![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW212_CAGATC_L003_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 5533233 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
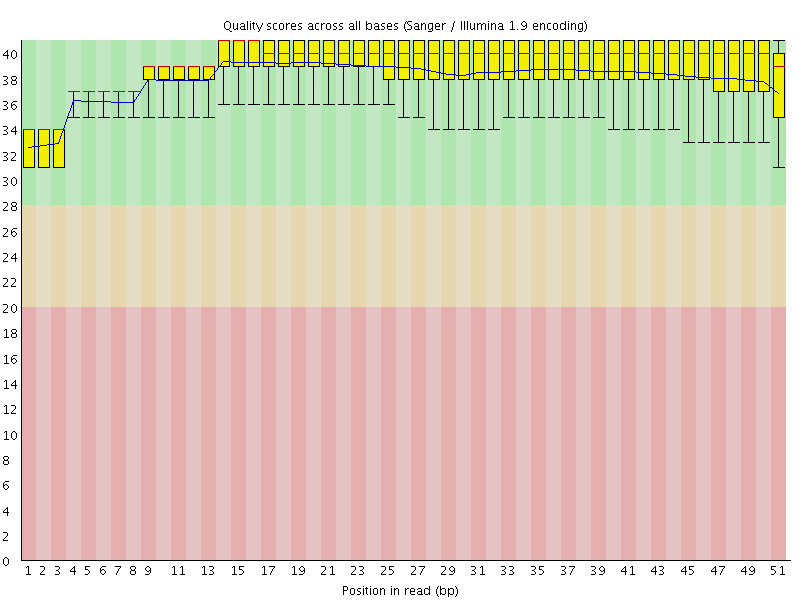
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
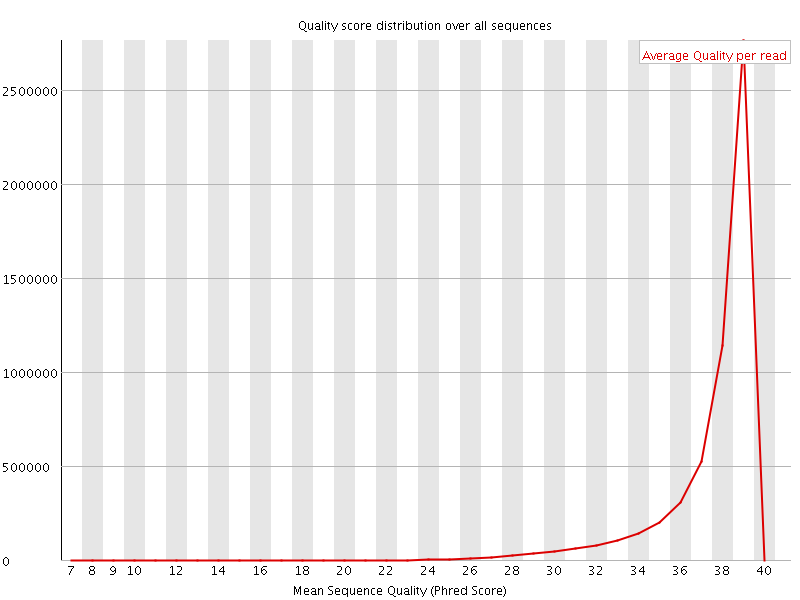
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
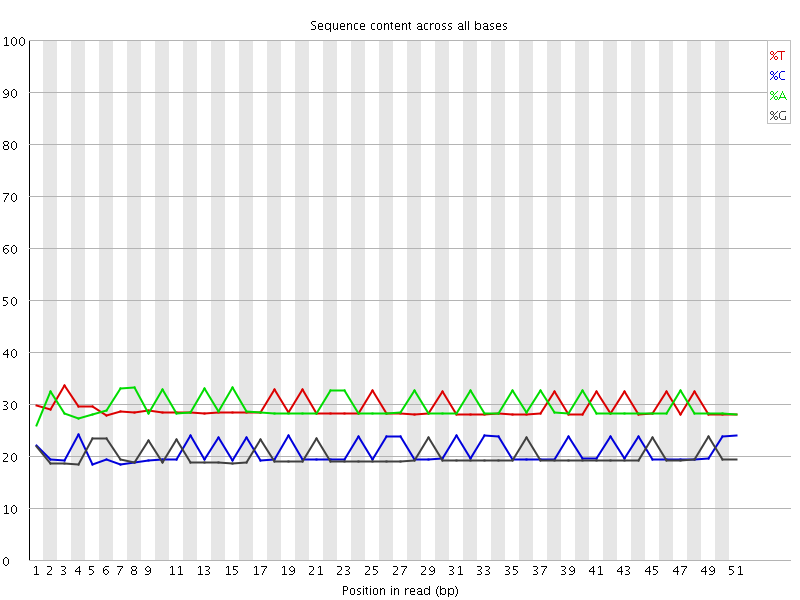
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
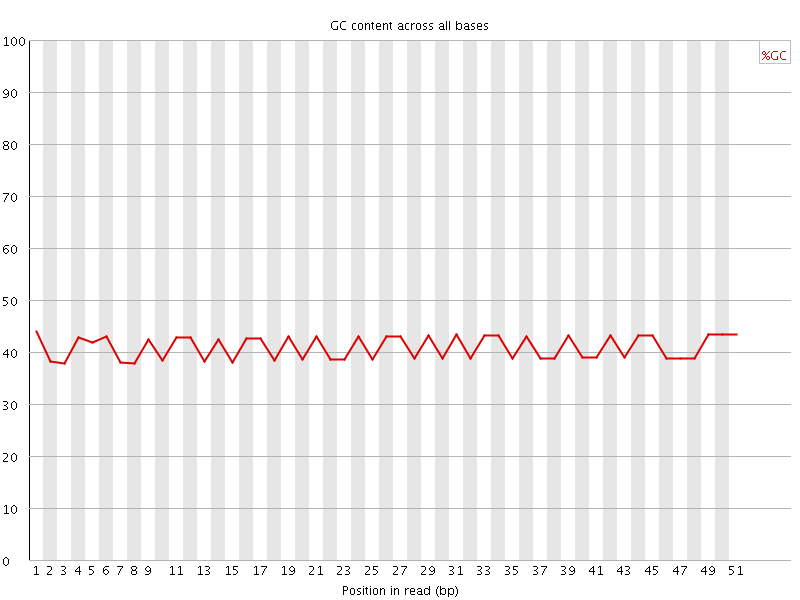
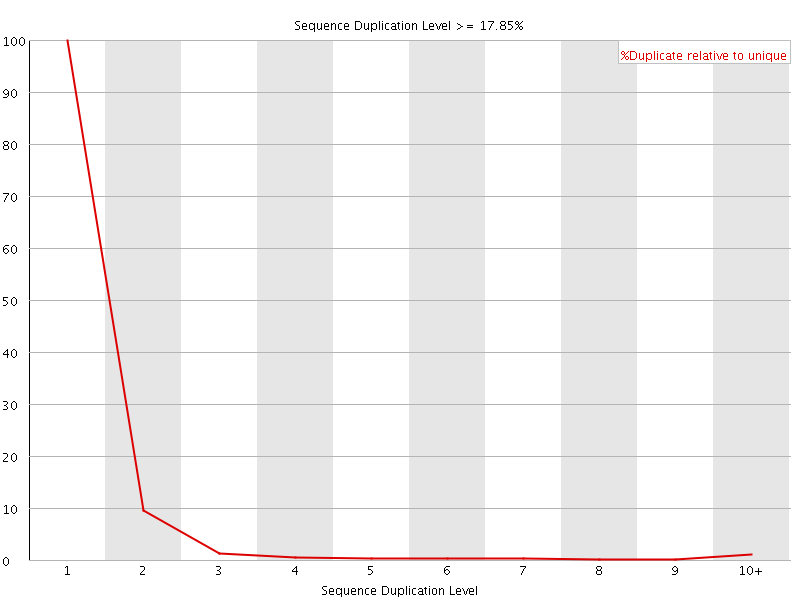
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
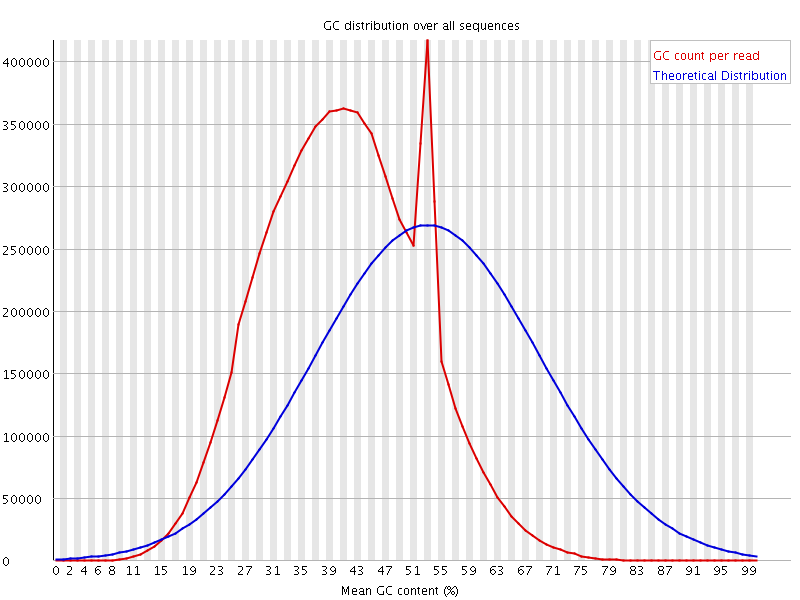
![[OK]](Icons/tick.png) Per base N content
Per base N content
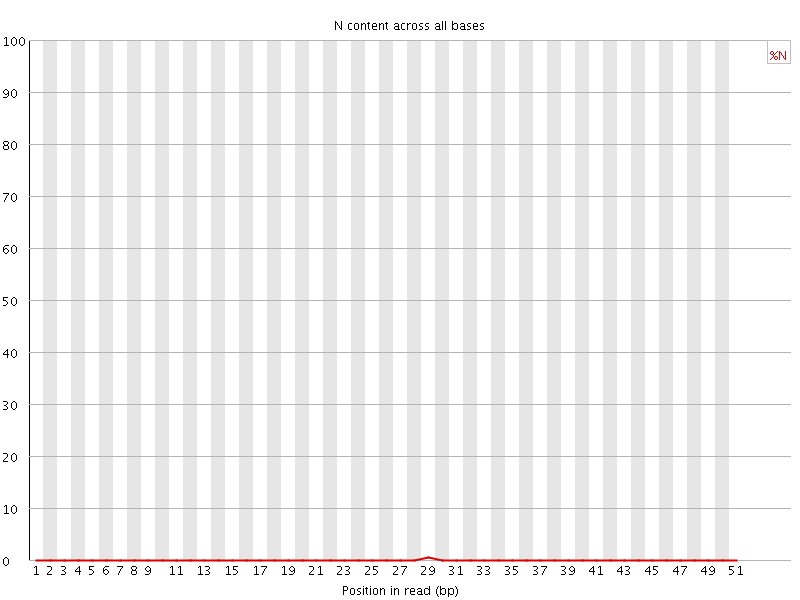
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
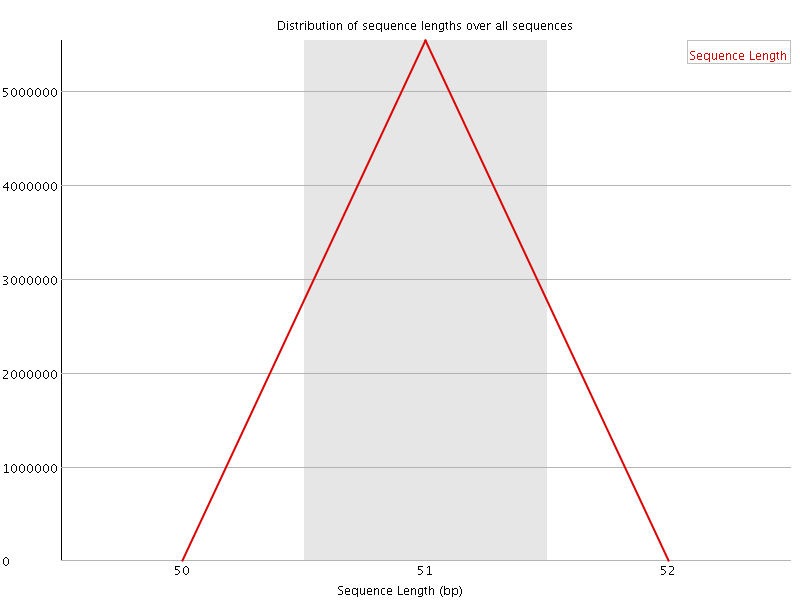
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCAGATCATCTCGTATGCC | 215849 | 3.9009562763758545 | TruSeq Adapter, Index 7 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
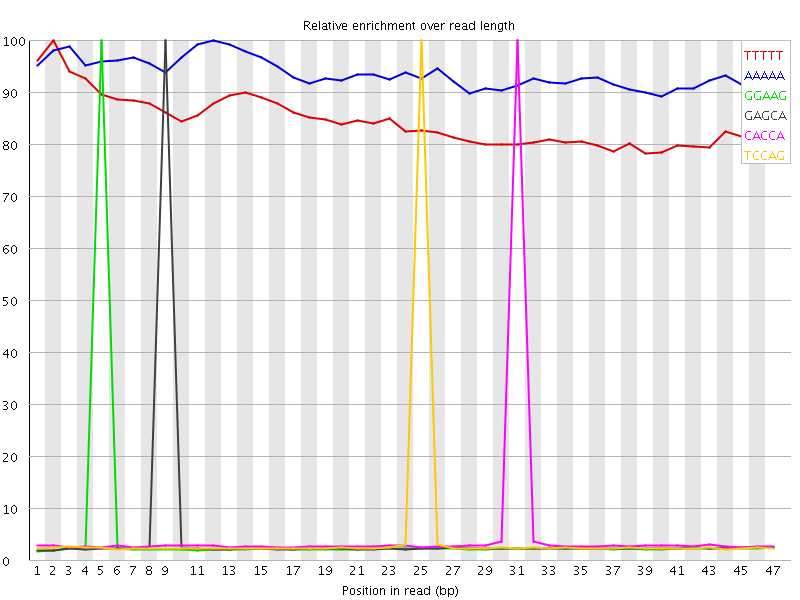
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 2002605 | 3.5444372 | 4.199184 | 2 |
| AAAAA | 2021980 | 3.4212046 | 3.6516829 | 12 |
| GGAAG | 530360 | 2.8711295 | 65.28296 | 5 |
| GAGCA | 533940 | 2.7709332 | 62.175354 | 9 |
| CACCA | 575725 | 2.7456923 | 55.69168 | 31 |
| TCCAG | 537830 | 2.6998534 | 59.18149 | 25 |
| CGGAA | 510350 | 2.6485107 | 62.48847 | 4 |
| CTCCA | 549325 | 2.6434803 | 56.70562 | 24 |
| GAAGA | 709955 | 2.6082723 | 44.86345 | 6 |
| CCAGA | 507495 | 2.5247416 | 57.660294 | 33 |
| CTGAA | 692285 | 2.4601893 | 42.91797 | 19 |
| TCACC | 491125 | 2.363408 | 55.892822 | 30 |
| TCGGA | 451040 | 2.3618844 | 62.731503 | 3 |
| TCTCG | 462565 | 2.3430302 | 59.01729 | 41 |
| AGAGC | 449710 | 2.3338134 | 61.794018 | 8 |
| AGCAC | 461935 | 2.2980847 | 59.193295 | 10 |
| ATCGG | 434460 | 2.2750626 | 62.534805 | 2 |
| CCAGT | 448935 | 2.2536092 | 58.53834 | 26 |
| GCACA | 449870 | 2.2380621 | 59.101025 | 11 |
| ACCAG | 445395 | 2.2157998 | 57.51439 | 32 |
| GTCTG | 418745 | 2.2126014 | 62.327137 | 17 |
| CACAC | 456470 | 2.1769526 | 56.42769 | 12 |
| GATCG | 411870 | 2.1567693 | 62.18723 | 1 |
| CGTCT | 417305 | 2.1137745 | 59.54718 | 16 |
| ATGCC | 420125 | 2.1089857 | 57.913578 | 47 |
| TCTGA | 586865 | 2.1044168 | 42.94409 | 18 |
| GTCAC | 417950 | 2.0980675 | 58.46116 | 29 |
| CTCGT | 412740 | 2.0906515 | 58.421204 | 42 |
| CACGT | 412855 | 2.0724912 | 59.276073 | 14 |
| TCATC | 598255 | 2.0565128 | 40.10166 | 38 |
| AAGAG | 547685 | 2.0121157 | 44.28558 | 7 |
| ACGTC | 399775 | 2.0068307 | 59.112476 | 15 |
| ACTCC | 413340 | 1.9890884 | 56.31439 | 23 |
| CAGTC | 394560 | 1.980652 | 58.234566 | 27 |
| TGAAC | 552710 | 1.9641784 | 42.40553 | 20 |
| ACACG | 394780 | 1.9639949 | 58.700428 | 13 |
| GATCA | 513575 | 1.8251033 | 41.149548 | 36 |
| CAGAT | 506120 | 1.7986104 | 41.154278 | 34 |
| GAACT | 503790 | 1.7903303 | 42.19027 | 21 |
| CATCT | 517450 | 1.7787442 | 39.836033 | 39 |
| AACTC | 515220 | 1.7552052 | 40.402443 | 22 |
| ATCTC | 509680 | 1.7520347 | 40.145214 | 40 |
| AGATC | 480155 | 1.706338 | 41.09449 | 35 |
| AGTCA | 461095 | 1.6386039 | 41.4946 | 28 |
| TCGTA | 421405 | 1.5111002 | 41.325855 | 43 |
| GTATG | 395120 | 1.4779885 | 43.000343 | 45 |
| TATGC | 406510 | 1.4576888 | 41.227787 | 46 |
| ATCAT | 597635 | 1.4543524 | 28.432636 | 37 |
| CGTAT | 395455 | 1.4180471 | 41.27796 | 44 |