![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW213_ACTTGA_L003_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 5394063 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
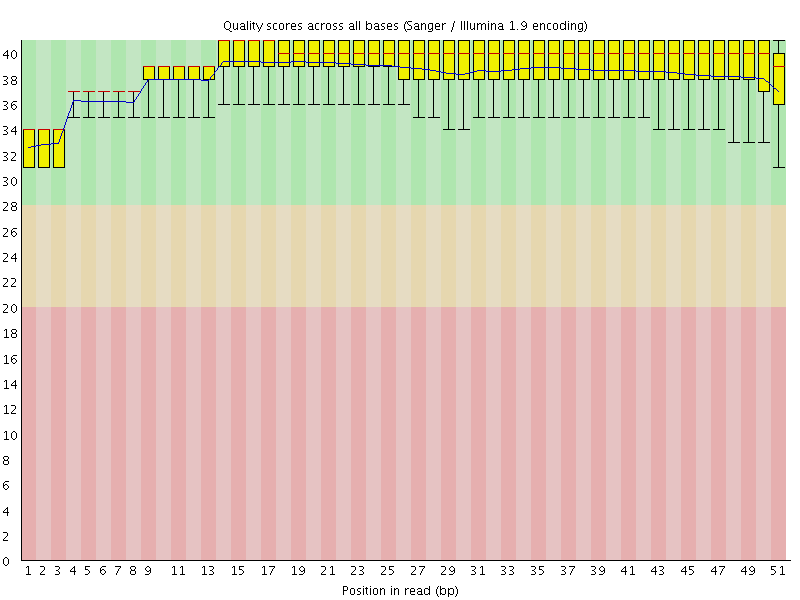
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
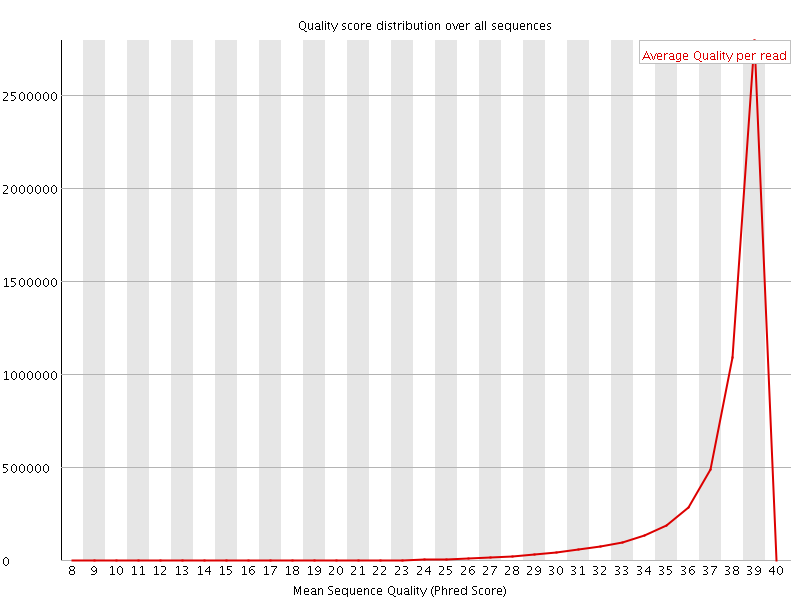
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
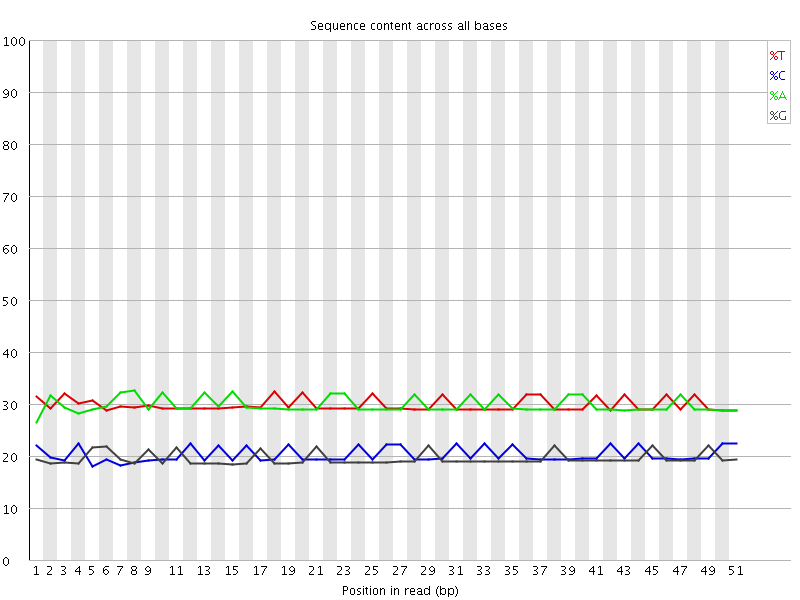
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
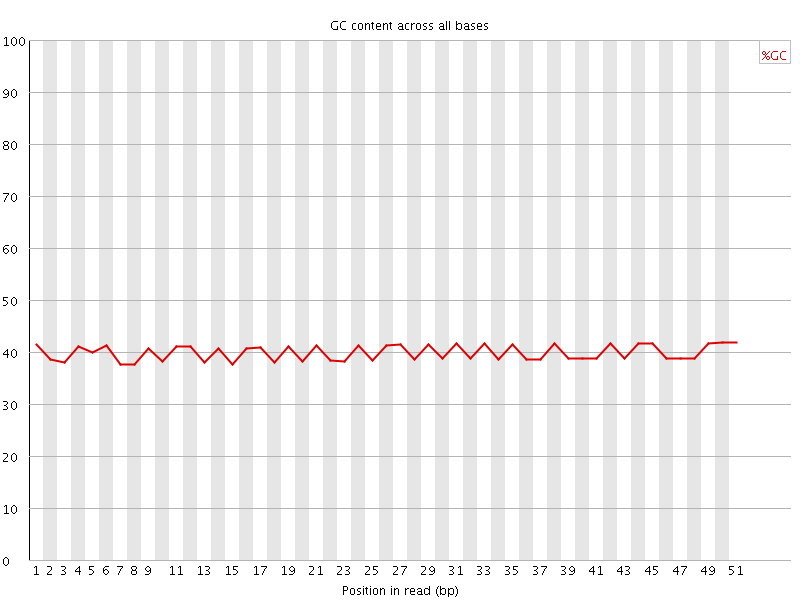
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
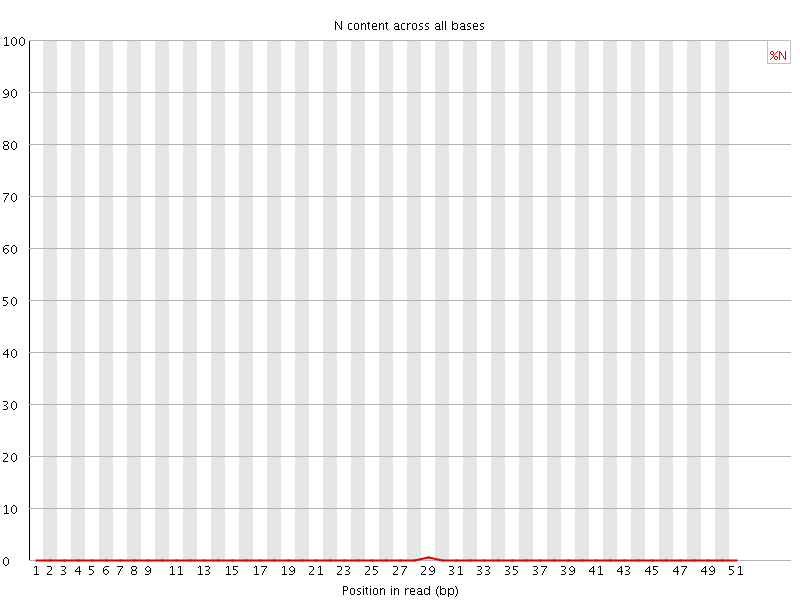
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
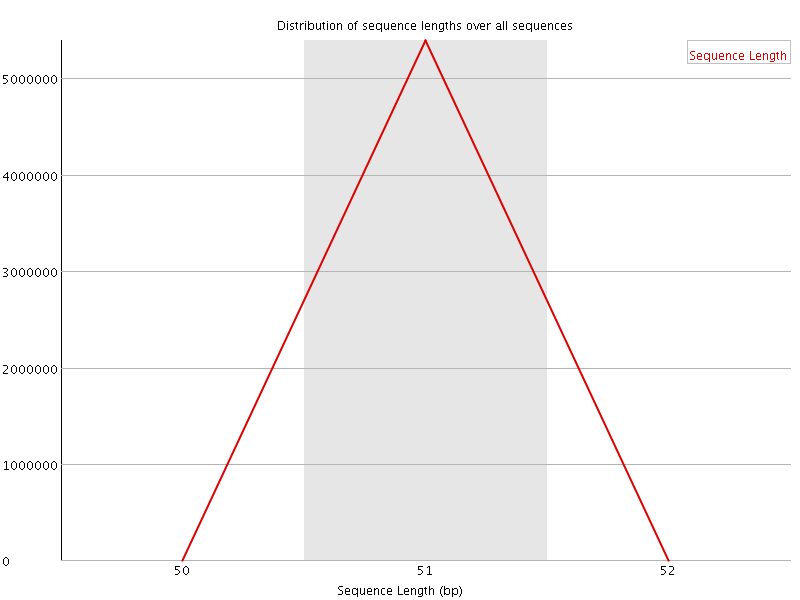
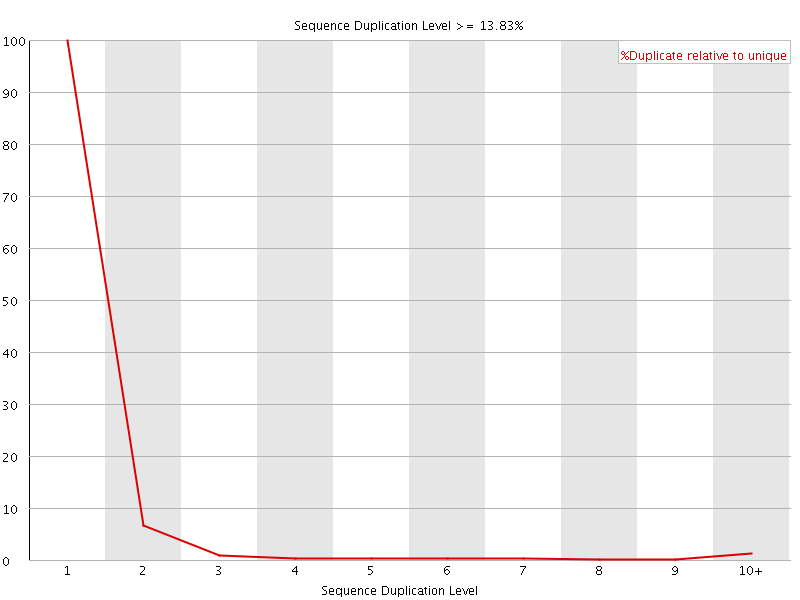
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACACTTGAATCTCGTATGCC | 143878 | 2.6673399995513587 | TruSeq Adapter, Index 8 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
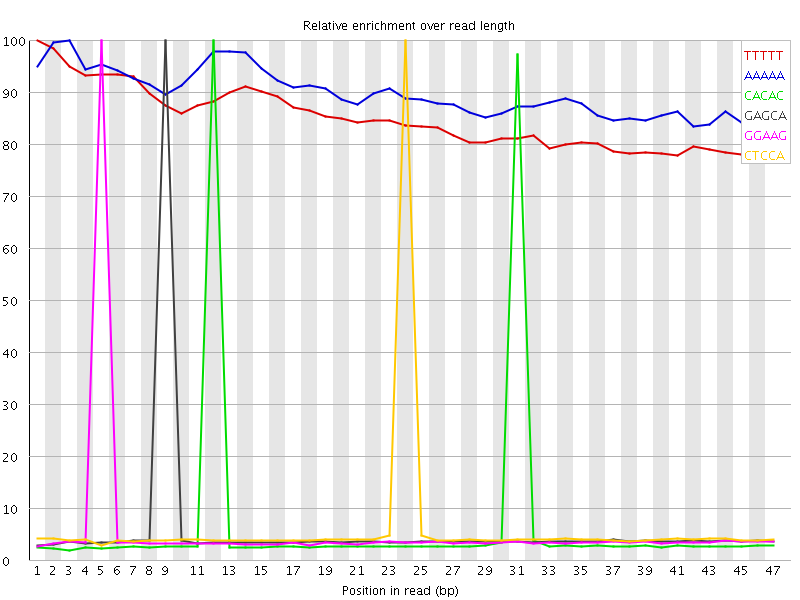
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 2047710 | 3.3384767 | 3.9346588 | 1 |
| AAAAA | 2040800 | 3.3379745 | 3.718208 | 3 |
| CACAC | 532360 | 2.7500305 | 39.98091 | 12 |
| GAGCA | 452520 | 2.5271187 | 43.968365 | 9 |
| GGAAG | 434330 | 2.5219452 | 45.639664 | 5 |
| CTCCA | 471135 | 2.4321868 | 39.82101 | 24 |
| GAAGA | 637670 | 2.427173 | 30.609568 | 6 |
| TCCAG | 440995 | 2.367081 | 41.241577 | 25 |
| CGGAA | 414820 | 2.3165812 | 43.84119 | 4 |
| TTGAA | 895505 | 2.2315192 | 19.966694 | 36 |
| CTGAA | 605530 | 2.215297 | 29.10959 | 19 |
| TCGGA | 372465 | 2.0787048 | 43.497112 | 3 |
| TCTCG | 386440 | 2.0729125 | 40.618015 | 41 |
| AGAGC | 362325 | 2.0234208 | 43.434757 | 8 |
| AGCAC | 373720 | 2.0072722 | 41.809635 | 10 |
| GCACA | 360935 | 1.9386035 | 41.764244 | 11 |
| ATCGG | 335510 | 1.8724611 | 43.18921 | 2 |
| CCAGT | 345510 | 1.8545566 | 40.63643 | 26 |
| TCTGA | 504155 | 1.8432314 | 28.706198 | 18 |
| CTCGT | 334520 | 1.7944069 | 40.21219 | 42 |
| GTCTG | 318970 | 1.7790029 | 42.69394 | 17 |
| CGTCT | 330380 | 1.7721999 | 41.041767 | 16 |
| AAGAG | 463560 | 1.7644554 | 29.945618 | 7 |
| CACGT | 328290 | 1.7621268 | 41.260937 | 14 |
| GTCAC | 322685 | 1.7320412 | 40.466427 | 29 |
| GATCG | 310145 | 1.7309006 | 42.821503 | 1 |
| ATGCC | 315870 | 1.6954612 | 40.153305 | 47 |
| ACTCC | 325955 | 1.6827097 | 39.336662 | 23 |
| ACGTC | 310795 | 1.6682205 | 41.096012 | 15 |
| ACACG | 310460 | 1.6674993 | 41.264904 | 13 |
| TGAAC | 455520 | 1.6664938 | 28.52624 | 20 |
| CTTGA | 455540 | 1.665491 | 27.794495 | 35 |
| CAGTC | 304570 | 1.6348073 | 40.382698 | 27 |
| TCACA | 463465 | 1.6307423 | 26.903728 | 30 |
| CACTT | 454145 | 1.5969172 | 26.819246 | 33 |
| GAATC | 431945 | 1.5802461 | 27.609095 | 38 |
| ACTTG | 422940 | 1.5463026 | 27.693056 | 34 |
| TGAAT | 615270 | 1.5331982 | 19.310072 | 37 |
| AACTC | 435525 | 1.532433 | 27.290232 | 22 |
| GAACT | 415130 | 1.5187294 | 28.369053 | 21 |
| ATCTC | 427530 | 1.5033305 | 26.737421 | 40 |
| AGTCA | 374350 | 1.3695381 | 27.700436 | 28 |
| ACACT | 373025 | 1.3125212 | 26.478117 | 32 |
| TCGTA | 343435 | 1.2556261 | 27.44391 | 43 |
| AATCT | 517920 | 1.2412733 | 18.337858 | 39 |
| GTATG | 303220 | 1.1526607 | 28.377289 | 45 |
| TATGC | 314695 | 1.1505504 | 27.292437 | 46 |
| CGTAT | 309895 | 1.133001 | 27.299677 | 44 |