![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW211_GCCAAT_L003_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 7109579 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
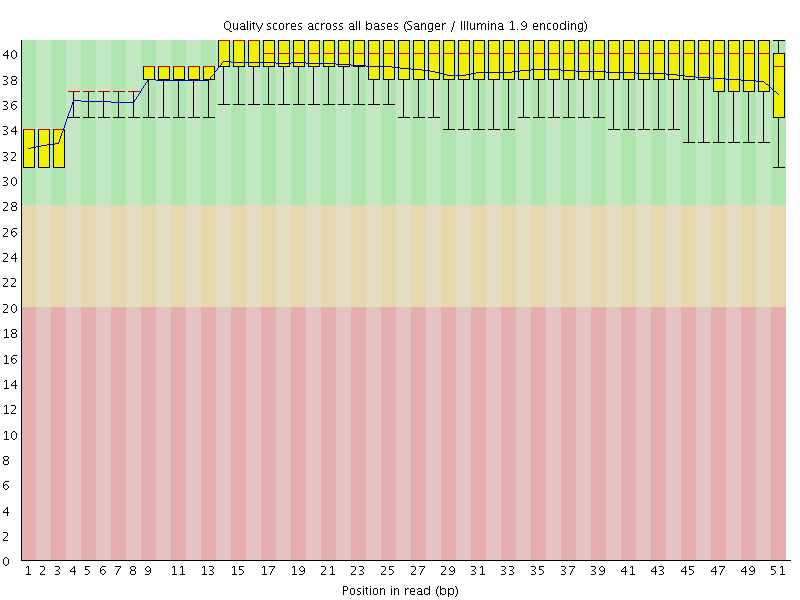
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
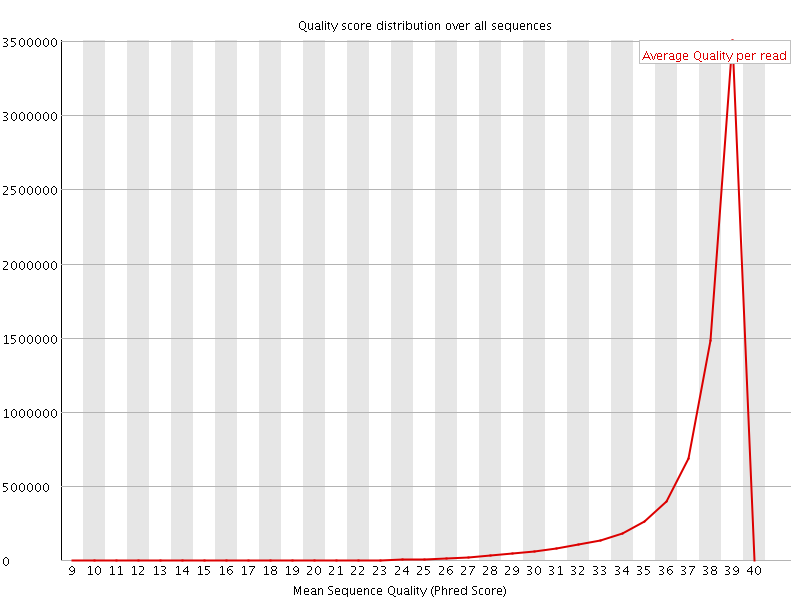
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
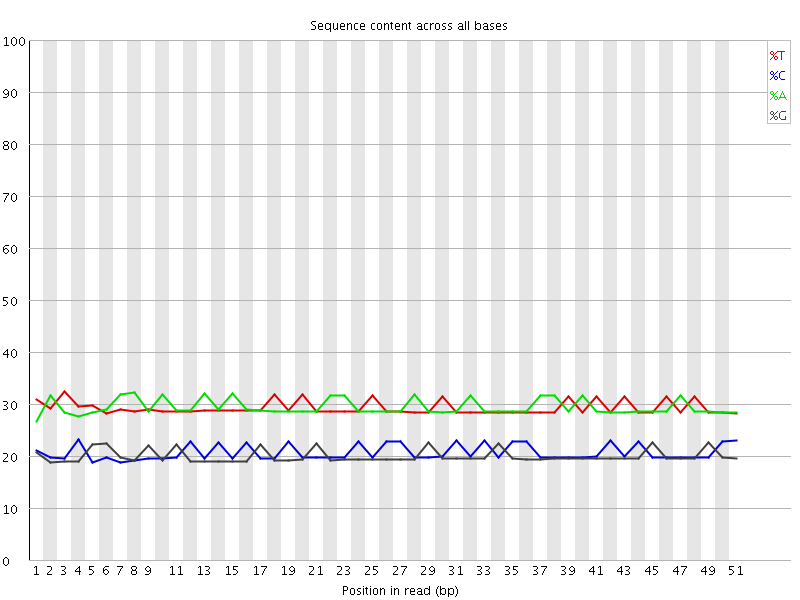
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
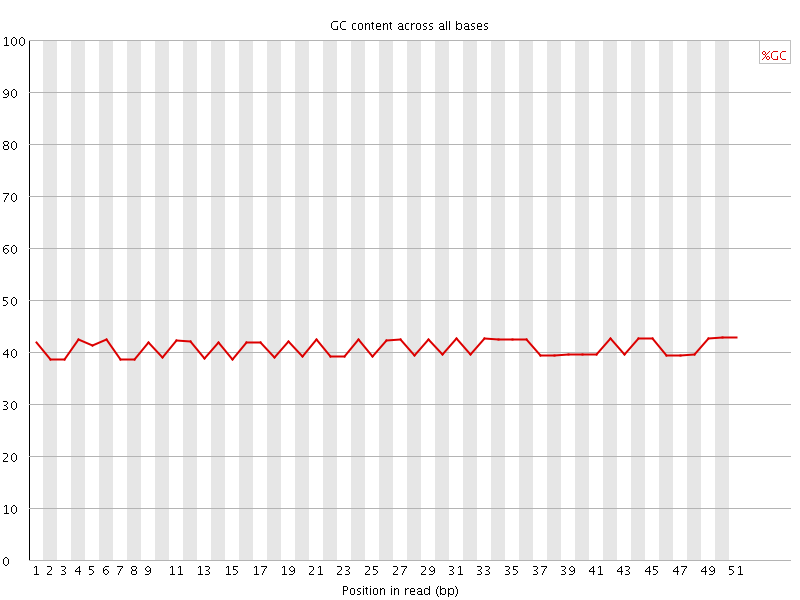
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
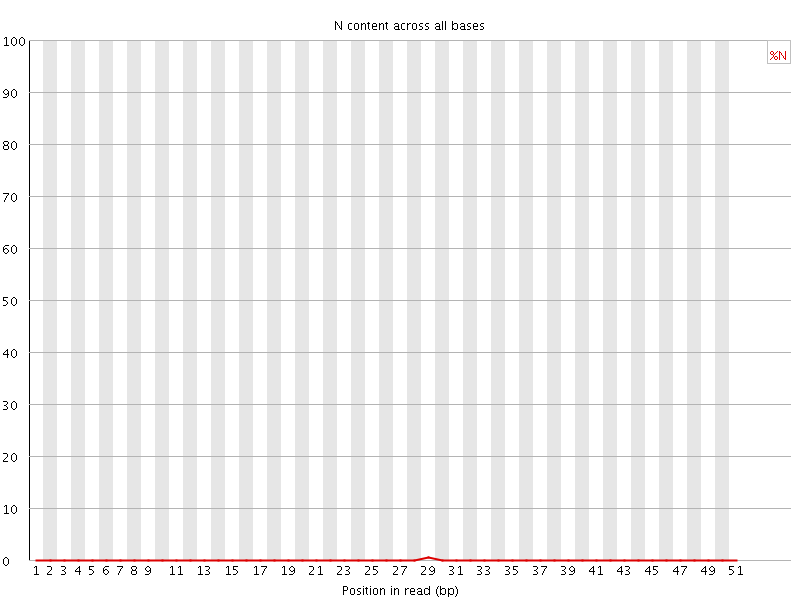
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
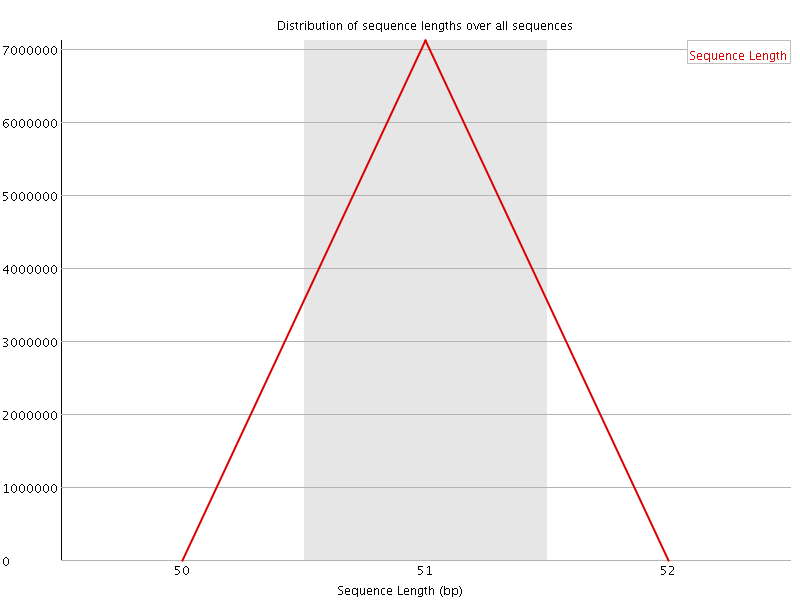
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
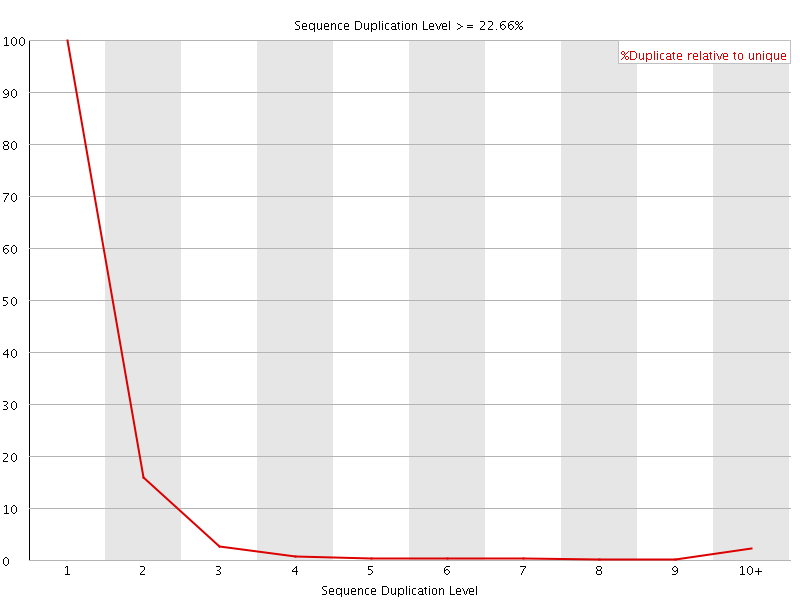
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGCCAATATCTCGTATGCC | 196281 | 2.760796384708574 | TruSeq Adapter, Index 6 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
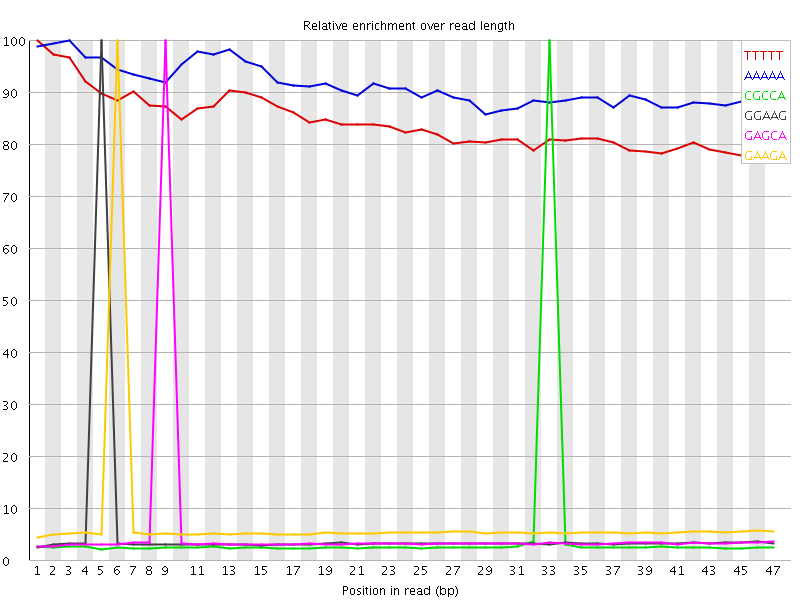
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 2544415 | 3.4619336 | 4.106589 | 1 |
| AAAAA | 2585570 | 3.4168656 | 3.7421825 | 3 |
| CGCCA | 488545 | 2.7044356 | 57.781544 | 33 |
| GGAAG | 593565 | 2.4886096 | 46.303898 | 5 |
| GAGCA | 589340 | 2.384346 | 44.41319 | 9 |
| GAAGA | 827665 | 2.3615878 | 32.08157 | 6 |
| TCCAG | 600825 | 2.3593812 | 42.019943 | 25 |
| CTCCA | 611030 | 2.315408 | 40.53638 | 24 |
| CGGAA | 568330 | 2.2993438 | 44.49297 | 4 |
| CTGAA | 801455 | 2.2196033 | 30.621231 | 19 |
| ACGCC | 396305 | 2.1938236 | 57.508896 | 32 |
| GCCAA | 554950 | 2.1665668 | 40.958107 | 34 |
| CACGC | 384355 | 2.127672 | 57.756897 | 31 |
| TCTCG | 501840 | 1.982199 | 41.86822 | 41 |
| TCGGA | 486970 | 1.9816978 | 44.460773 | 3 |
| AGAGC | 484555 | 1.9604077 | 44.032307 | 8 |
| AGCAC | 499855 | 1.9514719 | 42.467052 | 10 |
| CCAGT | 488925 | 1.9199609 | 41.540936 | 26 |
| ATCGG | 467280 | 1.9015704 | 44.258236 | 2 |
| GCACA | 486645 | 1.899899 | 42.302074 | 11 |
| CACAC | 492180 | 1.8542024 | 40.679714 | 12 |
| TCTGA | 663835 | 1.8492185 | 30.490904 | 18 |
| GTCTG | 445410 | 1.8231695 | 43.89014 | 17 |
| GATCG | 439870 | 1.7900268 | 43.99035 | 1 |
| ATGCC | 455110 | 1.7871724 | 41.3844 | 47 |
| CGTCT | 444225 | 1.7546276 | 42.223507 | 16 |
| AAGAG | 612260 | 1.7469697 | 31.443863 | 7 |
| GTCAC | 444035 | 1.7436821 | 41.562286 | 29 |
| CTCGT | 438745 | 1.7329823 | 41.468243 | 42 |
| CACGT | 435875 | 1.7116387 | 42.04697 | 14 |
| TGAAC | 615925 | 1.7057841 | 30.072319 | 20 |
| TCACG | 430880 | 1.6920238 | 41.30307 | 30 |
| ACGTC | 424150 | 1.6655958 | 42.001053 | 15 |
| ACTCC | 436960 | 1.6557955 | 40.02457 | 23 |
| CCAAT | 617350 | 1.6498431 | 28.299595 | 35 |
| CAGTC | 417990 | 1.6414059 | 41.40169 | 27 |
| ACACG | 415835 | 1.6234516 | 41.874054 | 13 |
| GAACT | 554340 | 1.5352265 | 29.85965 | 21 |
| AACTC | 571875 | 1.5283129 | 28.734411 | 22 |
| ATCTC | 565430 | 1.5199239 | 28.567636 | 40 |
| AGTCA | 503685 | 1.3949391 | 29.47961 | 28 |
| AATAT | 961650 | 1.2857375 | 14.683524 | 37 |
| TCGTA | 449060 | 1.2509284 | 29.29176 | 43 |
| TATGC | 436425 | 1.2157316 | 29.272171 | 46 |
| CAATA | 643480 | 1.212809 | 20.034342 | 36 |
| GTATG | 415930 | 1.2006971 | 30.308231 | 45 |
| CGTAT | 418420 | 1.1655759 | 29.238111 | 44 |
| ATATC | 601385 | 1.1400968 | 20.122032 | 38 |
| TATCT | 515560 | 0.9831056 | 20.139376 | 39 |