![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW214_GATCAG_L003_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 9608336 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
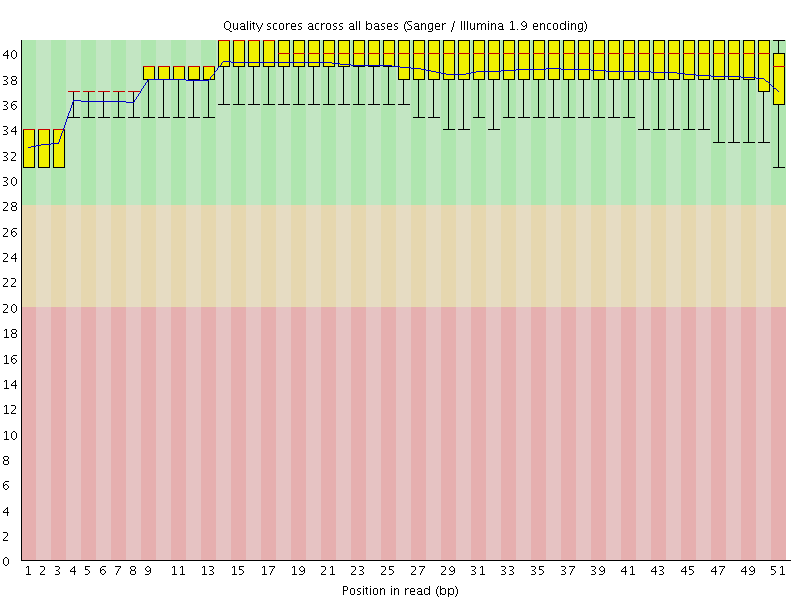
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
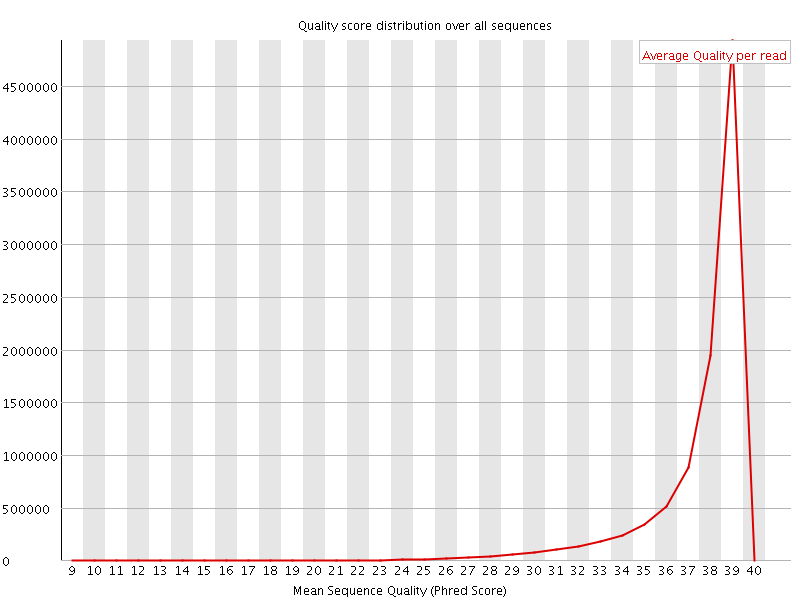
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
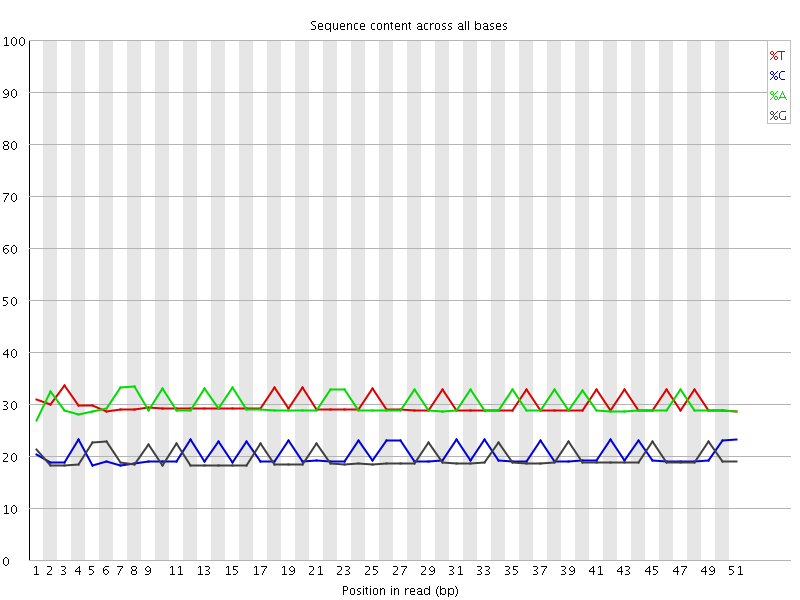
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
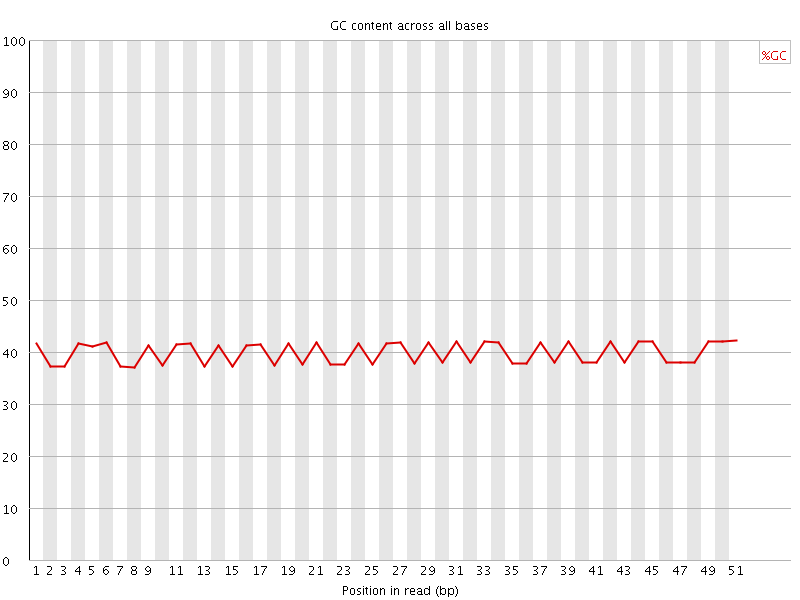
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
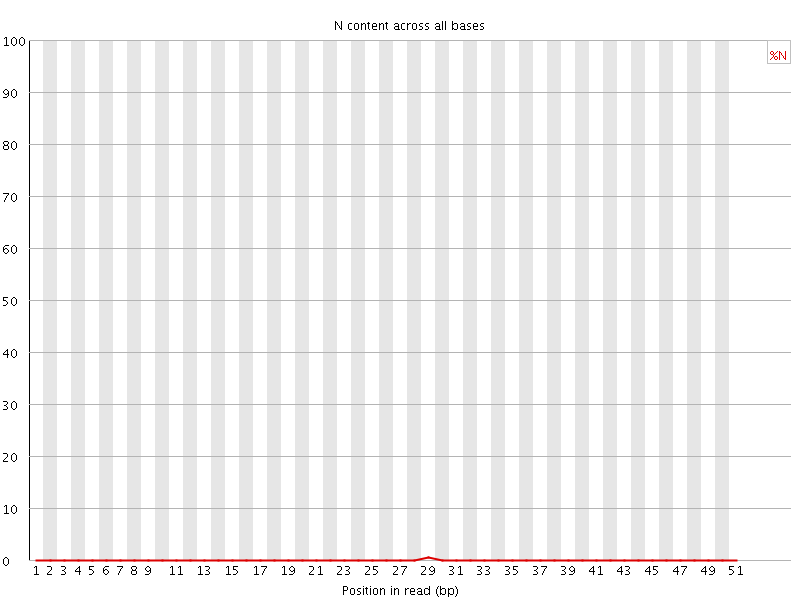
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
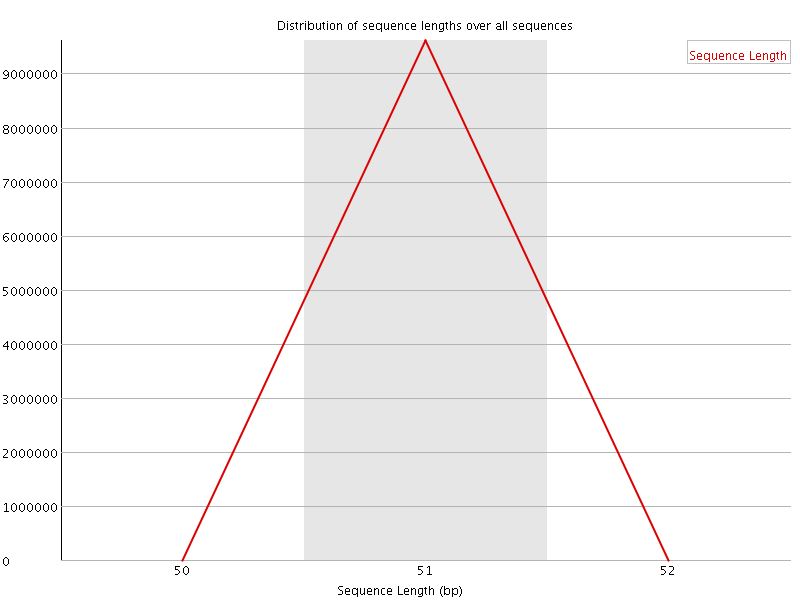
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
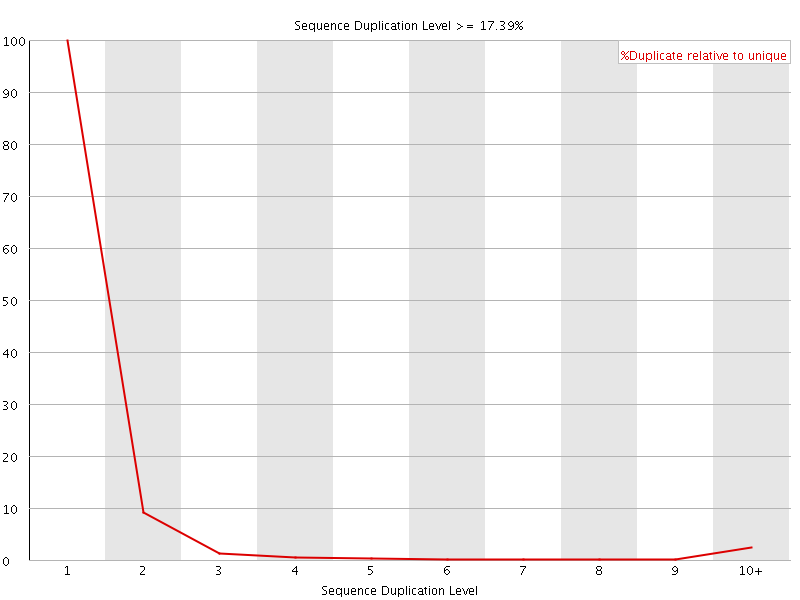
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGATCAGATCTCGTATGCC | 350543 | 3.648321624056444 | TruSeq Adapter, Index 9 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
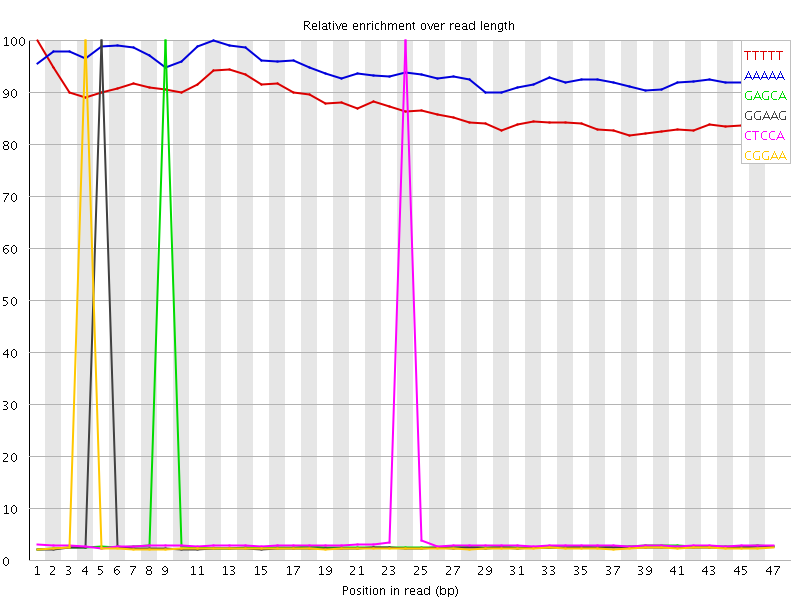
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 3778640 | 3.4609442 | 3.961253 | 1 |
| AAAAA | 3717520 | 3.3750203 | 3.5858076 | 12 |
| GAGCA | 900310 | 2.8191345 | 59.341103 | 9 |
| GGAAG | 867685 | 2.817027 | 61.56899 | 5 |
| CTCCA | 934990 | 2.72827 | 54.179848 | 24 |
| CGGAA | 850190 | 2.662194 | 59.380062 | 4 |
| TCCAG | 872320 | 2.6391344 | 56.079666 | 25 |
| GAAGA | 1242260 | 2.6373806 | 41.01144 | 6 |
| TCTCG | 801005 | 2.4276612 | 55.748413 | 41 |
| TCGGA | 769215 | 2.4128962 | 59.23172 | 3 |
| CTGAA | 1171610 | 2.4032855 | 39.01232 | 19 |
| AGAGC | 751760 | 2.3539805 | 58.894142 | 8 |
| CACAC | 796170 | 2.3190978 | 54.603786 | 12 |
| AGCAC | 761680 | 2.3003345 | 56.68969 | 10 |
| GCACA | 757865 | 2.288813 | 56.618793 | 11 |
| ATCGG | 702755 | 2.2044227 | 58.89003 | 2 |
| CTCGT | 705245 | 2.137435 | 55.282764 | 42 |
| CCAGT | 703790 | 2.1292603 | 55.39671 | 26 |
| CGTCT | 699990 | 2.1215081 | 56.32978 | 16 |
| CACGA | 701695 | 2.1191752 | 54.87398 | 31 |
| GTCTG | 673220 | 2.1155105 | 58.349964 | 17 |
| CACGT | 690175 | 2.0880692 | 56.247997 | 14 |
| TCAGA | 997945 | 2.0470521 | 37.675556 | 36 |
| GTCAC | 676475 | 2.046621 | 55.446053 | 29 |
| TCTGA | 995495 | 2.045637 | 38.758034 | 18 |
| GATCG | 650170 | 2.039472 | 58.546276 | 1 |
| ACACG | 664105 | 2.00565 | 56.17495 | 13 |
| ATGCC | 660215 | 1.9974277 | 55.19769 | 47 |
| AAGAG | 940080 | 1.9958372 | 40.361004 | 7 |
| ACTCC | 683850 | 1.9954518 | 53.77838 | 23 |
| ACGTC | 655315 | 1.9826031 | 56.115982 | 15 |
| TCACG | 655195 | 1.9822401 | 55.208588 | 30 |
| CAGTC | 646935 | 1.9572501 | 55.248676 | 27 |
| CGATC | 641510 | 1.9408371 | 54.517937 | 33 |
| TGAAC | 911065 | 1.868838 | 38.46034 | 20 |
| ATCAG | 847685 | 1.7388285 | 37.312572 | 35 |
| GATCA | 840075 | 1.7232183 | 37.337425 | 34 |
| AACTC | 869830 | 1.7208831 | 36.907185 | 22 |
| GAACT | 836820 | 1.7165414 | 38.26045 | 21 |
| ATCTC | 862265 | 1.7089329 | 36.428658 | 40 |
| CAGAT | 826980 | 1.6963571 | 37.28585 | 37 |
| ACGAT | 809580 | 1.660665 | 37.489468 | 32 |
| AGATC | 787980 | 1.6163577 | 37.37183 | 38 |
| GATCT | 775565 | 1.593704 | 37.489544 | 39 |
| AGTCA | 762625 | 1.5643477 | 37.659935 | 28 |
| TCGTA | 723535 | 1.4867878 | 37.580887 | 43 |
| GTATG | 649870 | 1.3845901 | 38.843235 | 45 |
| TATGC | 667525 | 1.3716933 | 37.44075 | 46 |
| CGTAT | 663070 | 1.3625387 | 37.47037 | 44 |