![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW220_ATGTCA_L003_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 6968942 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
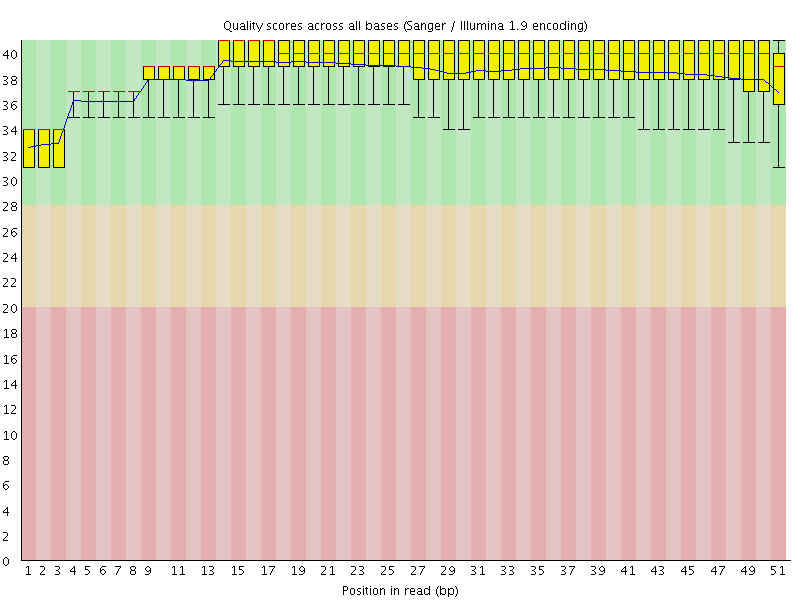
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
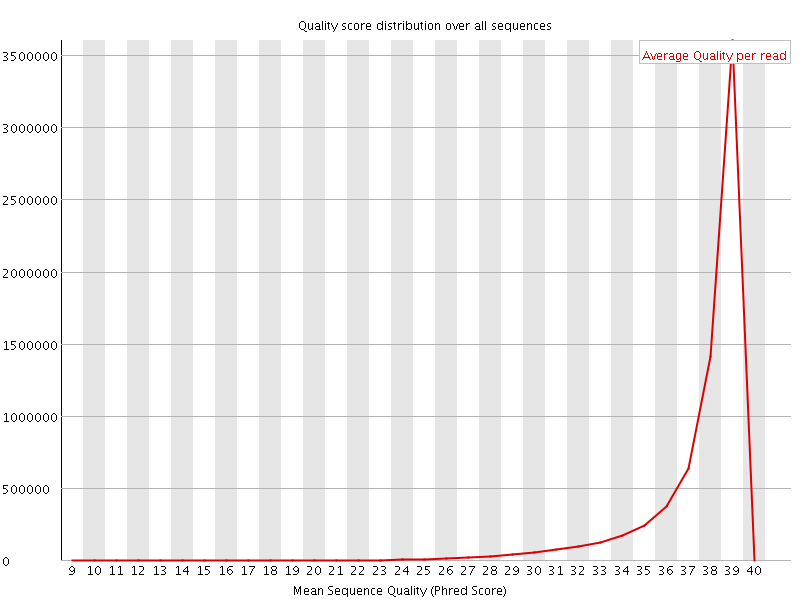
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
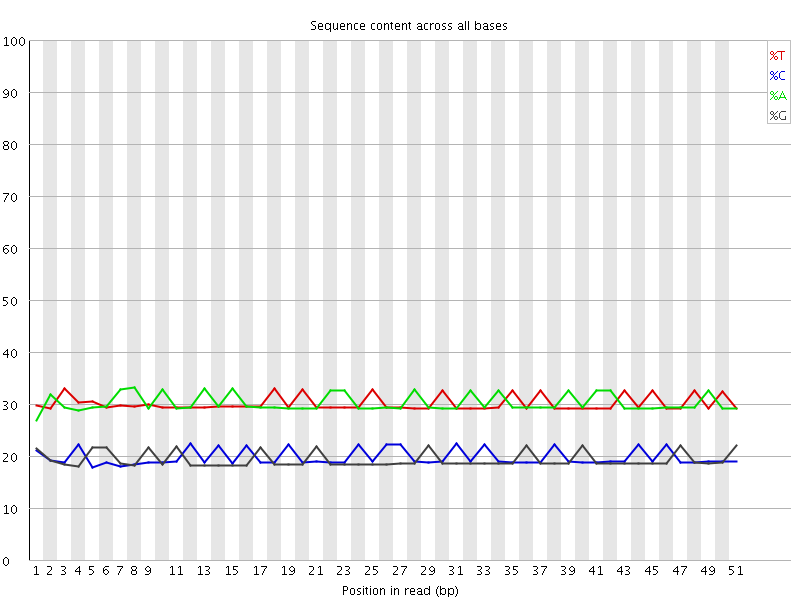
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
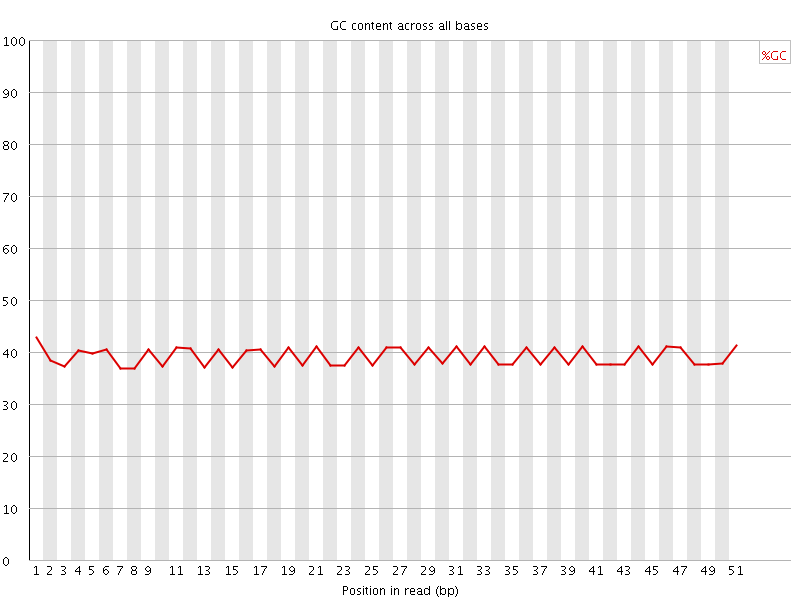
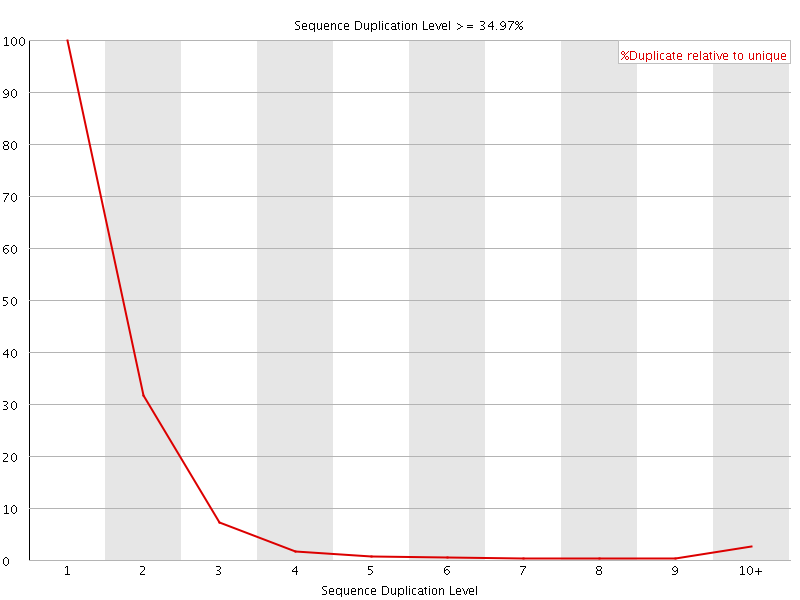
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
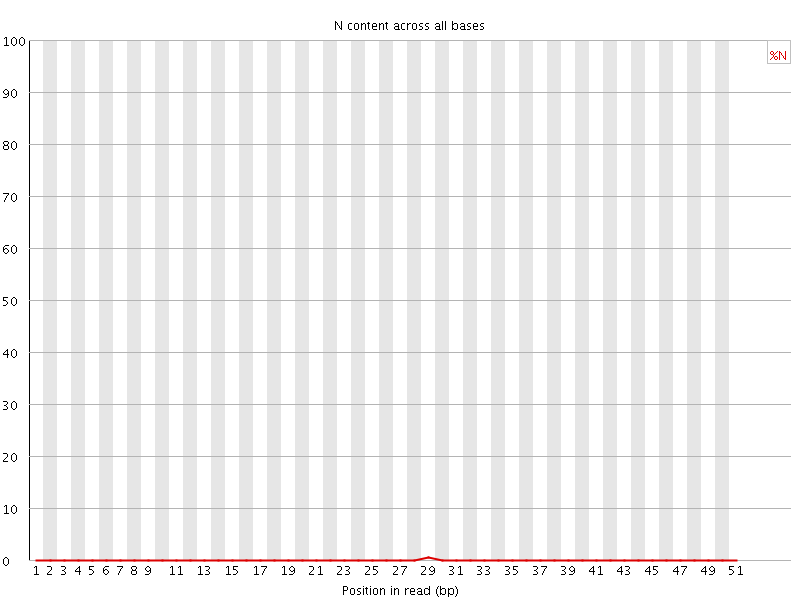
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
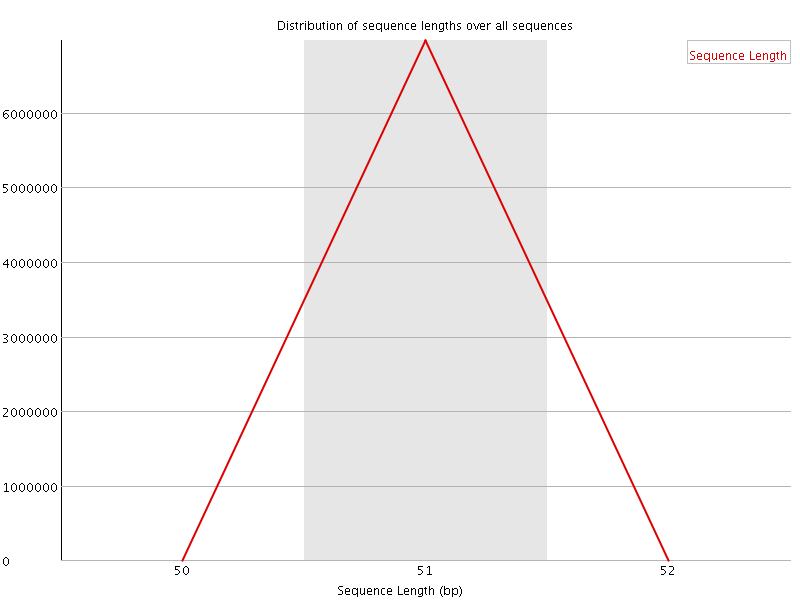
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACATGTCAGAATCTCGTATG | 206779 | 2.9671505373412494 | TruSeq Adapter, Index 1 (97% over 35bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
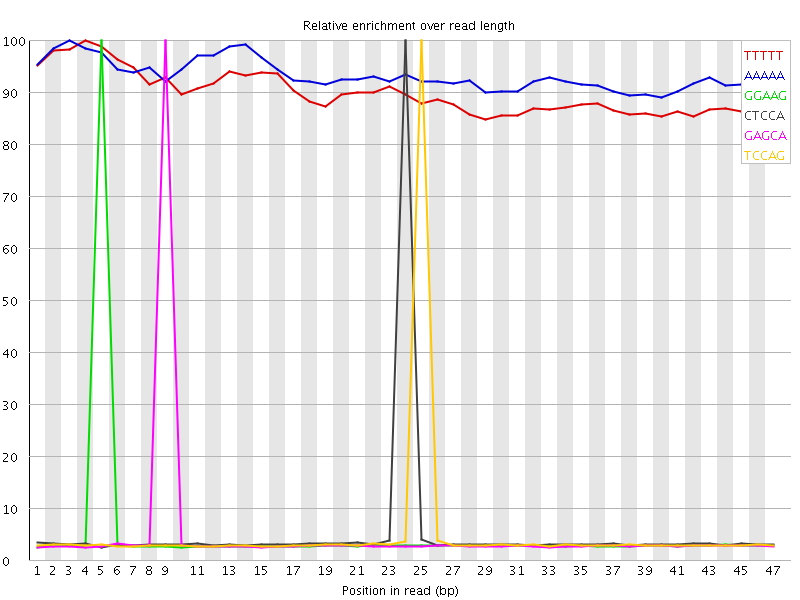
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 3007545 | 3.6153114 | 4.023951 | 4 |
| AAAAA | 2983205 | 3.5065615 | 3.7603533 | 3 |
| GGAAG | 589460 | 2.6547909 | 53.49294 | 5 |
| CTCCA | 607415 | 2.564248 | 48.498676 | 24 |
| GAGCA | 580330 | 2.554082 | 51.823895 | 9 |
| TCCAG | 587415 | 2.5376735 | 49.75191 | 25 |
| CGGAA | 568865 | 2.5036237 | 52.129726 | 4 |
| CTGAA | 820200 | 2.3172207 | 33.622032 | 19 |
| GAAGA | 801960 | 2.3081791 | 34.77771 | 6 |
| CAGAA | 791065 | 2.2249117 | 32.15161 | 38 |
| TCTCG | 505115 | 2.1919367 | 49.129234 | 43 |
| TCGGA | 489995 | 2.1662 | 52.0443 | 3 |
| AGAGC | 481925 | 2.1209931 | 51.34035 | 8 |
| GCACA | 488460 | 2.1007419 | 50.001225 | 11 |
| AGCAC | 487220 | 2.095409 | 50.000217 | 10 |
| CACAC | 497450 | 2.0906286 | 48.536633 | 12 |
| CCAGT | 476640 | 2.0591178 | 49.116722 | 26 |
| ATCGG | 458900 | 2.0287333 | 51.790028 | 2 |
| GTCTG | 444765 | 1.9750792 | 51.23913 | 17 |
| GTCAG | 444355 | 1.9644318 | 49.30167 | 36 |
| GTCAC | 451845 | 1.9520015 | 49.08911 | 29 |
| TCTGA | 685125 | 1.9443057 | 33.45726 | 18 |
| TCAGA | 683465 | 1.9309183 | 32.13499 | 37 |
| CGTCT | 439505 | 1.9072235 | 49.797787 | 16 |
| CTCGT | 439035 | 1.9051839 | 48.59719 | 44 |
| CACGT | 435395 | 1.8809365 | 49.660988 | 14 |
| ACTCC | 444935 | 1.8783267 | 48.00668 | 23 |
| GATCG | 416660 | 1.841996 | 51.360497 | 1 |
| CAGTC | 417550 | 1.8038447 | 48.842014 | 27 |
| ACACG | 418840 | 1.8013238 | 49.47519 | 13 |
| ACGTC | 412685 | 1.7828276 | 49.53236 | 15 |
| TGAAC | 625945 | 1.7684134 | 33.004368 | 20 |
| AAGAG | 609470 | 1.7541598 | 34.133102 | 7 |
| TCACA | 623950 | 1.7225875 | 31.71716 | 30 |
| TGTCA | 578730 | 1.6423689 | 32.036816 | 35 |
| AACTC | 583745 | 1.6115903 | 31.999853 | 22 |
| GAATC | 569955 | 1.610231 | 31.823765 | 40 |
| GAACT | 562200 | 1.5883217 | 32.818596 | 21 |
| ATCTC | 567150 | 1.5728106 | 31.431057 | 42 |
| CACAT | 565750 | 1.5619103 | 31.33347 | 31 |
| AGTCA | 522965 | 1.4774754 | 32.275043 | 28 |
| CATGT | 507070 | 1.4390061 | 32.053387 | 33 |
| ACATG | 507450 | 1.4336426 | 31.968376 | 32 |
| ATGTC | 502045 | 1.4247457 | 31.867987 | 34 |
| AGAAT | 766330 | 1.4158604 | 21.210466 | 39 |
| AATCT | 708615 | 1.2851263 | 20.710442 | 41 |
| TCGTA | 447750 | 1.2706628 | 31.726702 | 45 |
| GTATG | 418105 | 1.2142168 | 32.19638 | 47 |
| CGTAT | 416335 | 1.1815107 | 31.571814 | 46 |