![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW217_CTTGTA_L003_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 7224626 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
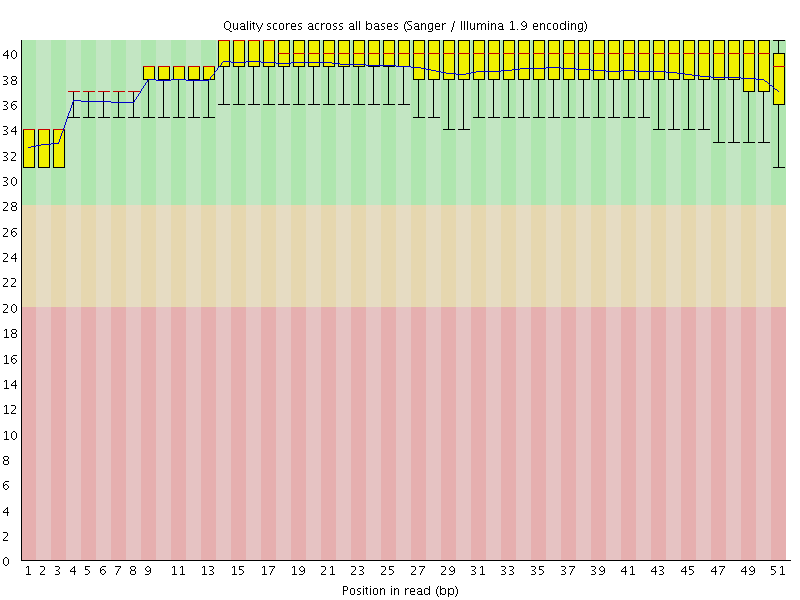
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
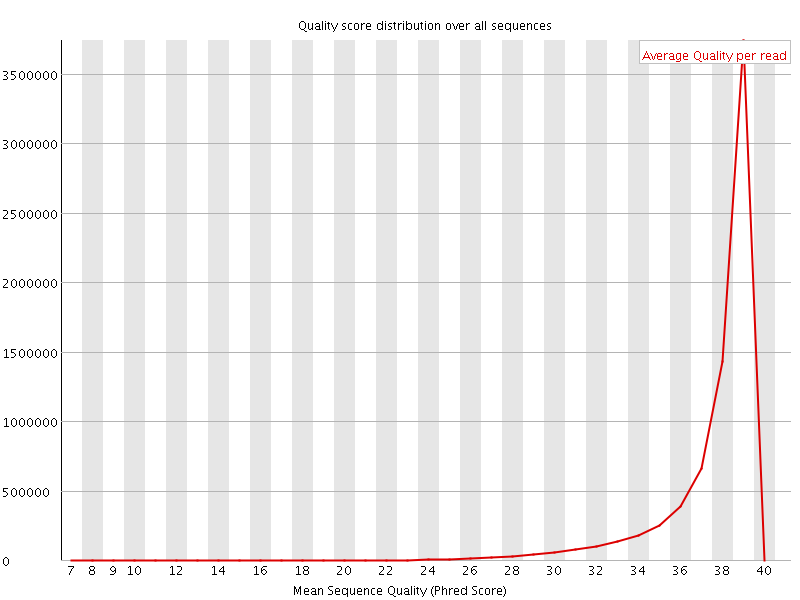
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
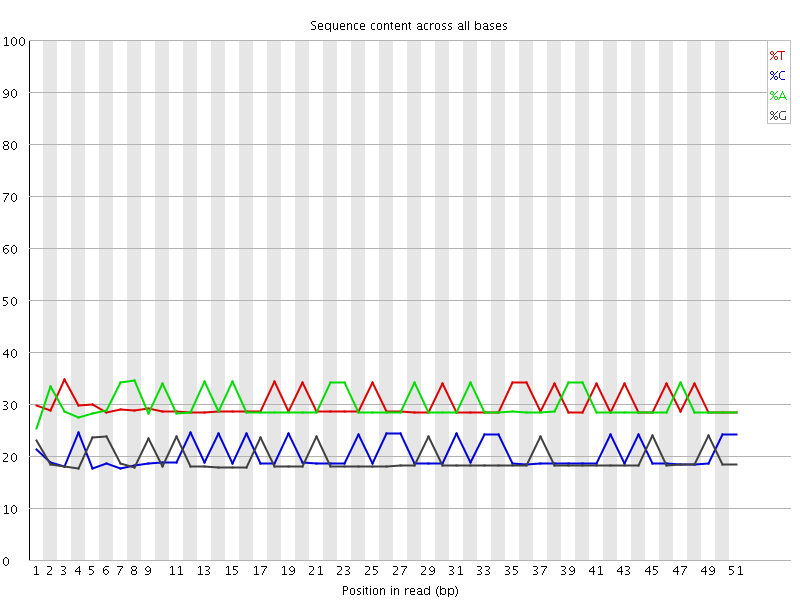
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
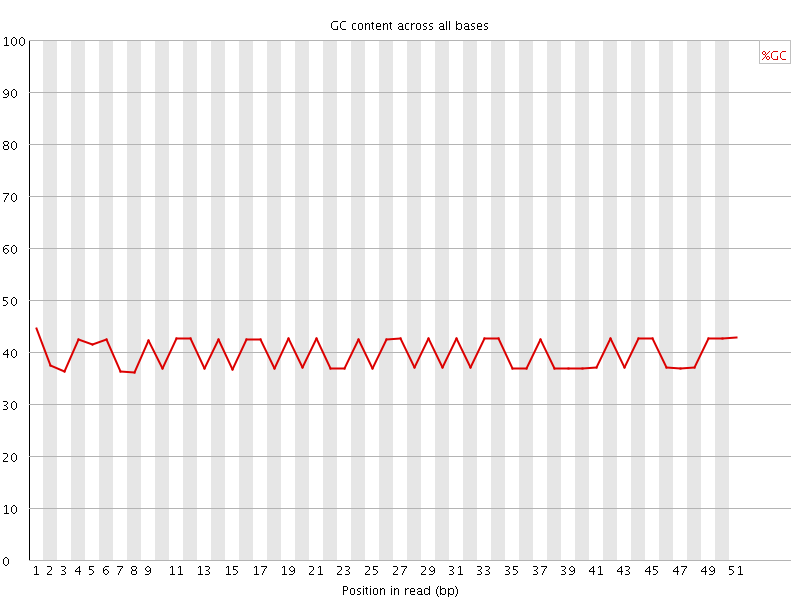
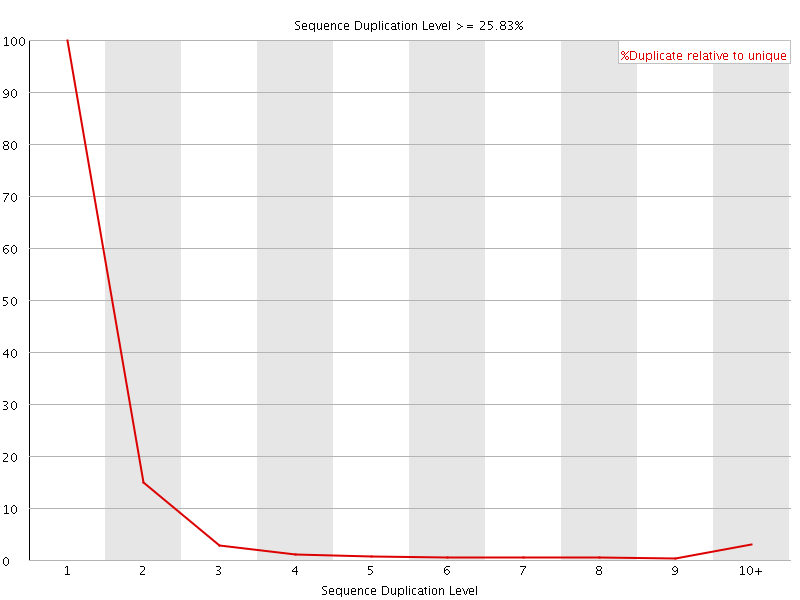
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
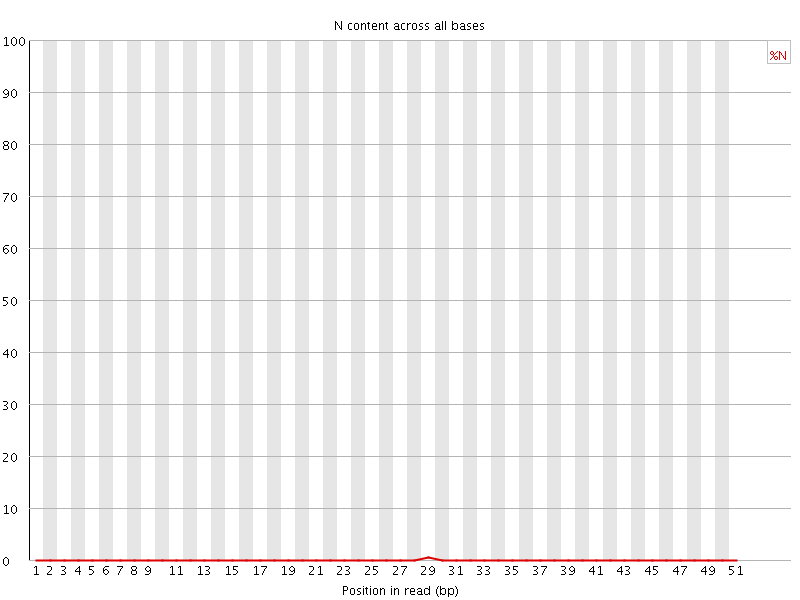
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
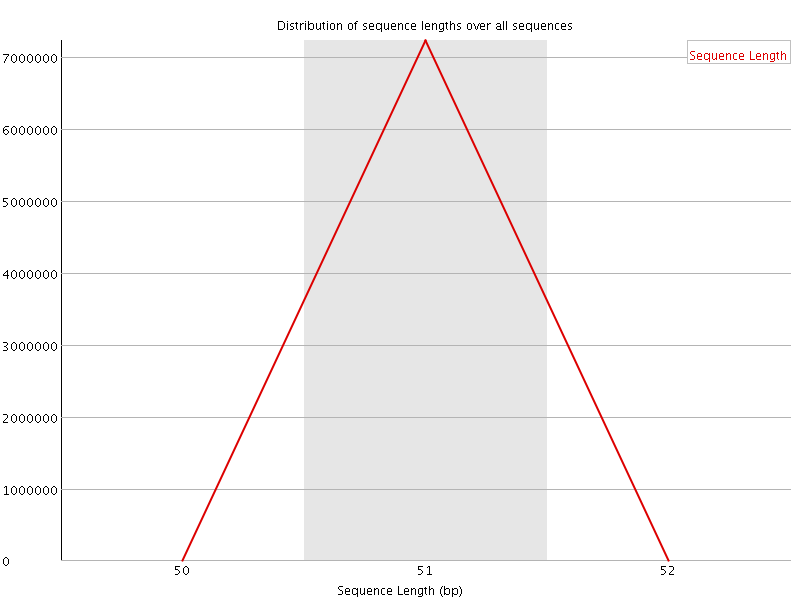
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCTTGTAATCTCGTATGCC | 375812 | 5.2018194436639345 | TruSeq Adapter, Index 12 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
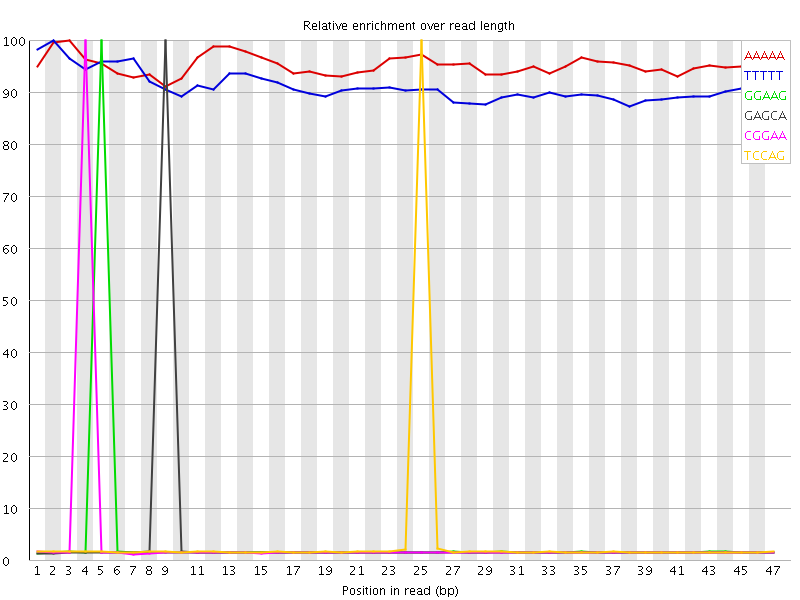
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 3046115 | 3.7252295 | 3.909003 | 3 |
| TTTTT | 3106060 | 3.6991026 | 4.0594277 | 2 |
| GGAAG | 740565 | 3.2836359 | 87.58208 | 5 |
| GAGCA | 734965 | 3.1083007 | 83.43193 | 9 |
| CGGAA | 731260 | 3.0926316 | 83.65235 | 4 |
| TCCAG | 743410 | 2.9829452 | 78.029594 | 25 |
| CTCCA | 752655 | 2.8805628 | 74.45761 | 24 |
| TCACC | 736515 | 2.8187919 | 73.981804 | 30 |
| GAAGA | 965180 | 2.7857265 | 57.619495 | 6 |
| TCGGA | 647855 | 2.7253993 | 82.784065 | 3 |
| TCTCG | 682720 | 2.7249308 | 76.914955 | 41 |
| GCACA | 673150 | 2.7153933 | 79.19421 | 11 |
| AGAGC | 641455 | 2.71283 | 83.068535 | 8 |
| CACAC | 704420 | 2.7102983 | 75.58095 | 12 |
| CTGAA | 972945 | 2.6642742 | 54.327663 | 19 |
| ATCGG | 626690 | 2.6363623 | 82.585556 | 2 |
| AGCAC | 651335 | 2.6273947 | 79.12508 | 10 |
| CCAGT | 653680 | 2.622902 | 77.570564 | 26 |
| GTCTG | 621760 | 2.601783 | 81.60211 | 17 |
| GATCG | 596410 | 2.5089803 | 82.27571 | 1 |
| GTCAC | 624860 | 2.507261 | 77.55444 | 29 |
| CACGT | 616240 | 2.4726732 | 78.33902 | 14 |
| ATGCC | 615730 | 2.4706268 | 76.94274 | 47 |
| CGTCT | 617920 | 2.4662955 | 77.807816 | 16 |
| CTCGT | 616025 | 2.458732 | 76.32963 | 42 |
| ACACG | 602390 | 2.4299574 | 78.72164 | 13 |
| CCTTG | 607785 | 2.4258437 | 76.50212 | 33 |
| CACCT | 633365 | 2.4240158 | 73.40961 | 31 |
| CAGTC | 596780 | 2.3945897 | 77.331566 | 27 |
| ACGTC | 595310 | 2.3886912 | 78.105774 | 15 |
| ACTCC | 610210 | 2.335397 | 74.28785 | 23 |
| AAGAG | 787765 | 2.2736666 | 57.14517 | 7 |
| TCTGA | 833220 | 2.2695842 | 53.64557 | 18 |
| TGAAC | 803465 | 2.2001767 | 53.84712 | 20 |
| GAACT | 737790 | 2.020335 | 53.612408 | 21 |
| CTTGT | 726555 | 1.9685711 | 52.11509 | 34 |
| AACTC | 753330 | 1.9676164 | 51.10258 | 22 |
| ATCTC | 732720 | 1.903659 | 50.07114 | 40 |
| AGTCA | 691960 | 1.8948361 | 52.970108 | 28 |
| ACCTT | 665095 | 1.7279646 | 49.917465 | 32 |
| GTATG | 601640 | 1.718141 | 54.681477 | 45 |
| TATGC | 612415 | 1.6681397 | 52.123642 | 46 |
| TCGTA | 608335 | 1.6570263 | 52.133205 | 43 |
| CGTAT | 589735 | 1.6063622 | 52.121616 | 44 |
| AATCT | 871320 | 1.5449079 | 34.26623 | 39 |
| TTGTA | 828555 | 1.5320667 | 35.735306 | 35 |
| TGTAA | 770855 | 1.4329567 | 35.750088 | 36 |
| GTAAT | 705600 | 1.311653 | 35.578503 | 37 |
| TAATC | 690170 | 1.223717 | 33.86762 | 38 |