![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW219_AGTTCC_L003_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 6890497 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 46 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
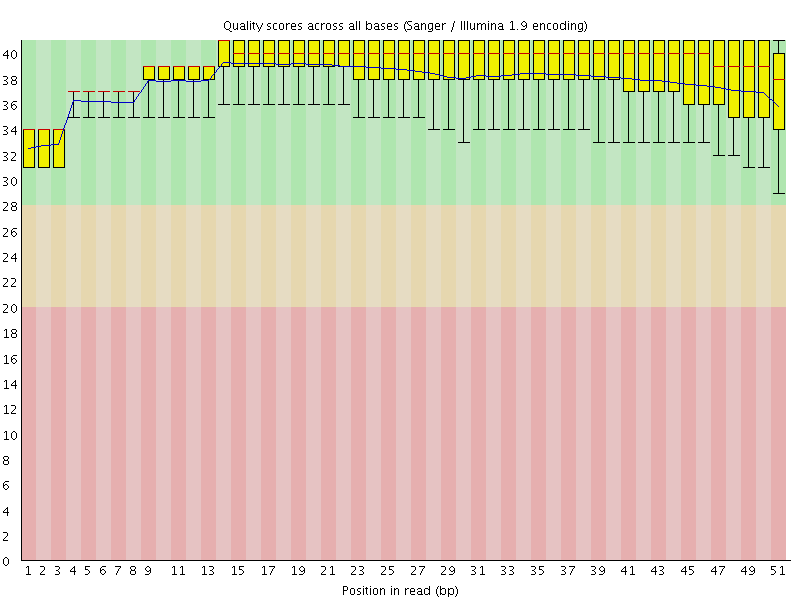
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
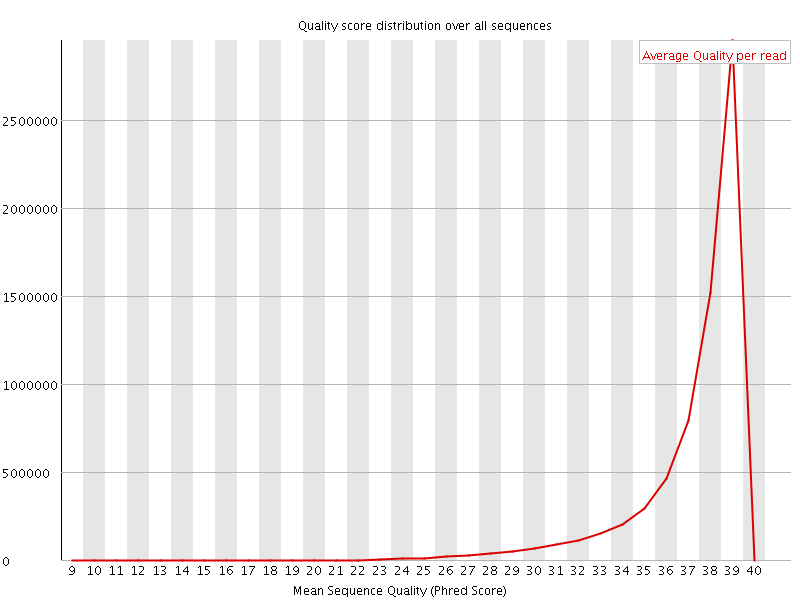
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
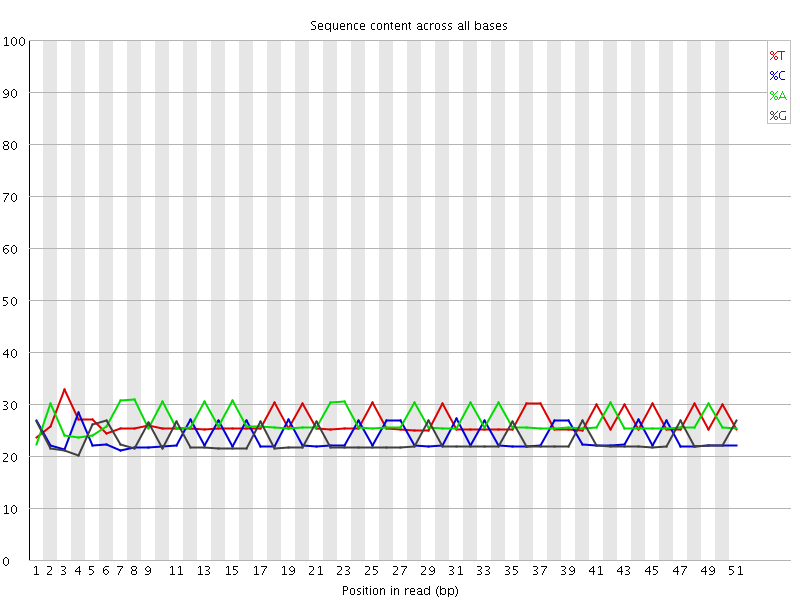
![[WARN]](Icons/warning.png) Per base GC content
Per base GC content
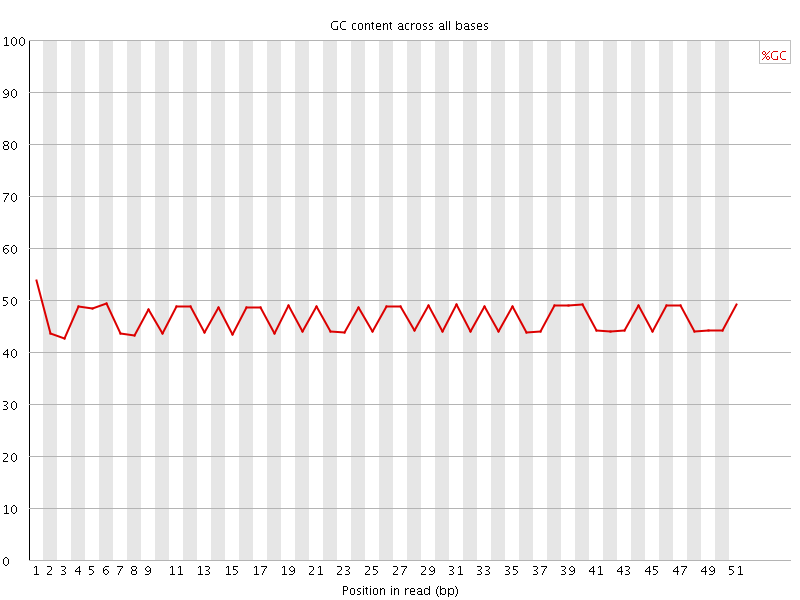
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
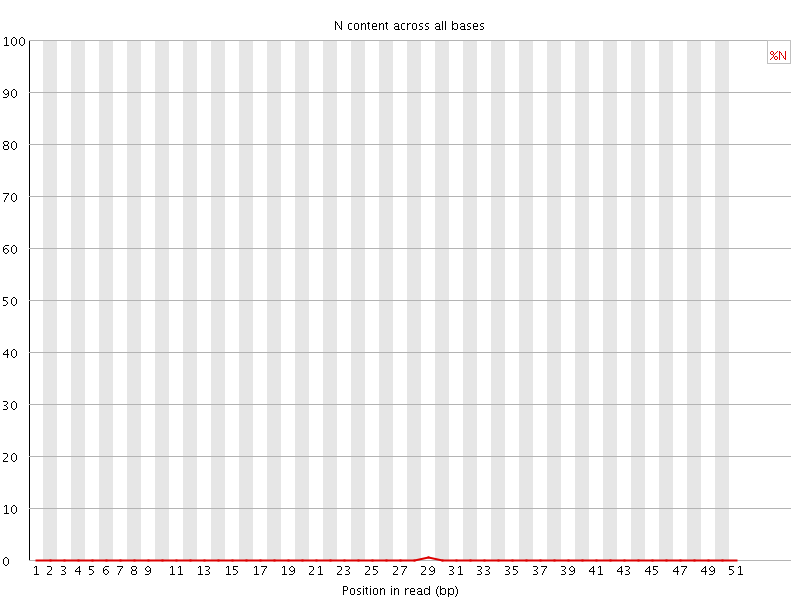
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
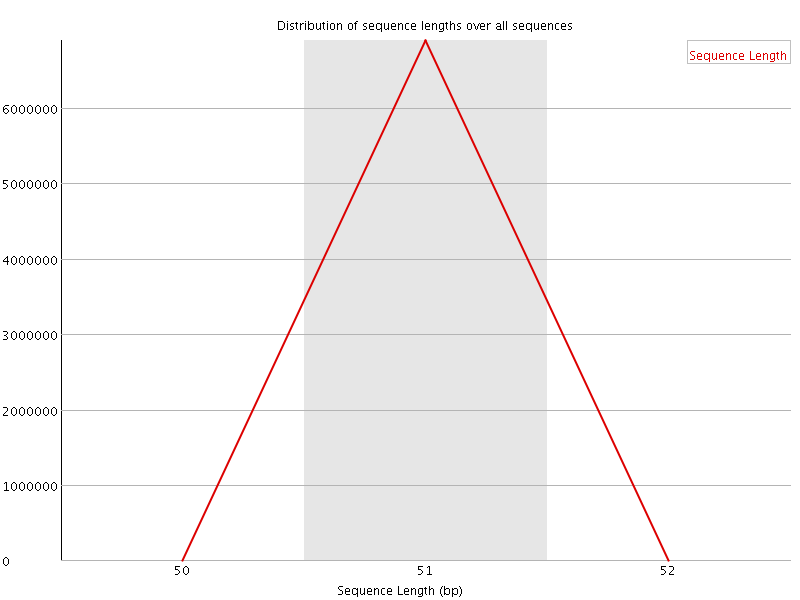
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
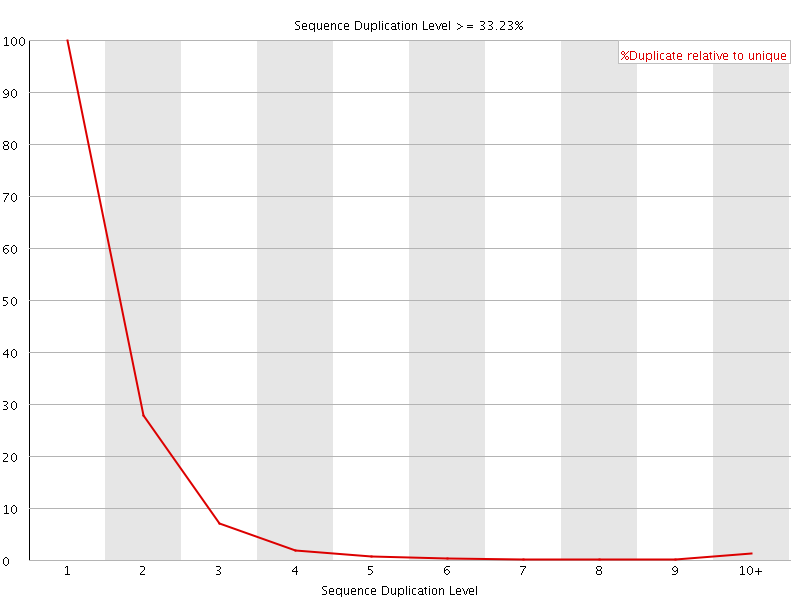
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACAGTTCCGTATCTCGTATG | 309667 | 4.4941170426458354 | TruSeq Adapter, Index 8 (97% over 37bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
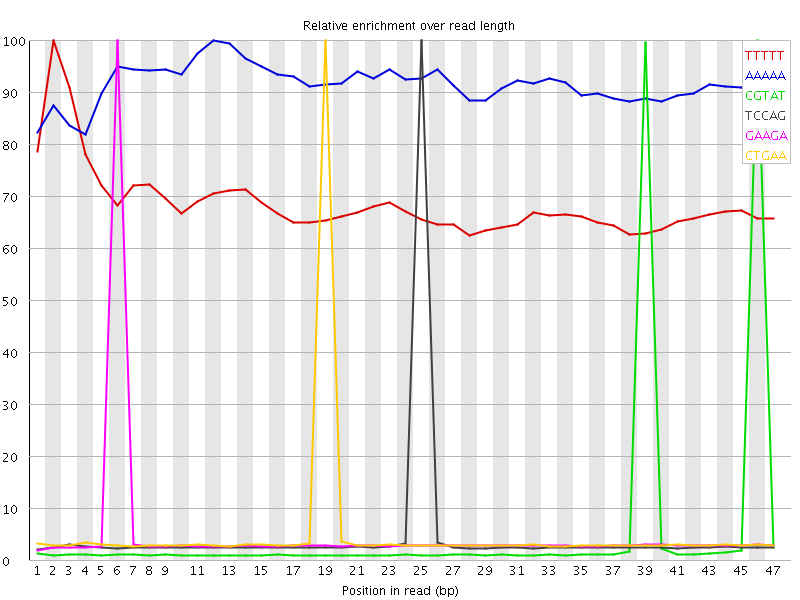
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 1663975 | 3.852325 | 5.6200457 | 2 |
| AAAAA | 1706320 | 3.8435473 | 4.197841 | 12 |
| CGTAT | 871295 | 2.6146784 | 47.82584 | 46 |
| TCCAG | 772245 | 2.6119618 | 55.70075 | 25 |
| GAAGA | 841045 | 2.5612447 | 51.482037 | 6 |
| CTGAA | 857935 | 2.5605106 | 49.983288 | 19 |
| GGAAG | 709935 | 2.5137982 | 59.498253 | 5 |
| CGGAA | 696315 | 2.4031289 | 58.025448 | 4 |
| GAGCA | 687585 | 2.373 | 57.875553 | 9 |
| TTCCG | 684900 | 2.3292694 | 54.478786 | 36 |
| CCAGT | 684370 | 2.3147428 | 55.330593 | 26 |
| CAGTT | 759975 | 2.280617 | 48.77742 | 33 |
| GATCG | 653690 | 2.268423 | 57.896248 | 1 |
| ATCGG | 640265 | 2.2218356 | 58.1269 | 2 |
| AGCAC | 646510 | 2.1747336 | 55.90676 | 10 |
| CTCCA | 658970 | 2.1723857 | 53.936882 | 24 |
| TGAAC | 714545 | 2.1325624 | 49.553444 | 20 |
| GTCTG | 601990 | 2.1004984 | 57.512367 | 17 |
| TCTGA | 688535 | 2.066232 | 49.7944 | 18 |
| GTTCC | 603505 | 2.052454 | 54.28831 | 35 |
| GCACA | 609425 | 2.049987 | 55.73145 | 11 |
| TCTCG | 590795 | 2.0092287 | 54.875748 | 43 |
| GTCAC | 588695 | 1.9911413 | 55.039574 | 29 |
| CACAG | 591795 | 1.9906831 | 54.25091 | 31 |
| AGAGC | 570885 | 1.9702438 | 57.195763 | 8 |
| AAGAG | 646225 | 1.9679569 | 50.74707 | 7 |
| CGTCT | 573205 | 1.9494071 | 55.91105 | 16 |
| GAACT | 644570 | 1.9237217 | 49.283665 | 21 |
| AGTTC | 639505 | 1.9190973 | 48.207733 | 34 |
| TCGGA | 552760 | 1.9181776 | 57.87924 | 3 |
| ACGTC | 566615 | 1.9164602 | 55.699074 | 15 |
| CACGT | 561615 | 1.8995486 | 55.70748 | 14 |
| CACAC | 579175 | 1.8988914 | 54.19839 | 12 |
| ATCTC | 642640 | 1.8796644 | 47.083473 | 42 |
| CAGTC | 548735 | 1.8559847 | 54.88128 | 27 |
| TCCGT | 543285 | 1.8476526 | 54.176857 | 37 |
| TCACA | 628840 | 1.8292451 | 47.12746 | 30 |
| AACTC | 624435 | 1.8164313 | 47.850773 | 22 |
| ACAGT | 606325 | 1.8095794 | 48.09066 | 32 |
| CTCGT | 531445 | 1.807386 | 54.51073 | 44 |
| ACACG | 526140 | 1.7698324 | 55.39186 | 13 |
| CCGTA | 522730 | 1.7680281 | 53.577755 | 38 |
| ACTCC | 529920 | 1.7469542 | 53.80421 | 23 |
| AGTCA | 582360 | 1.7380557 | 48.43796 | 28 |
| GTATC | 550480 | 1.6519413 | 47.854637 | 40 |
| GTATG | 517400 | 1.5930153 | 48.922764 | 47 |
| TCGTA | 527120 | 1.5818402 | 47.930664 | 45 |
| TATCT | 592095 | 1.5365449 | 41.558052 | 41 |