![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW218_AGTCAA_L003_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 5380642 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
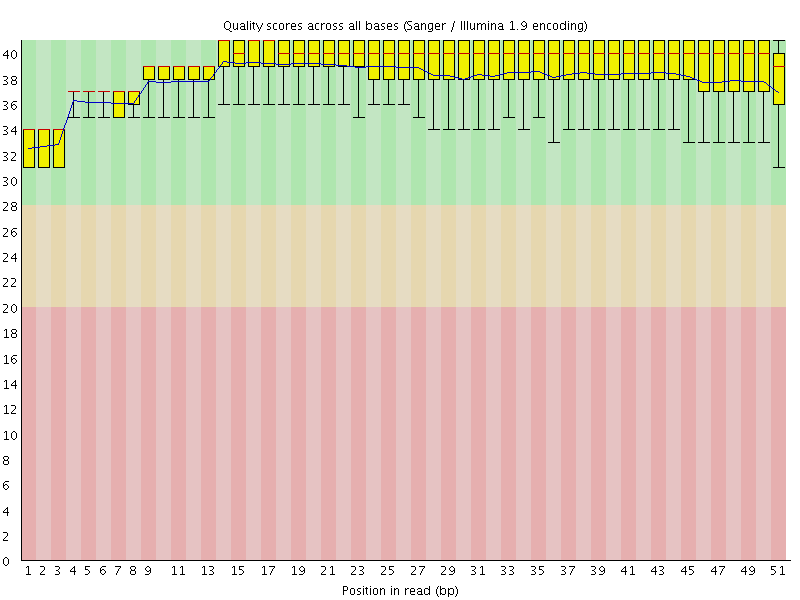
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
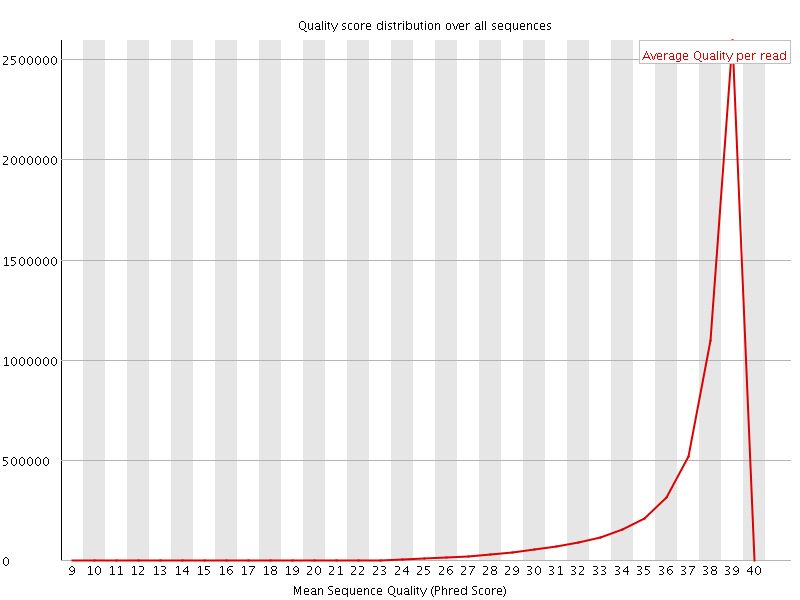
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
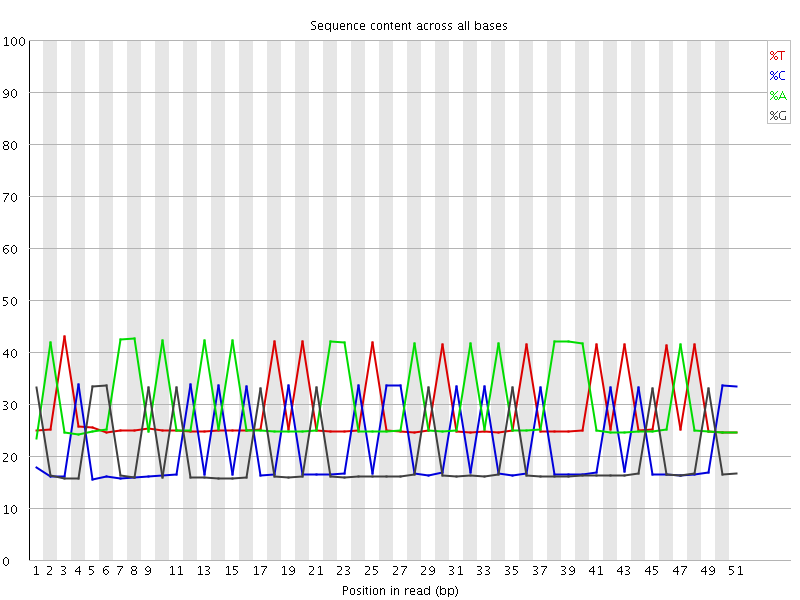
![[WARN]](Icons/warning.png) Per base GC content
Per base GC content
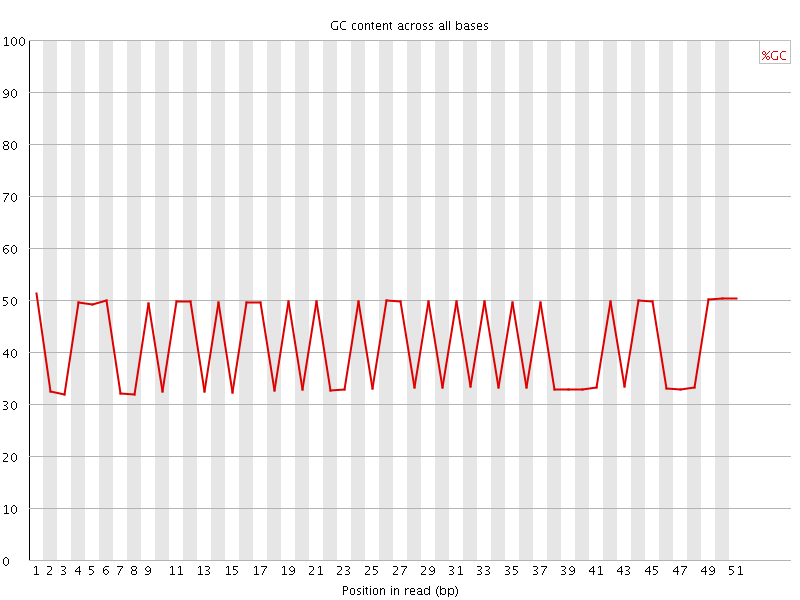
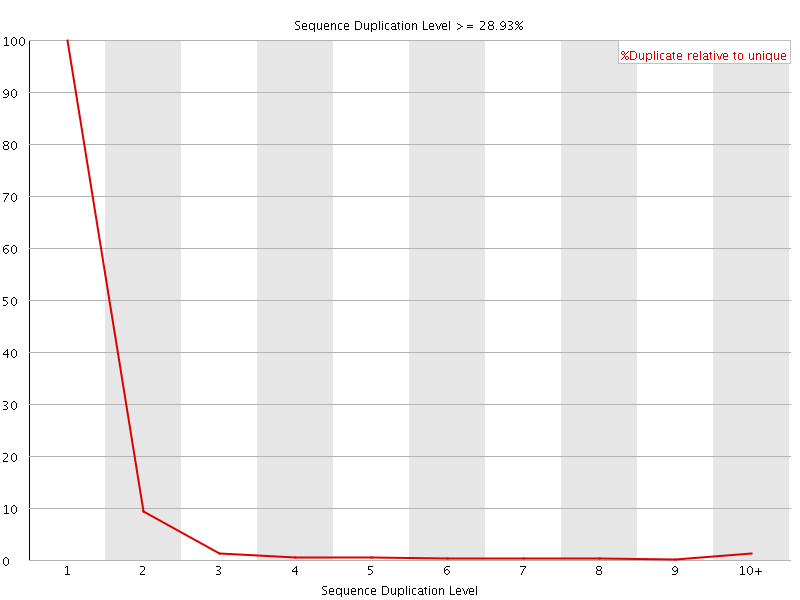
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
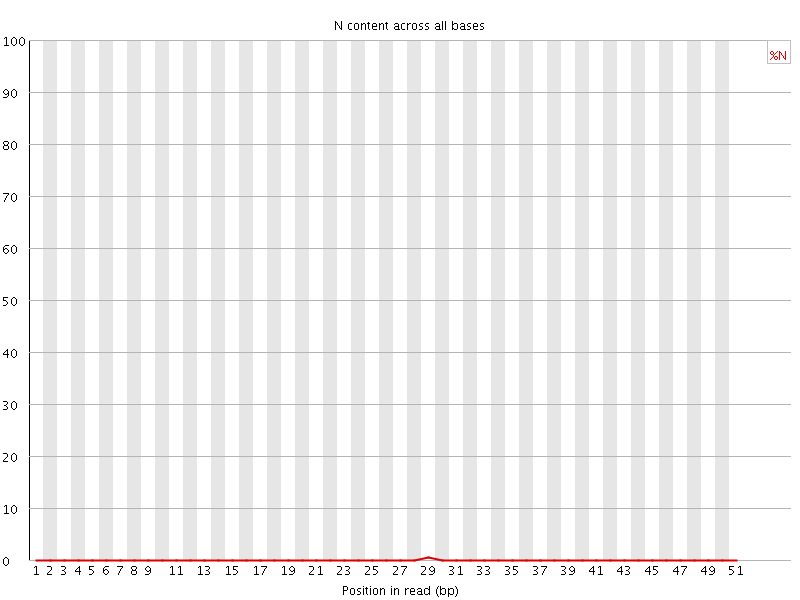
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
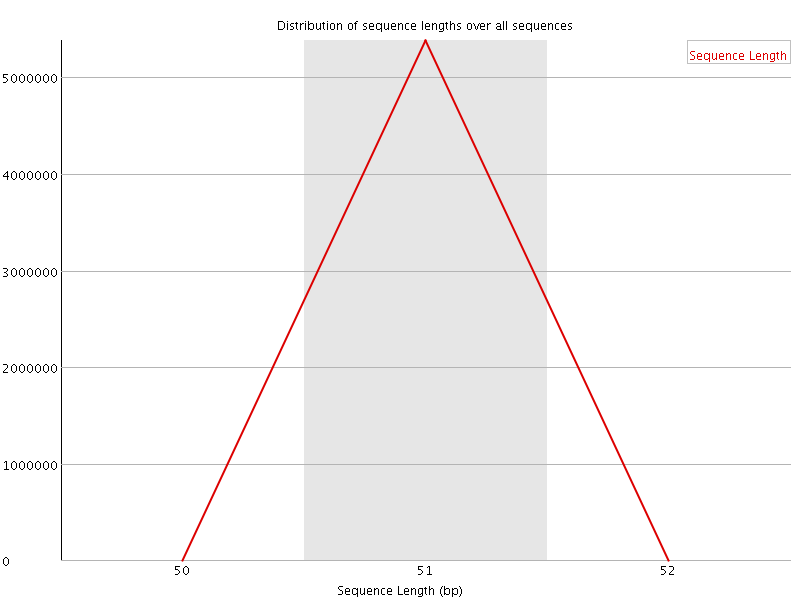
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACAGTCAAATCTCGTATGCC | 839900 | 15.609661449321473 | TruSeq Adapter, Index 8 (97% over 36bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
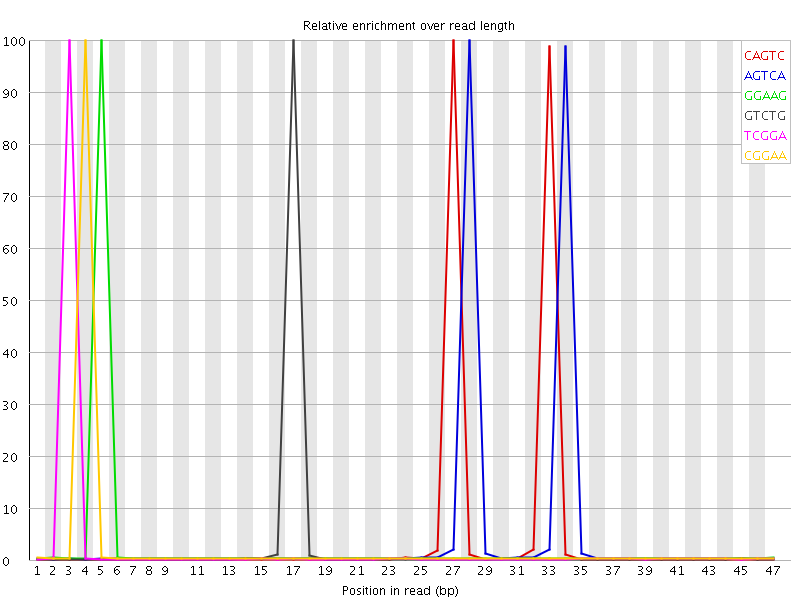
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CAGTC | 1971945 | 9.807588 | 212.029 | 27 |
| AGTCA | 2031650 | 7.2793884 | 152.61446 | 28 |
| GGAAG | 1162190 | 6.398566 | 242.40463 | 5 |
| GTCTG | 1071915 | 6.1012473 | 246.349 | 17 |
| TCGGA | 1101155 | 5.925962 | 237.22328 | 3 |
| CGGAA | 1160490 | 5.904783 | 224.44383 | 4 |
| GAGCA | 1158085 | 5.8925457 | 222.96979 | 9 |
| TCTCG | 1116510 | 5.873243 | 223.24428 | 41 |
| ATCGG | 1086145 | 5.845184 | 237.1045 | 2 |
| TCCAG | 1155760 | 5.7482424 | 213.18939 | 25 |
| GATCG | 1057900 | 5.693181 | 236.46698 | 1 |
| CGTCT | 1074385 | 5.651649 | 227.8833 | 16 |
| CTCGT | 1067200 | 5.6138535 | 222.08456 | 42 |
| AGAGC | 1094060 | 5.566775 | 222.66399 | 8 |
| CCAGT | 1086280 | 5.40268 | 212.42052 | 26 |
| CTCCA | 1170555 | 5.380421 | 197.2602 | 24 |
| GTCAC | 1072110 | 5.3322043 | 212.26886 | 29 |
| CACGT | 1066700 | 5.305297 | 215.8803 | 14 |
| ATGCC | 1061145 | 5.277669 | 209.81055 | 47 |
| ACGTC | 1054020 | 5.2422323 | 215.62482 | 15 |
| GCACA | 1104285 | 5.19279 | 205.28284 | 11 |
| CACAG | 1101545 | 5.179905 | 199.83614 | 31 |
| AGCAC | 1098980 | 5.167844 | 205.62286 | 10 |
| ACACG | 1054585 | 4.95908 | 204.32898 | 13 |
| ACTCC | 1068075 | 4.909375 | 197.79955 | 23 |
| CACAC | 1114465 | 4.8433175 | 189.4236 | 12 |
| GAAGA | 1307935 | 4.794323 | 161.5724 | 6 |
| CTGAA | 1311660 | 4.6996694 | 155.72604 | 19 |
| TCTGA | 1218390 | 4.617216 | 164.53592 | 18 |
| GTATG | 1053350 | 4.3192616 | 172.7991 | 45 |
| AAGAG | 1177330 | 4.315582 | 160.89085 | 7 |
| TTTTT | 2000185 | 4.3015184 | 4.7141995 | 1 |
| TGAAC | 1181575 | 4.233576 | 155.02429 | 20 |
| GTCAA | 1144030 | 4.099052 | 151.13895 | 35 |
| GAACT | 1141685 | 4.09065 | 154.69284 | 21 |
| ACAGT | 1136090 | 4.070603 | 151.49356 | 32 |
| TCGTA | 1067485 | 4.045346 | 159.9144 | 43 |
| TATGC | 1062695 | 4.0271935 | 159.46567 | 46 |
| ATCTC | 1149260 | 4.02503 | 148.55647 | 40 |
| CGTAT | 1048920 | 3.9749918 | 159.77362 | 44 |
| TCACA | 1177565 | 3.8993115 | 140.95662 | 30 |
| AACTC | 1158850 | 3.8373396 | 143.09499 | 22 |
| TCAAA | 1557805 | 3.7161632 | 101.20243 | 36 |
| CAAAT | 1448730 | 3.4559627 | 101.463486 | 37 |
| AAATC | 1436135 | 3.4259176 | 101.361 | 38 |
| AAAAA | 1997890 | 3.246265 | 3.4457302 | 5 |
| AATCT | 1231365 | 3.1068218 | 106.81786 | 39 |
| TGCCG | 211470 | 1.5797217 | 9.146477 | 47 |