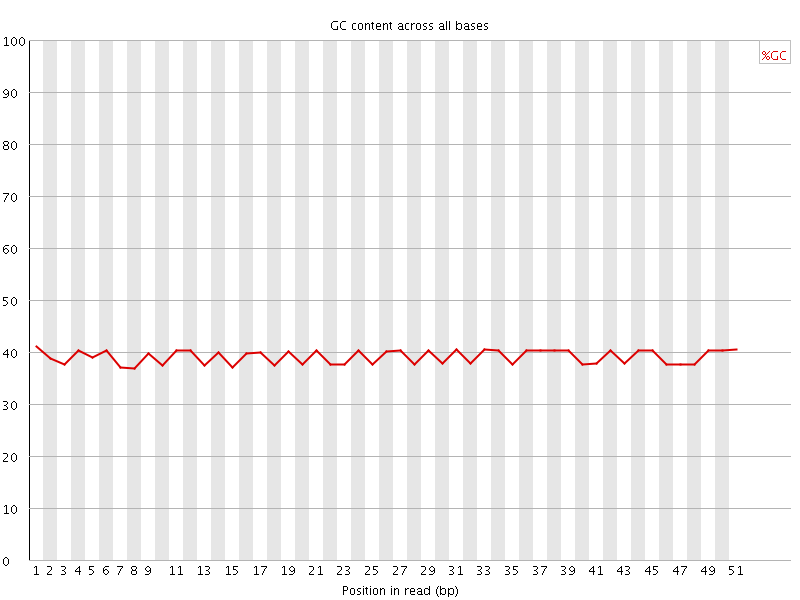
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
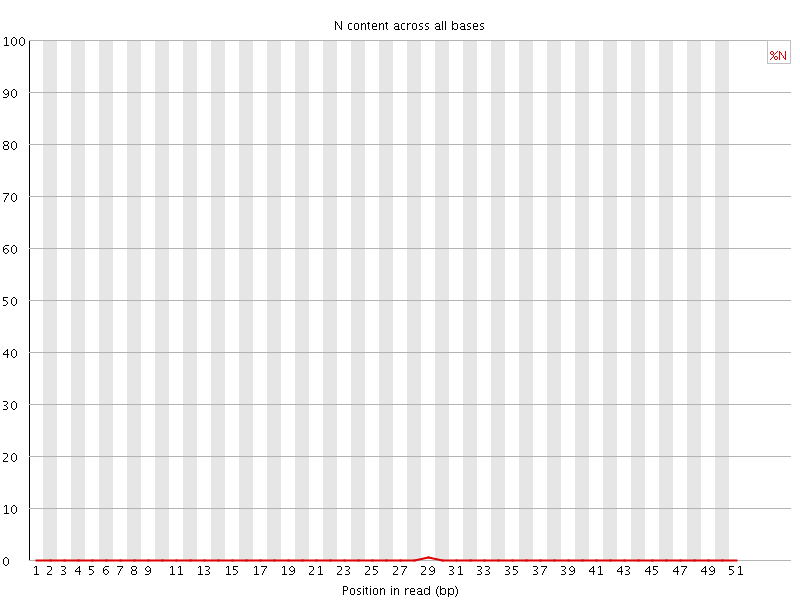
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
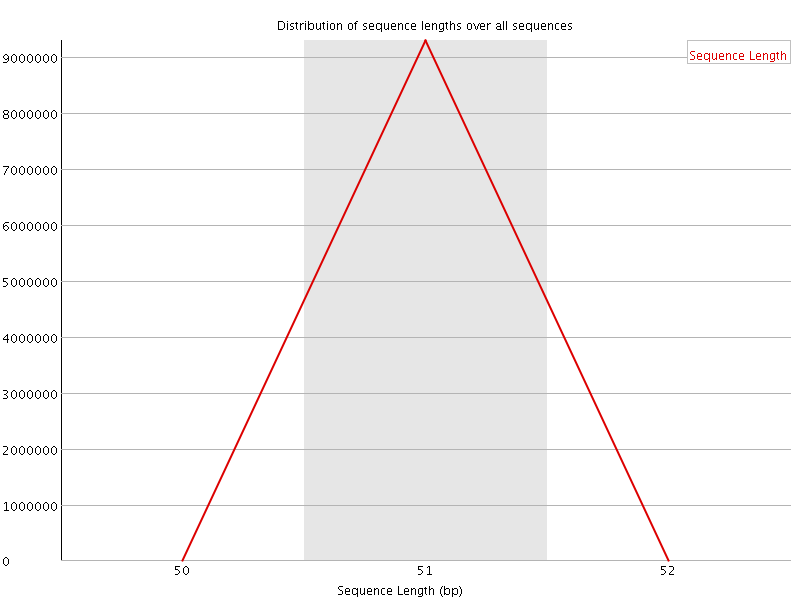
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
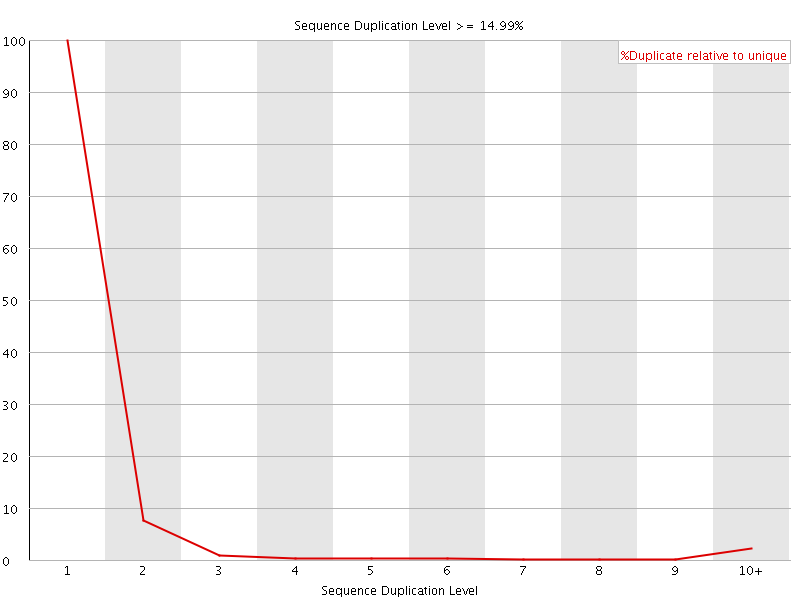
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGTGGCCATCTCGTATGCC | 218624 | 2.3538564716464108 | TruSeq Adapter, Index 1 (97% over 35bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
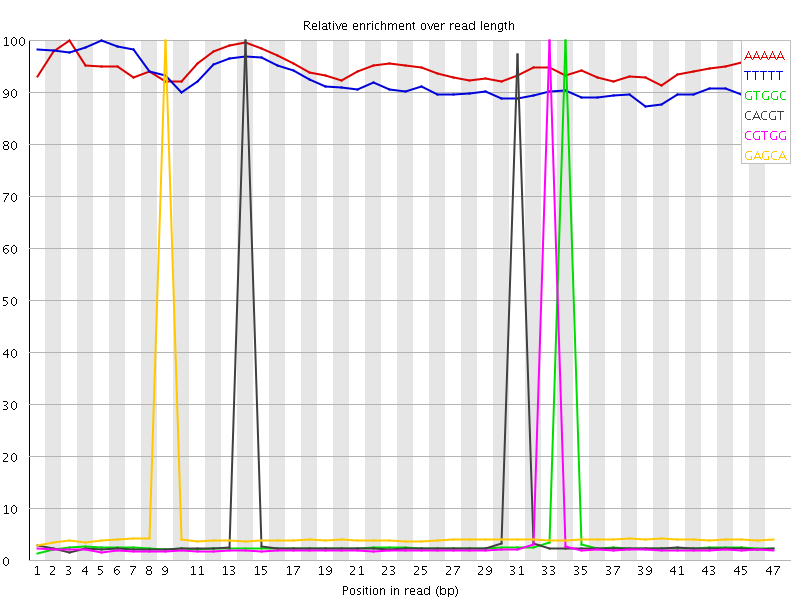
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 3814250 | 3.4279227 | 3.6232622 | 3 |
| TTTTT | 3851445 | 3.3731549 | 3.6561723 | 5 |
| GTGGC | 527780 | 2.7729576 | 60.745415 | 34 |
| CACGT | 785955 | 2.5407634 | 38.86751 | 14 |
| CGTGG | 481935 | 2.5320878 | 60.93336 | 33 |
| GAGCA | 740130 | 2.489049 | 41.17301 | 9 |
| GGAAG | 705650 | 2.4560187 | 42.791542 | 5 |
| CTCCA | 783795 | 2.4482293 | 37.882607 | 24 |
| CGGAA | 709180 | 2.384964 | 41.66402 | 4 |
| GGCCA | 467040 | 2.3832483 | 58.400063 | 36 |
| TGGCC | 465595 | 2.3636417 | 57.93553 | 35 |
| TCCAG | 715315 | 2.312405 | 38.99145 | 25 |
| GAAGA | 1042375 | 2.310253 | 27.893745 | 6 |
| CTGAA | 1039195 | 2.2139804 | 26.612173 | 19 |
| TCGGA | 614325 | 2.0553308 | 41.032406 | 3 |
| TCTCG | 624450 | 2.008271 | 37.90778 | 41 |
| AGCAC | 615790 | 2.0009725 | 39.614143 | 10 |
| AGAGC | 589540 | 1.9826164 | 40.734016 | 8 |
| CCATC | 626480 | 1.9568468 | 36.253468 | 38 |
| GCACA | 597065 | 1.9401264 | 39.415024 | 11 |
| CACAC | 606750 | 1.9050275 | 38.070934 | 12 |
| ATCGG | 547915 | 1.8331447 | 40.187225 | 2 |
| TCTGA | 853830 | 1.8096985 | 26.119131 | 18 |
| CCAGT | 554265 | 1.7917771 | 38.34144 | 26 |
| ACGTG | 527690 | 1.765478 | 39.075653 | 32 |
| GTCAC | 536805 | 1.735334 | 38.236134 | 29 |
| CTCGT | 538365 | 1.7314163 | 37.388317 | 42 |
| GTCTG | 513355 | 1.7086749 | 39.875423 | 17 |
| CGTCT | 528665 | 1.7002205 | 38.603317 | 16 |
| AAGAG | 754080 | 1.6712945 | 27.239668 | 7 |
| ACTCC | 530760 | 1.6578597 | 37.290203 | 23 |
| GCCAT | 512710 | 1.6574419 | 37.14452 | 37 |
| ACACG | 508755 | 1.6531684 | 39.07172 | 13 |
| TGAAC | 775495 | 1.6521738 | 26.034481 | 20 |
| ATGCC | 503505 | 1.6276848 | 37.370773 | 47 |
| GATCG | 483195 | 1.6166126 | 39.577644 | 1 |
| TCACG | 500055 | 1.6165322 | 37.9838 | 30 |
| ACGTC | 495760 | 1.6026475 | 38.731762 | 15 |
| CAGTC | 489670 | 1.5829602 | 38.161175 | 27 |
| AACTC | 724145 | 1.490683 | 24.99923 | 22 |
| GAACT | 690750 | 1.4716265 | 25.78085 | 21 |
| CATCT | 715935 | 1.466194 | 24.097904 | 39 |
| ATCTC | 696305 | 1.4259932 | 24.29403 | 40 |
| AGTCA | 628115 | 1.3381841 | 25.360622 | 28 |
| TCGTA | 564475 | 1.1964086 | 24.573133 | 43 |
| TATGC | 512275 | 1.0857704 | 24.417978 | 46 |
| GTATG | 491920 | 1.0790615 | 25.25489 | 45 |
| CGTAT | 508855 | 1.0785217 | 24.420204 | 44 |