![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW223_GTCCGC_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 6725563 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
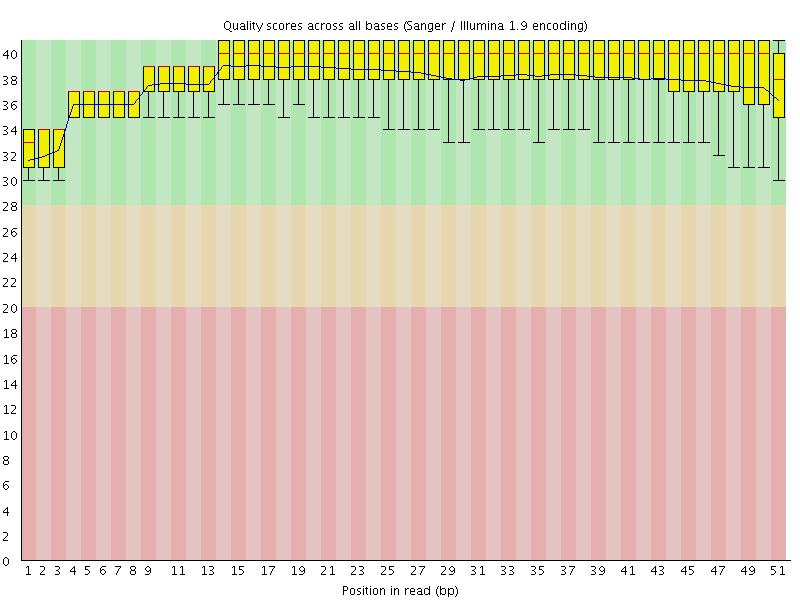
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
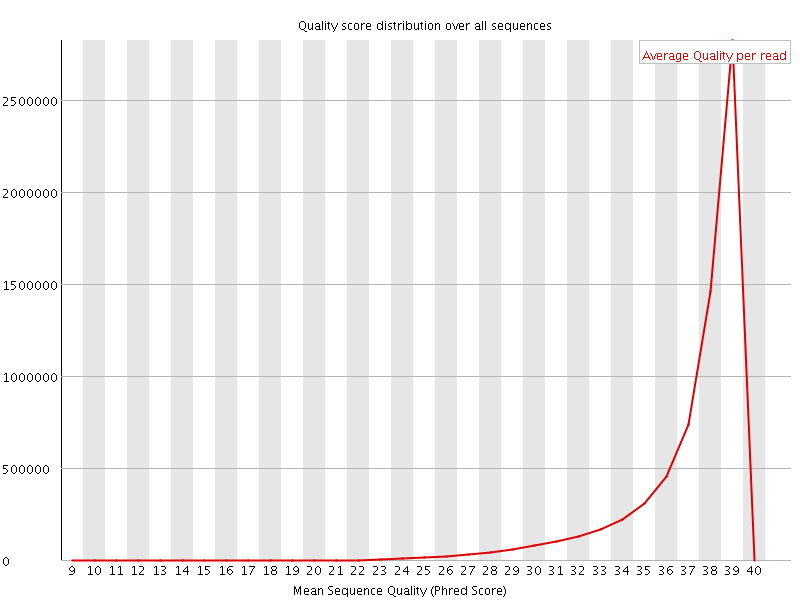
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
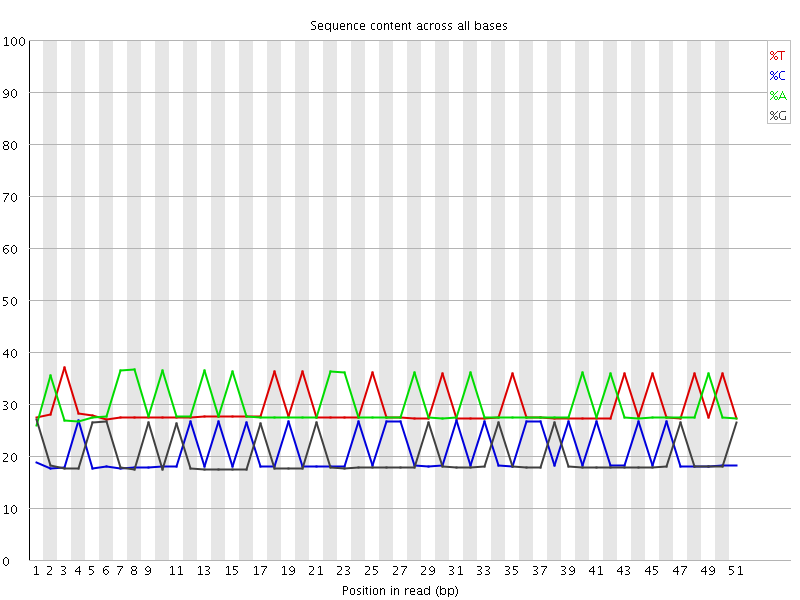
![[WARN]](Icons/warning.png) Per base GC content
Per base GC content
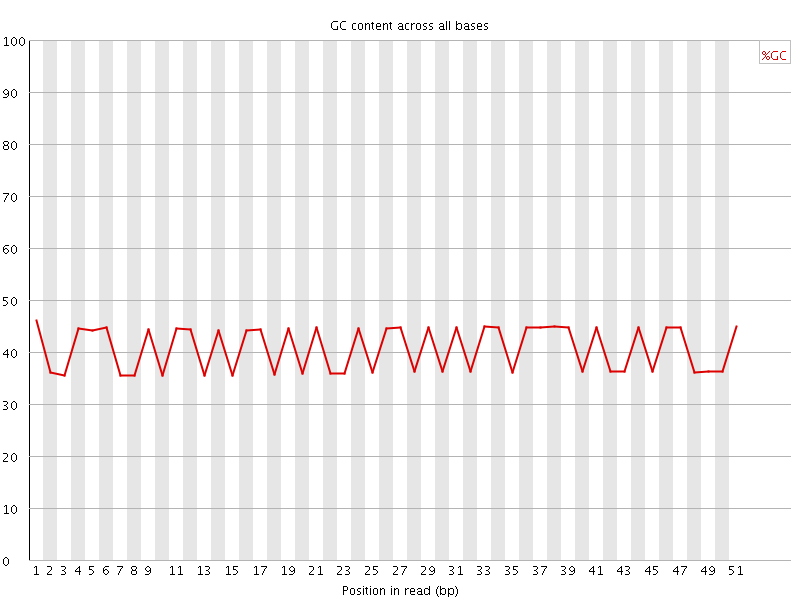
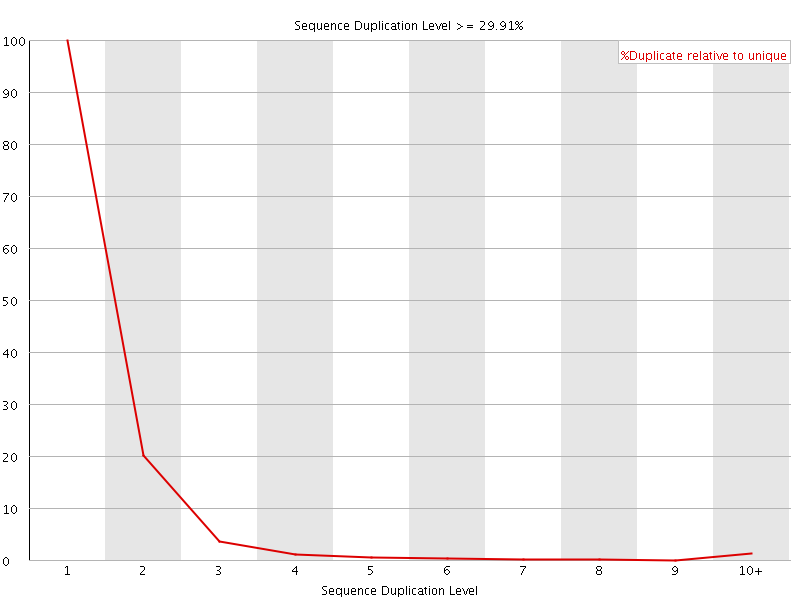
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
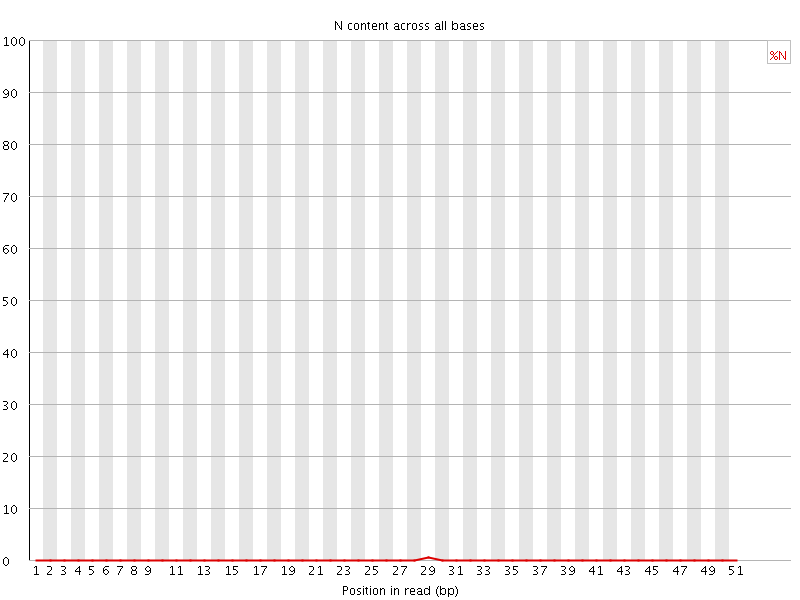
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
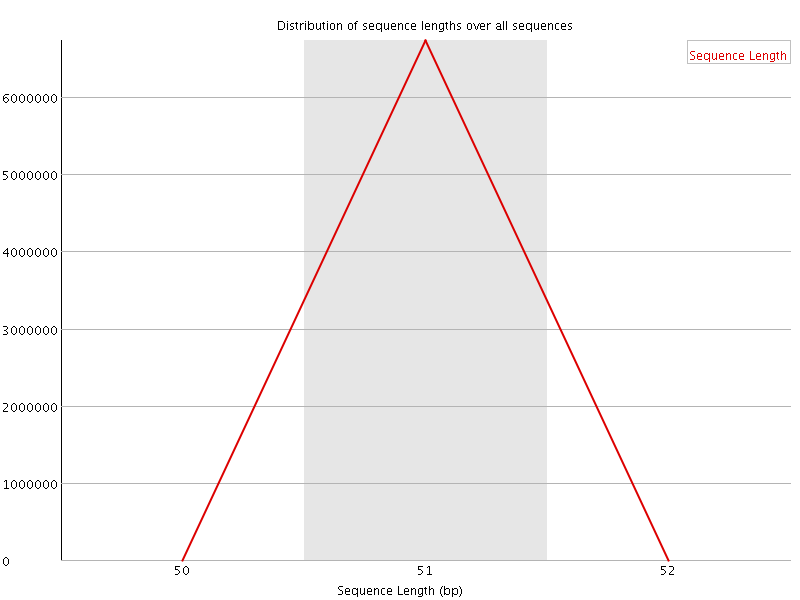
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGTCCGCACATCTCGTATG | 487960 | 7.255303385010296 | TruSeq Adapter, Index 1 (97% over 36bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
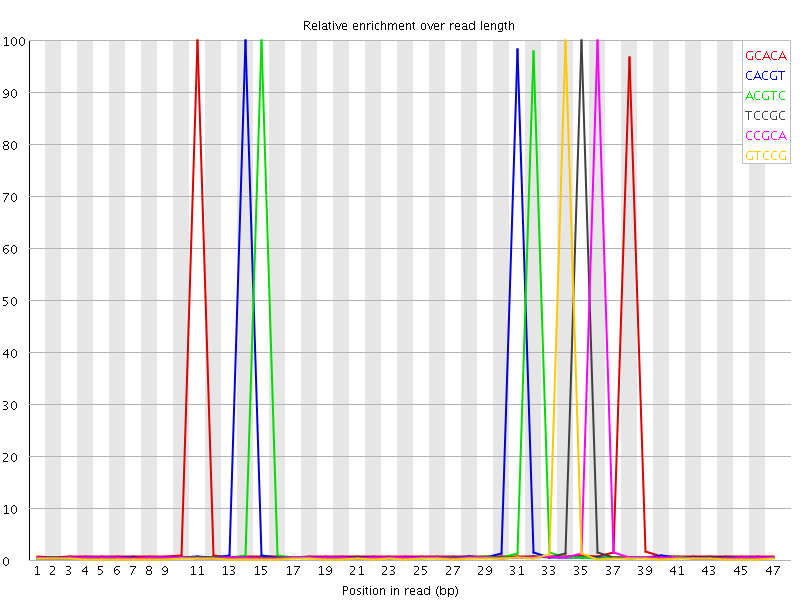
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GCACA | 1402825 | 5.73166 | 113.91549 | 11 |
| CACGT | 1359380 | 5.6376667 | 115.309 | 14 |
| ACGTC | 1341970 | 5.5654635 | 115.00791 | 15 |
| TCCGC | 744210 | 4.392114 | 158.10748 | 35 |
| CCGCA | 739315 | 4.2985897 | 156.3224 | 36 |
| GTCCG | 686020 | 4.2379556 | 165.96603 | 34 |
| CGCAC | 704570 | 4.0965724 | 156.03256 | 37 |
| CGTCC | 691665 | 4.0820084 | 158.83286 | 33 |
| TTTTT | 2794705 | 4.0786905 | 4.350996 | 2 |
| GGAAG | 903660 | 4.0454326 | 126.5805 | 5 |
| CGGAA | 900395 | 3.8508046 | 121.065125 | 4 |
| GAGCA | 893825 | 3.8227065 | 119.76279 | 9 |
| AAAAA | 2801695 | 3.7948756 | 4.020324 | 14 |
| TCCAG | 898565 | 3.7265592 | 114.0598 | 25 |
| CTCCA | 898745 | 3.5608487 | 108.25491 | 24 |
| TCGGA | 816895 | 3.5462246 | 122.40545 | 3 |
| TCTCG | 835625 | 3.5176415 | 115.70882 | 43 |
| GTCTG | 786435 | 3.465329 | 122.43281 | 17 |
| ATCGG | 792635 | 3.4409099 | 120.5413 | 2 |
| AGAGC | 803055 | 3.4345014 | 119.228584 | 8 |
| CCAGT | 804230 | 3.33533 | 113.402374 | 26 |
| AGCAC | 810750 | 3.312561 | 114.0047 | 10 |
| GAAGA | 1099670 | 3.304922 | 85.51058 | 6 |
| CGTCT | 780765 | 3.286703 | 116.59589 | 16 |
| GATCG | 753565 | 3.271303 | 119.423096 | 1 |
| CACAC | 837080 | 3.2674005 | 108.45242 | 12 |
| GTCAC | 787110 | 3.2643292 | 114.62164 | 29 |
| CTCGT | 770040 | 3.2415552 | 115.1185 | 44 |
| CTGAA | 1110550 | 3.2365108 | 81.93332 | 19 |
| TCACG | 765505 | 3.1747284 | 114.247154 | 30 |
| CAGTC | 747635 | 3.1006172 | 113.835434 | 27 |
| ACACG | 754510 | 3.0827758 | 113.62868 | 13 |
| ACTCC | 761375 | 3.0165856 | 108.22198 | 23 |
| TCTGA | 984995 | 2.9137652 | 82.836006 | 18 |
| AAGAG | 922805 | 2.7733767 | 84.66174 | 7 |
| TGAAC | 939395 | 2.7377083 | 81.15906 | 20 |
| GAACT | 878735 | 2.560925 | 80.91406 | 21 |
| CATCT | 885855 | 2.5034661 | 77.024025 | 41 |
| ATCTC | 883820 | 2.497715 | 77.43722 | 42 |
| AACTC | 886705 | 2.4687474 | 76.81365 | 22 |
| CACAT | 880775 | 2.4522371 | 75.52941 | 39 |
| AGTCA | 839910 | 2.4477763 | 80.555405 | 28 |
| ACATC | 848845 | 2.3633382 | 75.70396 | 40 |
| GTATG | 750235 | 2.3230543 | 84.26392 | 47 |
| TCGTA | 764050 | 2.2601762 | 80.84955 | 45 |
| CGTAT | 743855 | 2.2004366 | 80.68423 | 46 |