![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW224_GTGAAA_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 8965702 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
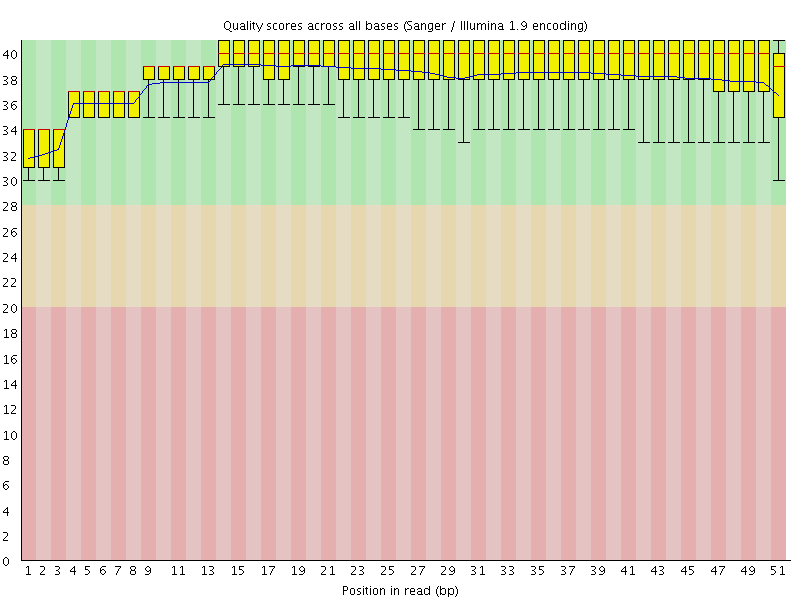
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
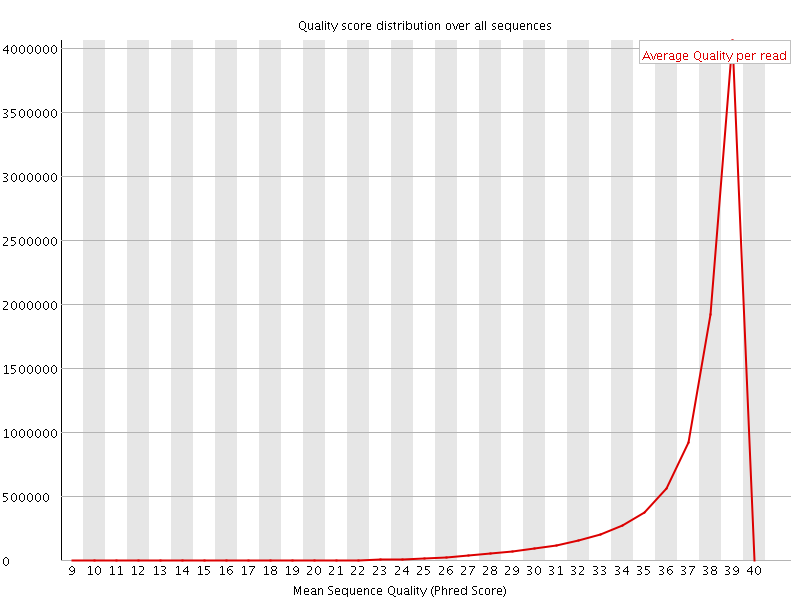
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
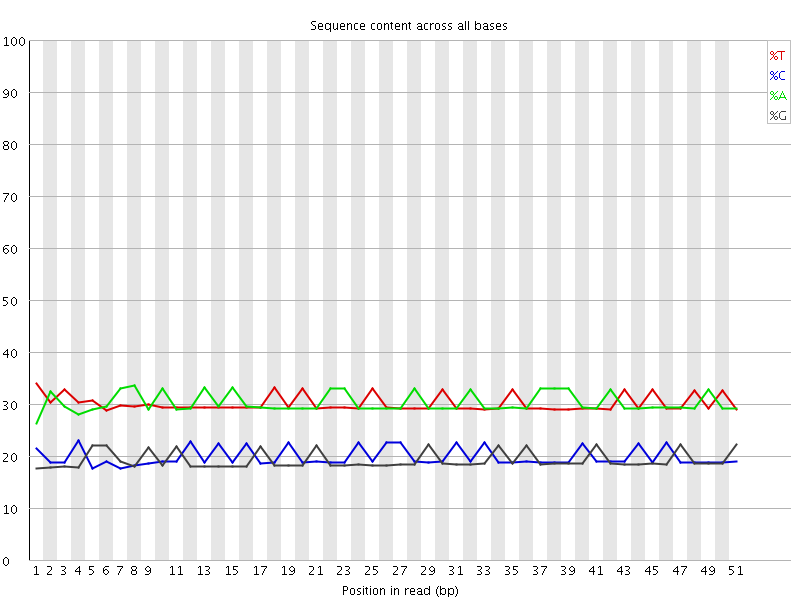
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
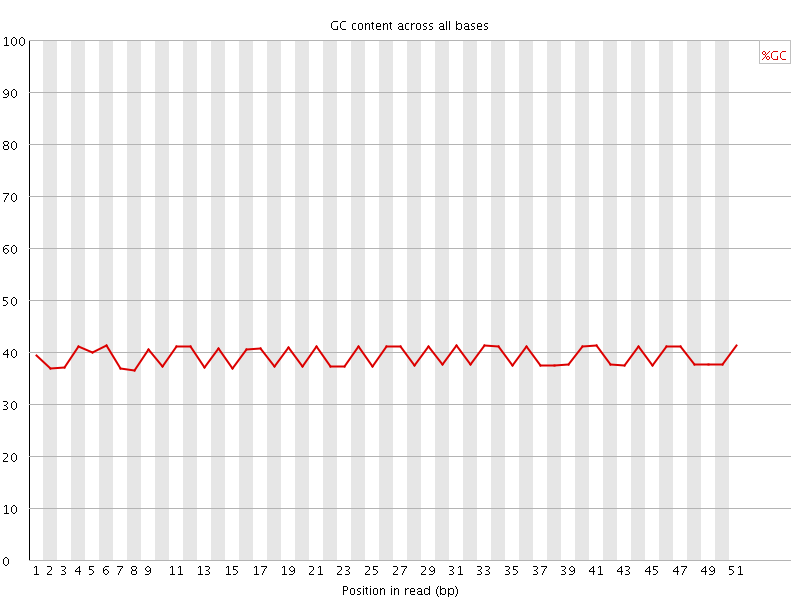
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
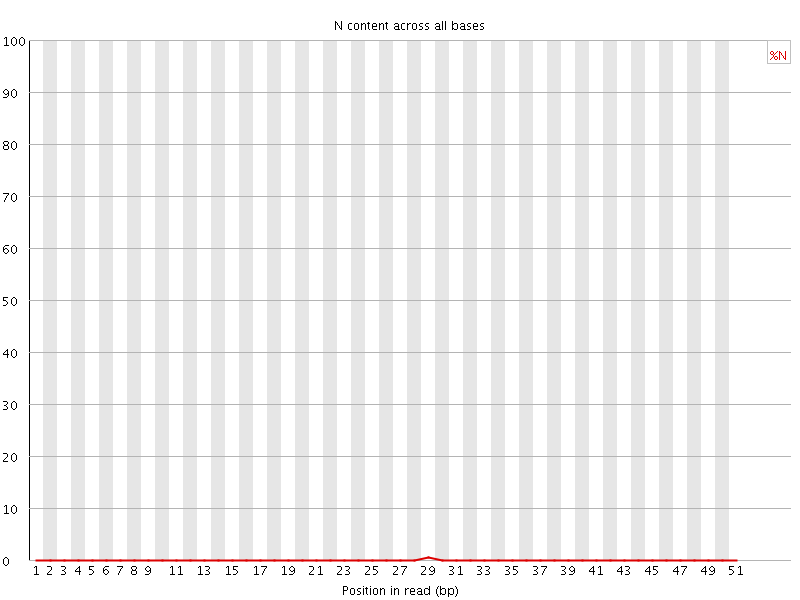
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
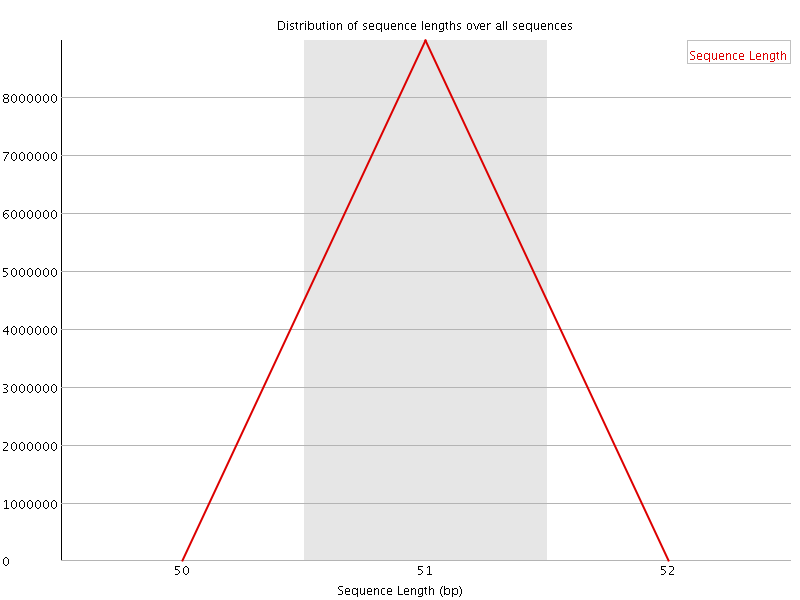
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGTGAAACGATCTCGTATG | 279804 | 3.1208264561994143 | TruSeq Adapter, Index 1 (97% over 35bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
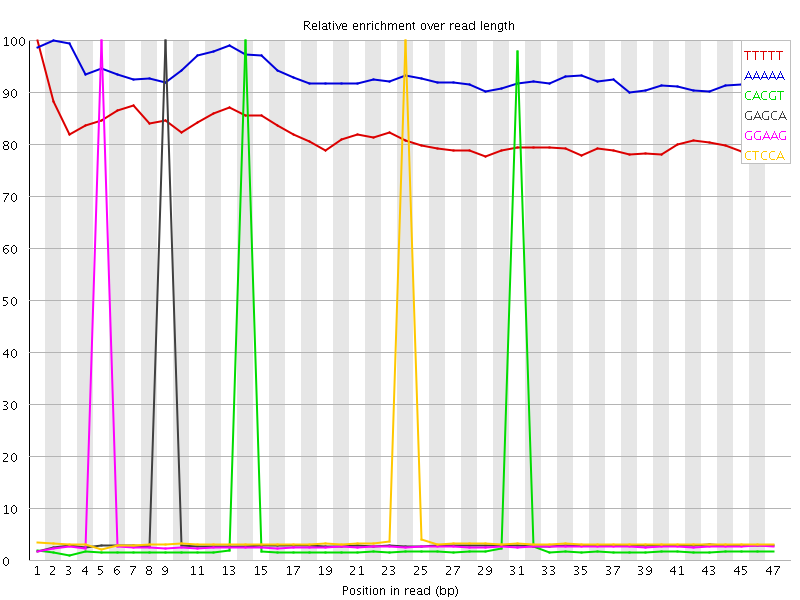
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 3606995 | 3.3680084 | 4.113982 | 1 |
| AAAAA | 3604525 | 3.2965405 | 3.5382955 | 2 |
| CACGT | 943985 | 3.1574132 | 53.574253 | 14 |
| GAGCA | 807855 | 2.77536 | 55.97389 | 9 |
| GGAAG | 770695 | 2.7308104 | 57.894814 | 5 |
| CTCCA | 841885 | 2.7302098 | 51.61545 | 24 |
| CGGAA | 769275 | 2.6428196 | 56.32474 | 4 |
| TCCAG | 782000 | 2.6156106 | 53.319977 | 25 |
| TGAAA | 1791655 | 2.5842493 | 24.247047 | 35 |
| GAAGA | 1089865 | 2.4587564 | 37.501366 | 6 |
| CTGAA | 1078025 | 2.3678381 | 35.943527 | 19 |
| TCGGA | 682135 | 2.3532047 | 56.14283 | 3 |
| TCTCG | 691120 | 2.321257 | 52.380486 | 43 |
| AGCAC | 689635 | 2.2971117 | 54.080296 | 10 |
| AGAGC | 663575 | 2.2796907 | 55.585125 | 8 |
| GCACA | 665190 | 2.2156878 | 53.8496 | 11 |
| CACAC | 674880 | 2.1795475 | 52.047676 | 12 |
| ATCGG | 621265 | 2.1432176 | 55.090775 | 2 |
| CCAGT | 626495 | 2.0954819 | 52.475925 | 26 |
| ACGTG | 599895 | 2.0694962 | 54.017437 | 32 |
| GAAAC | 940980 | 2.0582592 | 34.236073 | 36 |
| CTCGT | 608610 | 2.0441315 | 51.971127 | 44 |
| GTCAC | 607610 | 2.032316 | 52.796238 | 29 |
| GTCTG | 584995 | 2.0264926 | 55.026497 | 17 |
| GTGAA | 893925 | 2.0251038 | 36.0642 | 34 |
| CGTCT | 599410 | 2.0132318 | 53.33622 | 16 |
| TCTGA | 910455 | 2.008099 | 35.76545 | 18 |
| CGTGA | 571375 | 1.9711089 | 53.87411 | 33 |
| ACACG | 585015 | 1.948632 | 53.369442 | 13 |
| ACTCC | 600465 | 1.9472915 | 51.08366 | 23 |
| GATCG | 559160 | 1.9289699 | 54.267654 | 1 |
| TCACG | 576010 | 1.9266212 | 52.56071 | 30 |
| ACGTC | 570465 | 1.9080745 | 53.14582 | 15 |
| CAGTC | 564085 | 1.8867348 | 52.34157 | 27 |
| TGAAC | 841040 | 1.8473102 | 35.49324 | 20 |
| AAGAG | 818190 | 1.845852 | 36.89543 | 7 |
| CGATC | 550005 | 1.8396404 | 50.218002 | 40 |
| AAACG | 826060 | 1.8068883 | 33.95805 | 37 |
| AACTC | 789590 | 1.6815189 | 34.161858 | 22 |
| GAACT | 757690 | 1.6642354 | 35.24877 | 21 |
| AACGA | 746085 | 1.6319544 | 33.56402 | 38 |
| ATCTC | 757875 | 1.6206944 | 33.266632 | 42 |
| ACGAT | 705435 | 1.5494593 | 33.55546 | 39 |
| AGTCA | 695710 | 1.5280988 | 34.79288 | 28 |
| GATCT | 689595 | 1.5209702 | 34.0768 | 41 |
| TCGTA | 629890 | 1.3892848 | 34.425083 | 45 |
| GTATG | 563830 | 1.2826195 | 35.270107 | 47 |
| CGTAT | 578170 | 1.2752113 | 34.284485 | 46 |