![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW221_CCGTCC_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 5170297 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 43 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
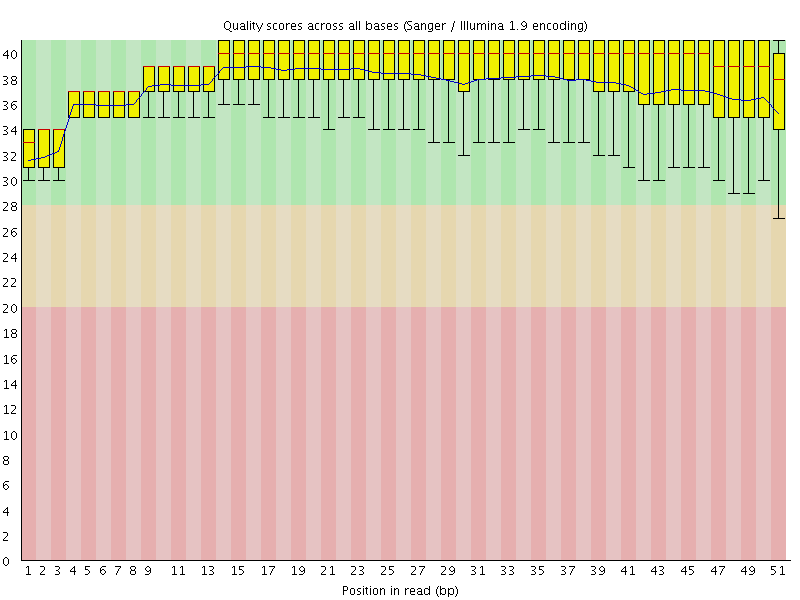
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
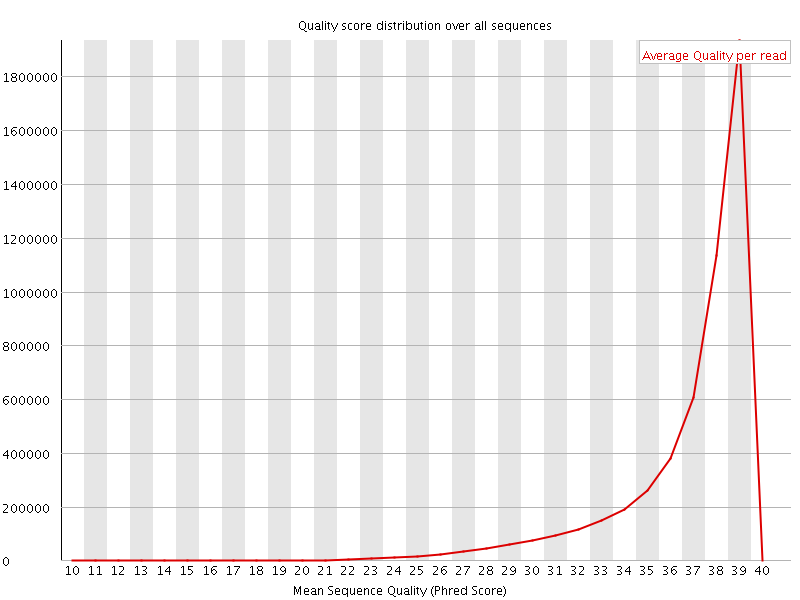
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
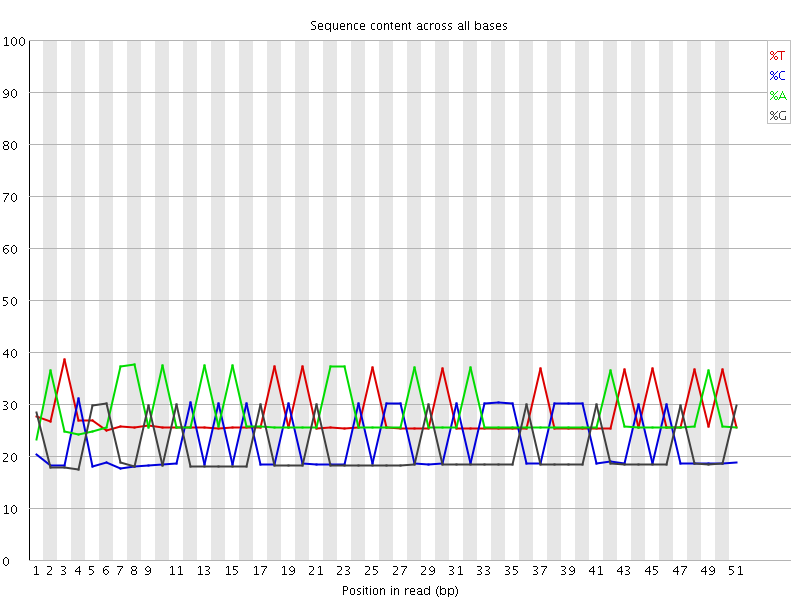
![[WARN]](Icons/warning.png) Per base GC content
Per base GC content
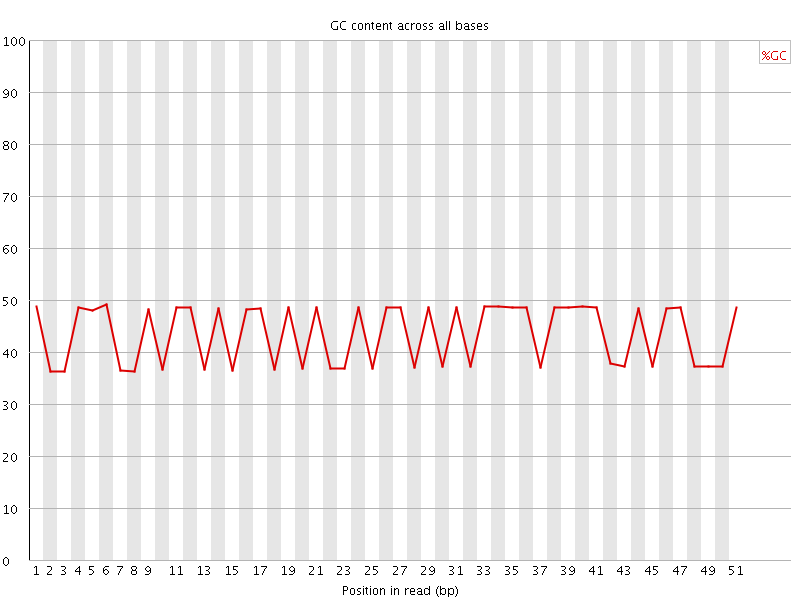
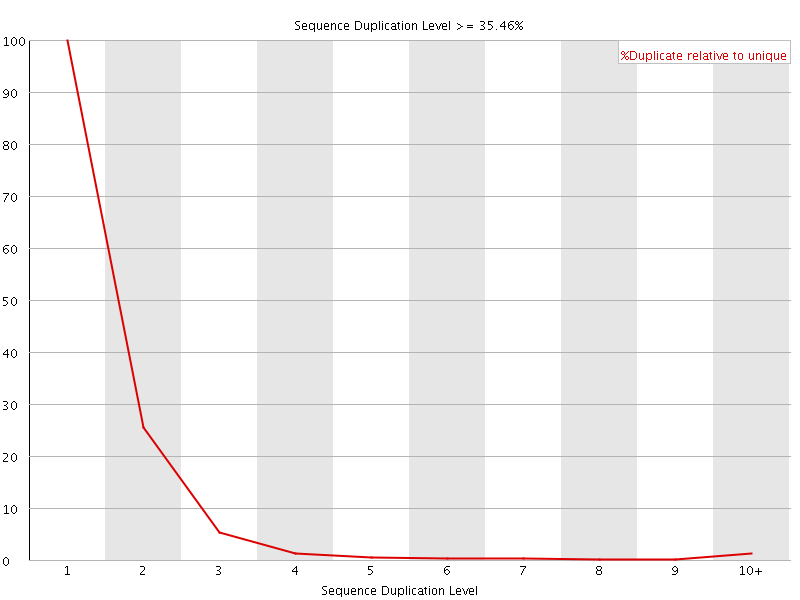
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
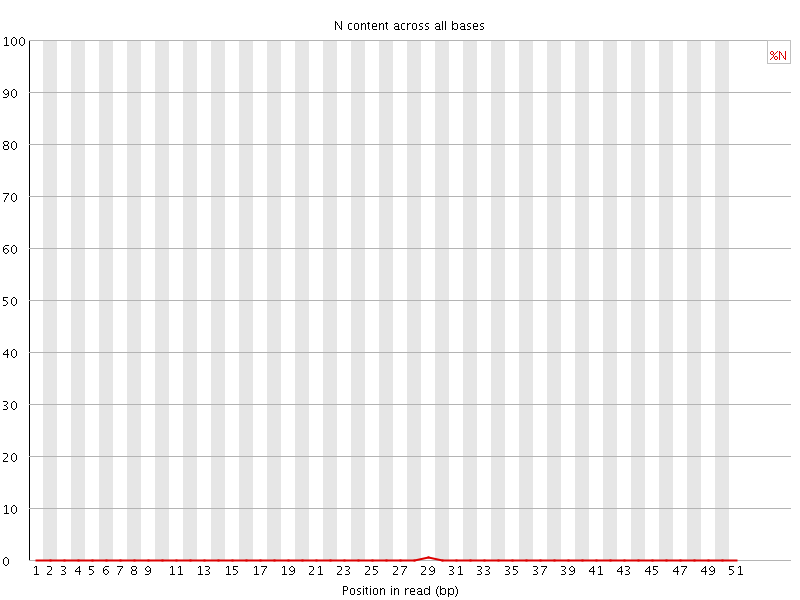
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
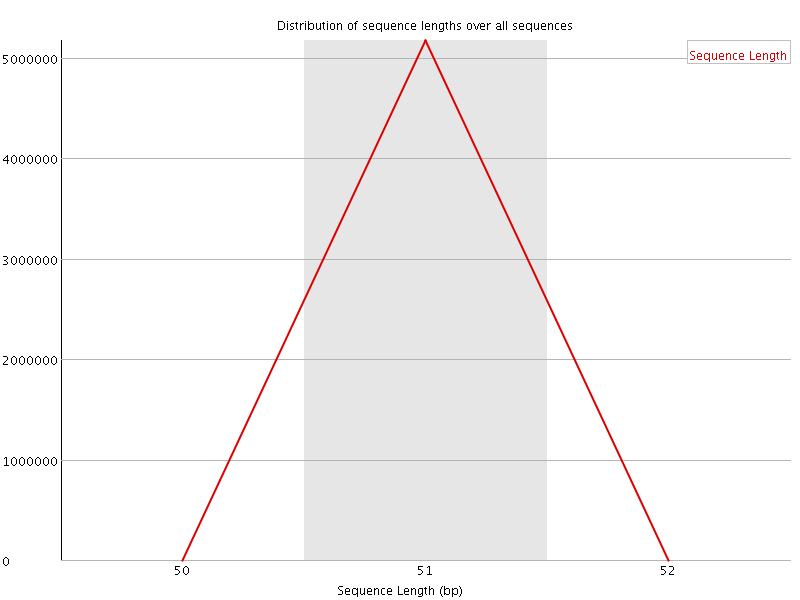
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCCGTCCCGATCTCGTATG | 475635 | 9.199374813477833 | TruSeq Adapter, Index 7 (97% over 36bp) |
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCCGTCCCGCTCTCGTATG | 10360 | 0.20037533627178475 | TruSeq Adapter, Index 7 (97% over 36bp) |
| GTTCGGAAGAGCACACGTCTGAACTCCAGTCACCCGTCCCGATCTCGTATG | 6651 | 0.12863864493664484 | TruSeq Adapter, Index 2 (97% over 34bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
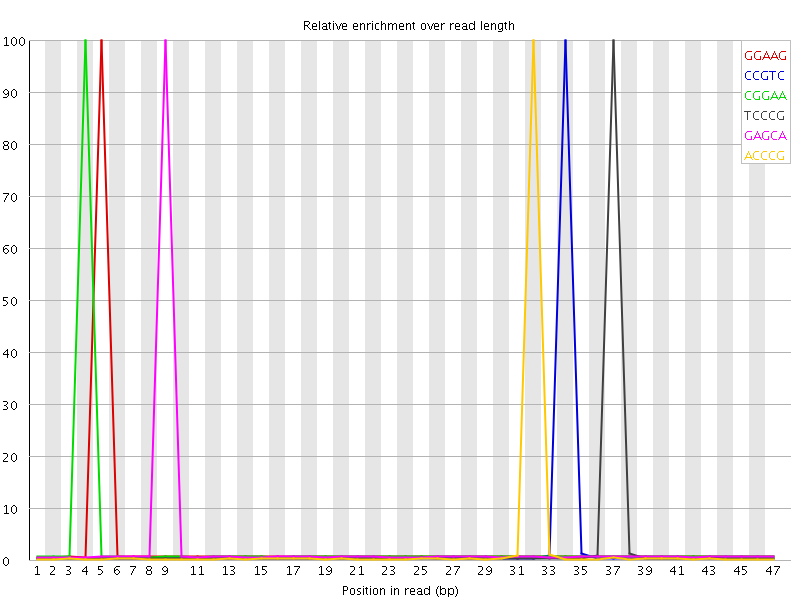
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGAAG | 849930 | 4.635663 | 157.61714 | 5 |
| CCGTC | 725890 | 4.396942 | 168.18602 | 34 |
| CGGAA | 852395 | 4.3473387 | 148.24825 | 4 |
| TCCCG | 713965 | 4.324709 | 165.35805 | 37 |
| GAGCA | 835160 | 4.259437 | 146.86069 | 9 |
| ACCCG | 711095 | 4.254722 | 167.89746 | 32 |
| TTTTT | 1770600 | 4.2517858 | 5.2224197 | 2 |
| CCCGT | 701510 | 4.249265 | 168.70135 | 33 |
| TCCAG | 869915 | 4.200003 | 137.16167 | 25 |
| CGTCC | 688415 | 4.1699443 | 166.98465 | 35 |
| GTCTG | 769955 | 4.024582 | 149.56287 | 17 |
| CCCGA | 672235 | 4.022209 | 156.1738 | 38 |
| GTCCC | 663910 | 4.02151 | 166.06627 | 36 |
| ATCGG | 778795 | 4.0210743 | 147.25307 | 2 |
| CACCC | 715835 | 4.005072 | 157.25146 | 31 |
| TCGGA | 772730 | 3.9897594 | 149.54405 | 3 |
| AAAAA | 1763250 | 3.9818296 | 4.2240825 | 13 |
| GAAGA | 971625 | 3.9497907 | 117.92391 | 6 |
| GATCG | 759865 | 3.9233353 | 145.76192 | 1 |
| AGAGC | 766885 | 3.9112248 | 146.54695 | 8 |
| CCAGT | 798465 | 3.8550382 | 136.18881 | 26 |
| CTGAA | 991745 | 3.8165033 | 110.9956 | 19 |
| TCACC | 842100 | 3.8018093 | 127.52778 | 30 |
| CTCCA | 832980 | 3.7606354 | 128.15189 | 24 |
| AGCAC | 783610 | 3.7371142 | 137.3312 | 10 |
| GCACA | 779920 | 3.7195165 | 137.18788 | 11 |
| TCTCG | 758850 | 3.7090707 | 132.00851 | 43 |
| CGTCT | 757435 | 3.702155 | 139.76846 | 16 |
| GTCAC | 755625 | 3.648204 | 136.49715 | 29 |
| CACGT | 741885 | 3.5818667 | 138.31041 | 14 |
| ACGTC | 738260 | 3.5643647 | 138.25426 | 15 |
| CCGAT | 734295 | 3.5452216 | 125.51653 | 39 |
| CAGTC | 729690 | 3.5229883 | 135.9744 | 27 |
| CGATC | 722915 | 3.4902782 | 125.54836 | 40 |
| ACACG | 728245 | 3.4730732 | 136.75008 | 13 |
| CTCGT | 708940 | 3.4651234 | 131.00159 | 44 |
| TCTGA | 882685 | 3.4388068 | 112.01653 | 18 |
| CACAC | 768125 | 3.4254858 | 128.00569 | 12 |
| AAGAG | 838240 | 3.4075623 | 117.258675 | 7 |
| ACTCC | 738065 | 3.3321254 | 128.38824 | 23 |
| TGAAC | 864445 | 3.3266187 | 110.44723 | 20 |
| GAACT | 828730 | 3.1891778 | 110.253845 | 21 |
| ATTTT | 1266355 | 3.0037925 | 3.4866662 | 1 |
| AGTCA | 776245 | 2.9872012 | 108.7304 | 28 |
| AACTC | 826435 | 2.9739127 | 102.938866 | 22 |
| GATCT | 756180 | 2.945963 | 101.65083 | 41 |
| GTATG | 680440 | 2.8349023 | 109.078094 | 47 |
| ATCTC | 777445 | 2.832211 | 95.17083 | 42 |
| TCGTA | 695970 | 2.7113934 | 102.79783 | 45 |
| CGTAT | 686665 | 2.675143 | 102.58612 | 46 |
| CCCGC | 145370 | 1.091253 | 7.487457 | 38 |
| CCGCT | 165265 | 1.0010617 | 5.9359083 | 39 |
| CGCTC | 141510 | 0.8571702 | 5.83621 | 40 |