![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW216_GGCTAC_L003_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 7880897 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
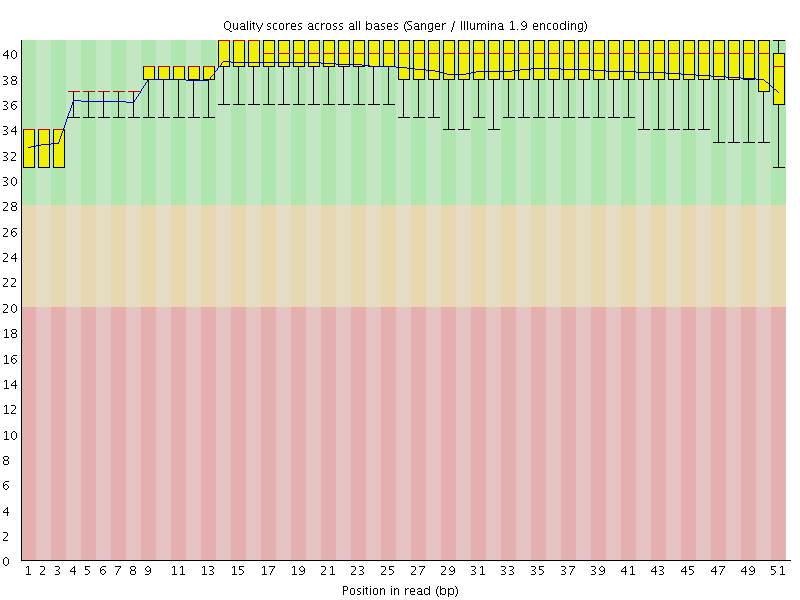
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
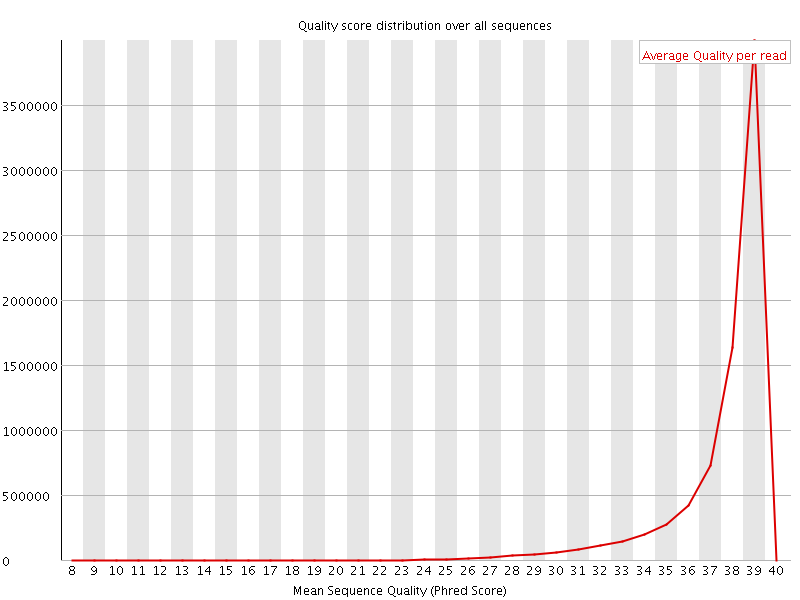
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
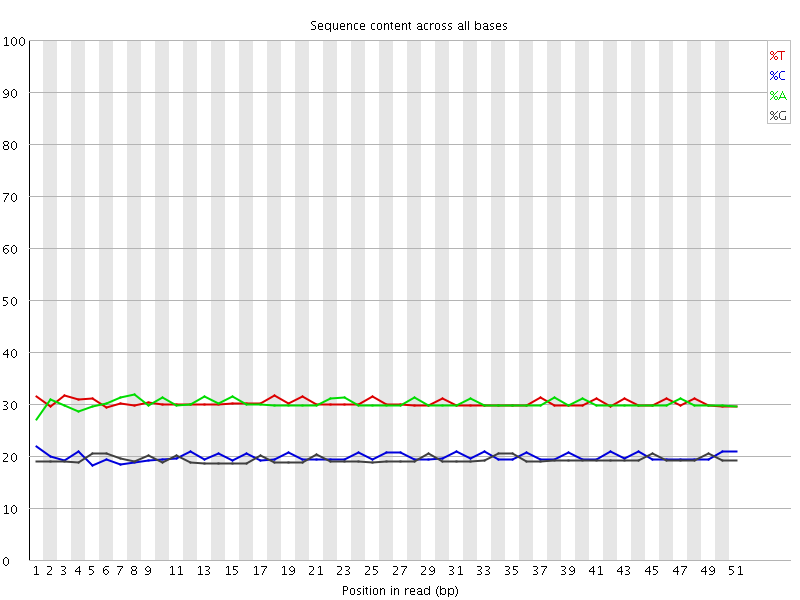
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
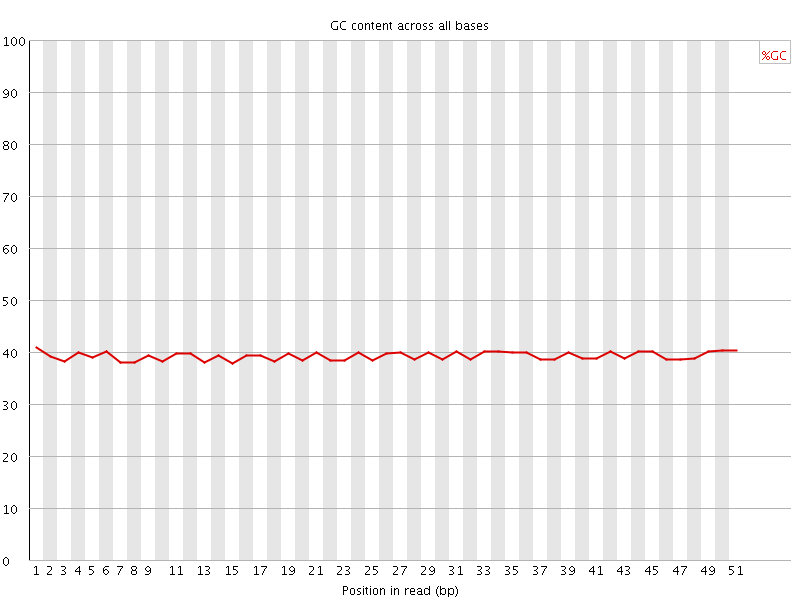
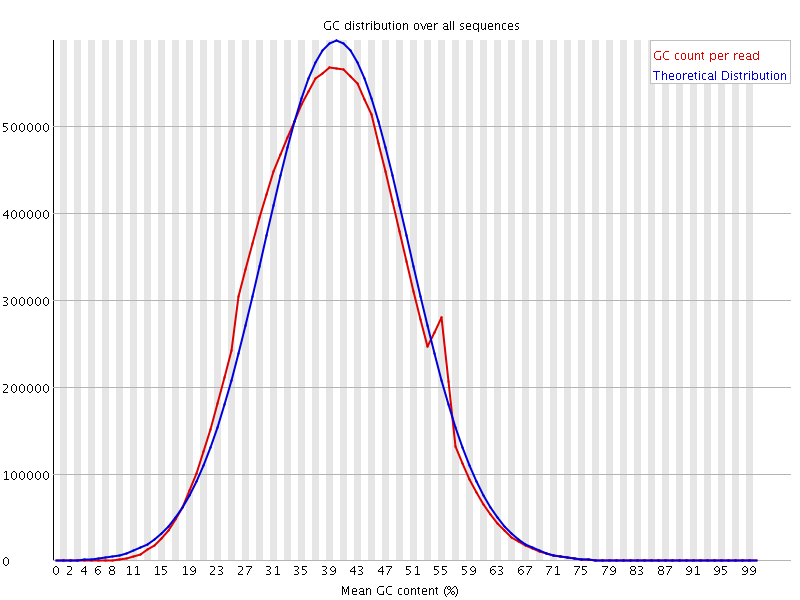
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
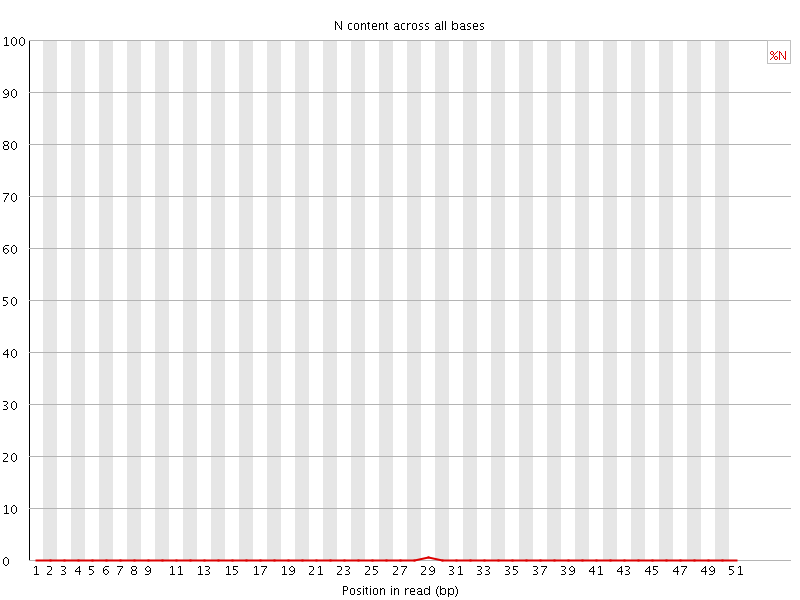
![[OK]](Icons/tick.png) Per base N content
Per base N content
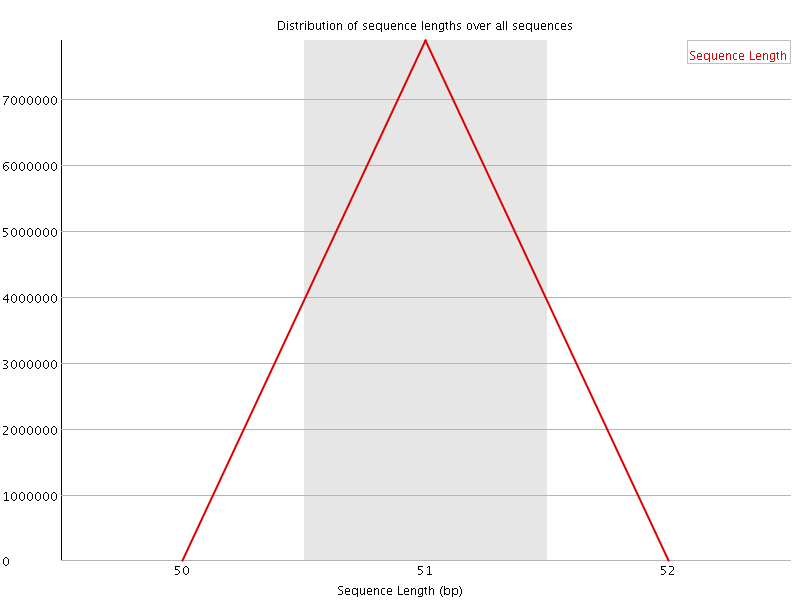
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
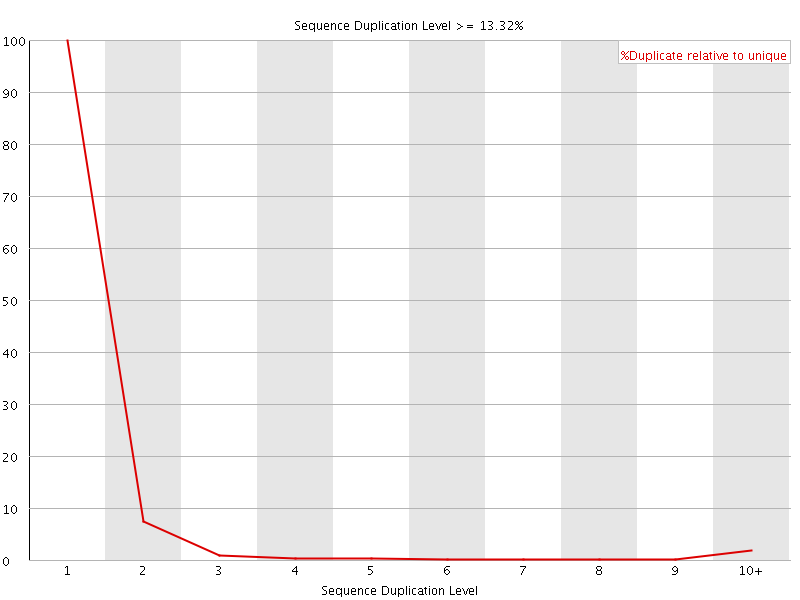
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGGCTACATCTCGTATGCC | 99460 | 1.262039080069185 | TruSeq Adapter, Index 11 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
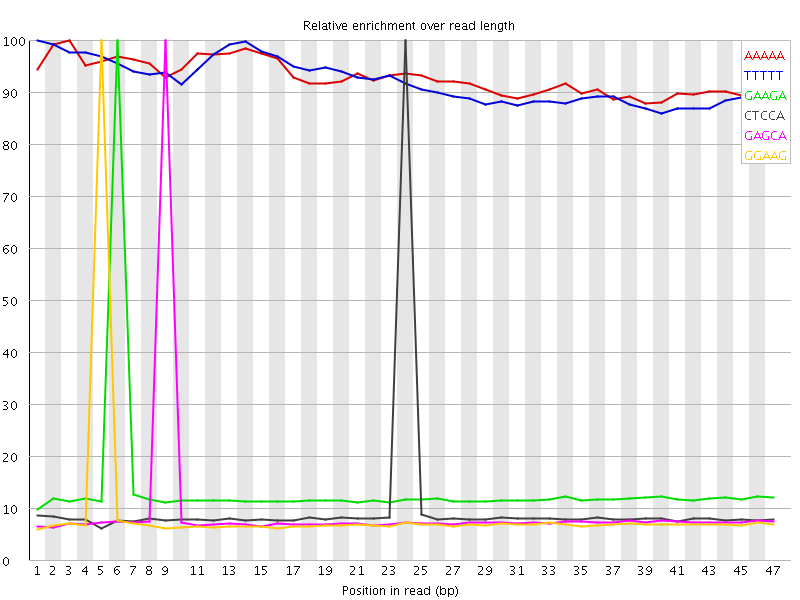
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 3187580 | 3.4068162 | 3.6683125 | 3 |
| TTTTT | 3208630 | 3.3656788 | 3.6574876 | 1 |
| GAAGA | 830245 | 2.1481683 | 15.799152 | 6 |
| CTCCA | 573545 | 2.11968 | 21.216581 | 24 |
| GAGCA | 538325 | 2.1088903 | 22.805517 | 9 |
| GGAAG | 516170 | 2.0779831 | 23.434853 | 5 |
| TCCAG | 523515 | 1.9882536 | 21.60454 | 25 |
| CTGAA | 784060 | 1.9667292 | 15.09735 | 19 |
| CGGAA | 497660 | 1.949585 | 22.720907 | 4 |
| CGGCT | 289840 | 1.7127272 | 31.458538 | 33 |
| TCTCG | 450385 | 1.704118 | 20.838692 | 41 |
| TCGGA | 429780 | 1.6773692 | 22.404139 | 3 |
| CACAC | 436185 | 1.6180817 | 21.063238 | 12 |
| ACGGC | 272400 | 1.6157117 | 31.428701 | 32 |
| AGAGC | 411385 | 1.6116023 | 22.3236 | 8 |
| TCTGA | 629670 | 1.573553 | 14.6518955 | 18 |
| AGCAC | 412545 | 1.5726817 | 21.739334 | 10 |
| GCACA | 410720 | 1.5657245 | 21.64447 | 11 |
| CACGG | 261020 | 1.5482124 | 31.52548 | 31 |
| AAGAG | 569465 | 1.4734284 | 15.129159 | 7 |
| CCAGT | 381630 | 1.4493896 | 21.050577 | 26 |
| ATCGG | 370595 | 1.4463787 | 22.055964 | 2 |
| CTCGT | 371820 | 1.4068522 | 20.508902 | 42 |
| TGAAC | 559460 | 1.4033445 | 14.537751 | 20 |
| CGTCT | 363850 | 1.3766961 | 21.174856 | 16 |
| CACGT | 360030 | 1.3673551 | 21.255043 | 14 |
| GTCAC | 356445 | 1.3537397 | 20.82163 | 29 |
| GTCTG | 345650 | 1.3439779 | 21.75509 | 17 |
| ACTCC | 356045 | 1.315854 | 20.493282 | 23 |
| ACACG | 336600 | 1.2831684 | 21.310654 | 13 |
| GATCG | 326655 | 1.2748872 | 21.75537 | 1 |
| ATGCC | 335435 | 1.273946 | 20.372213 | 47 |
| CATCT | 523585 | 1.273256 | 13.411323 | 39 |
| AACTC | 521355 | 1.2725912 | 14.01196 | 22 |
| TCACG | 331275 | 1.2581469 | 20.563227 | 30 |
| ACGTC | 330665 | 1.25583 | 21.127705 | 15 |
| CAGTC | 329445 | 1.2511966 | 20.828812 | 27 |
| GAACT | 494945 | 1.2415156 | 14.345161 | 21 |
| ATCTC | 507475 | 1.2340796 | 13.511457 | 40 |
| CTACA | 474925 | 1.159259 | 13.301231 | 36 |
| ACATC | 464745 | 1.1344103 | 13.299412 | 38 |
| AGTCA | 434355 | 1.0895323 | 13.991615 | 28 |
| GCTAC | 272140 | 1.0335584 | 19.985489 | 35 |
| GGCTA | 242065 | 0.94474477 | 20.560629 | 34 |
| TCGTA | 375600 | 0.938629 | 13.494111 | 43 |
| TATGC | 340485 | 0.85087615 | 13.357231 | 46 |
| GTATG | 326150 | 0.8375787 | 13.737426 | 45 |
| CGTAT | 328595 | 0.8211629 | 13.374365 | 44 |
| TACAT | 479955 | 0.7708709 | 8.811427 | 37 |