![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW210_ACAGTG_L003_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 4222456 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
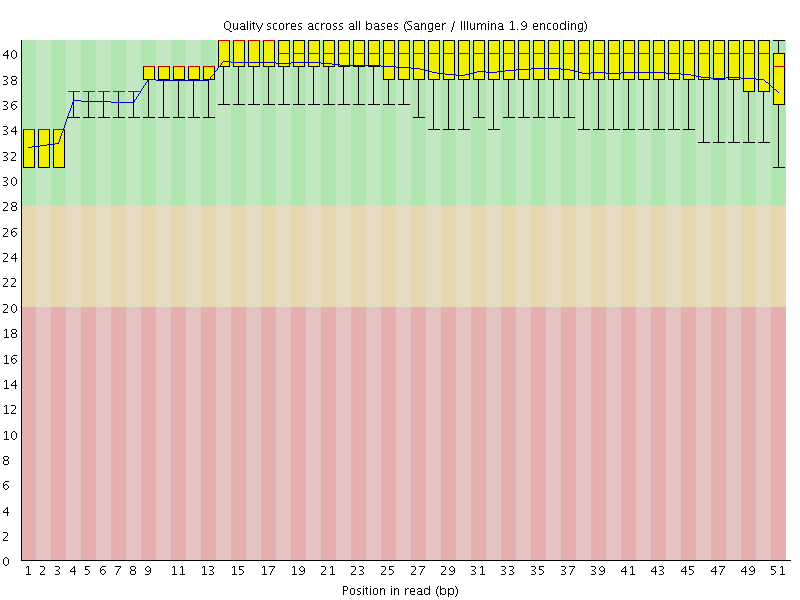
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
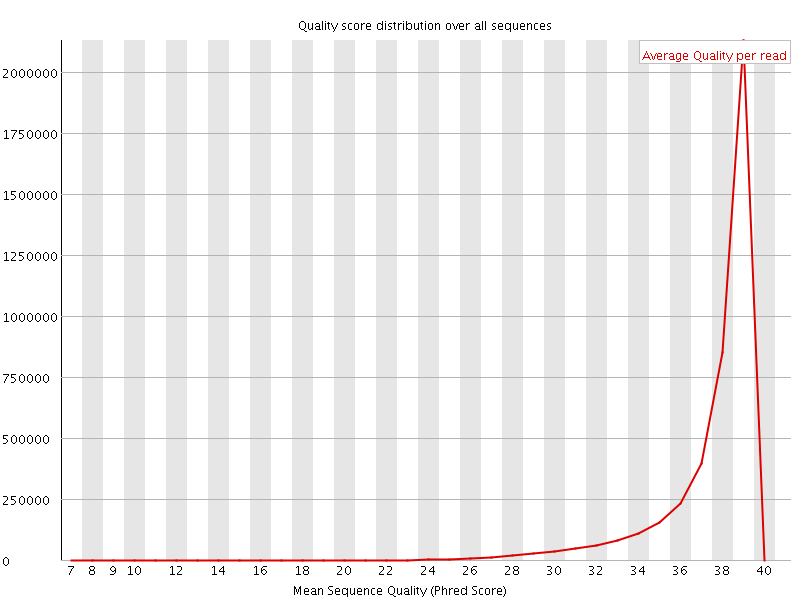
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
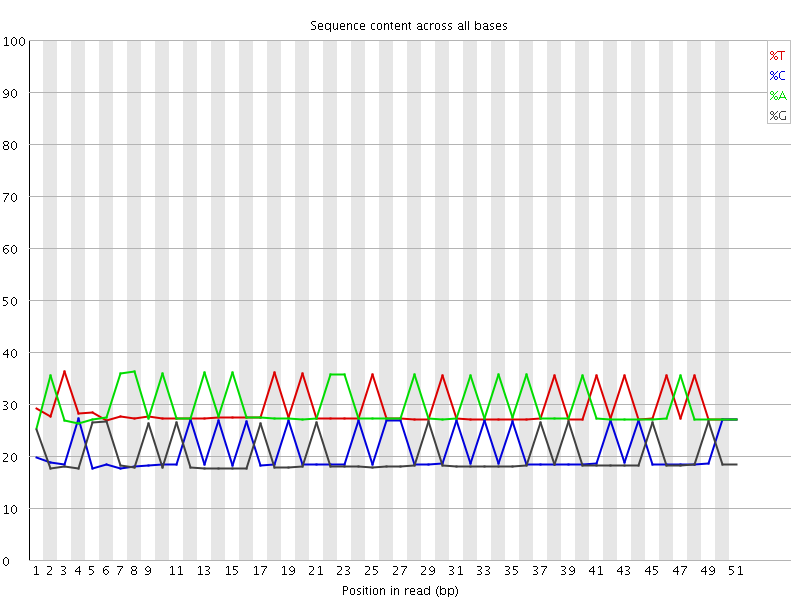
![[WARN]](Icons/warning.png) Per base GC content
Per base GC content
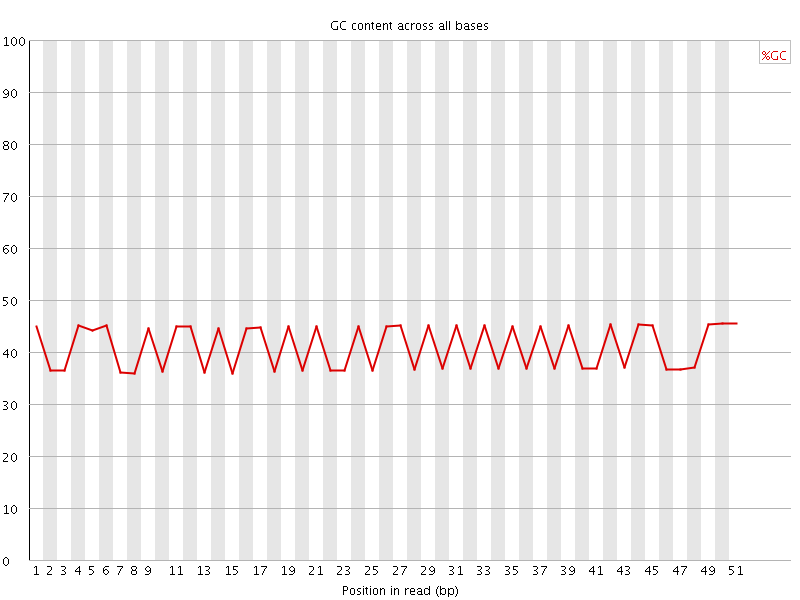
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
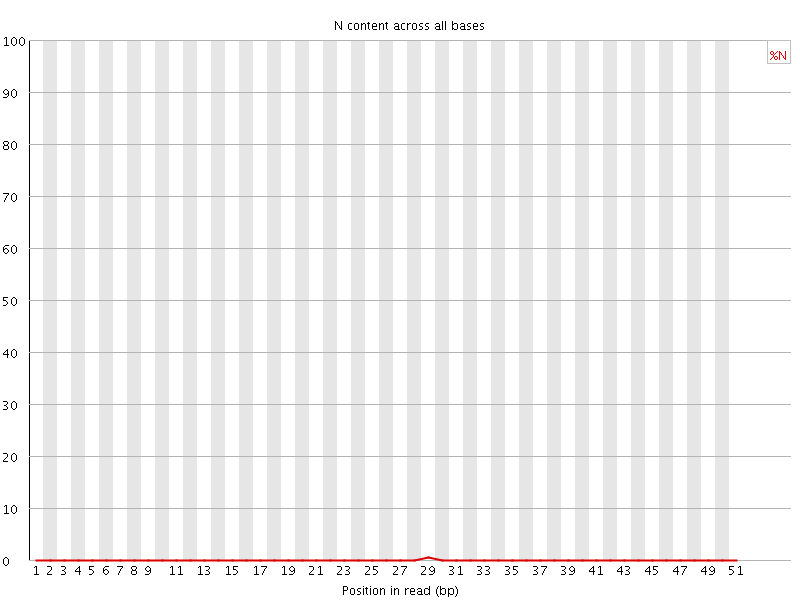
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
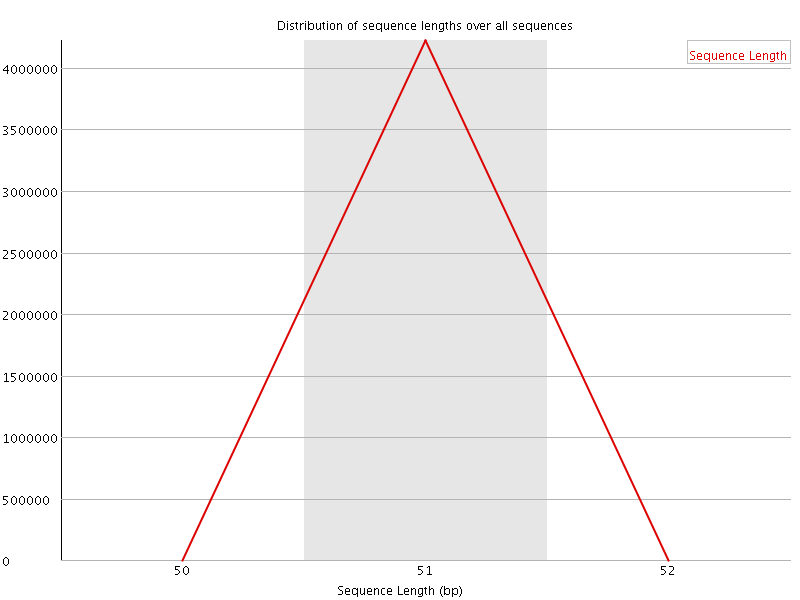
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
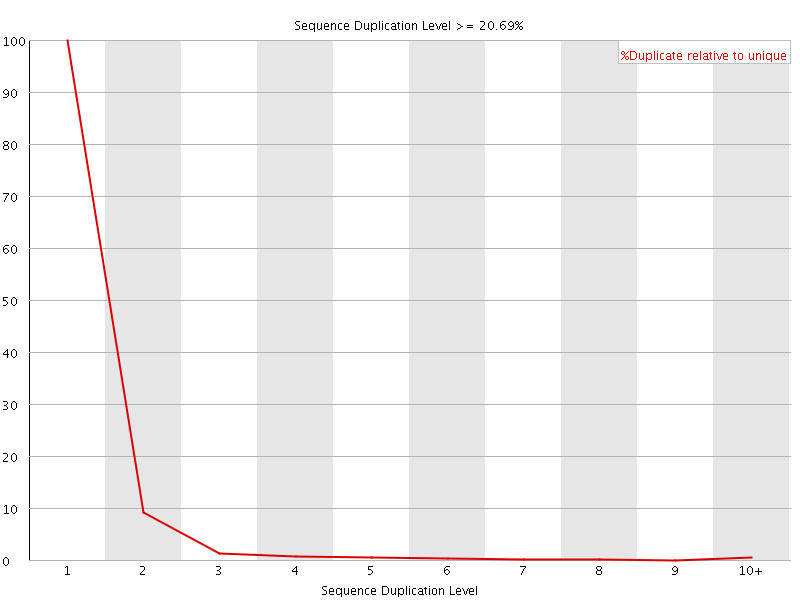
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACACAGTGATCTCGTATGCC | 321247 | 7.608060332659476 | TruSeq Adapter, Index 5 (100% over 51bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
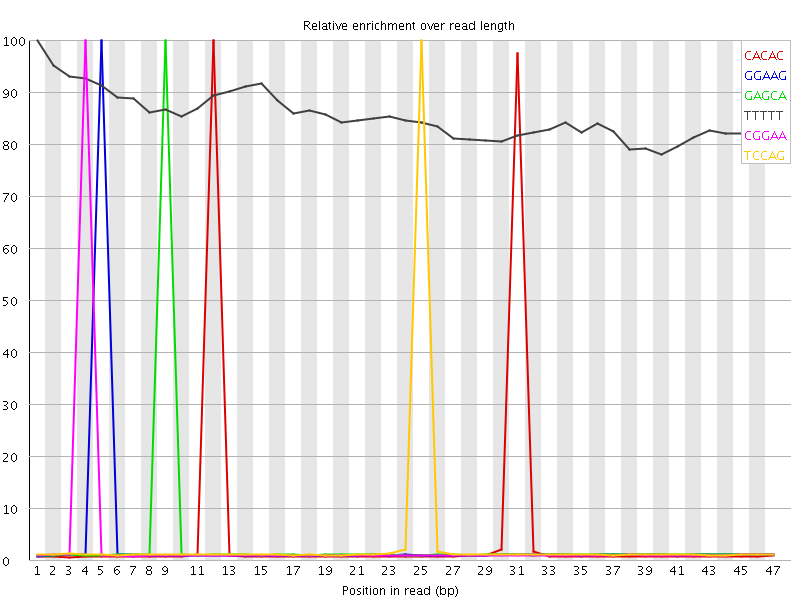
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CACAC | 880270 | 5.3822885 | 105.16645 | 12 |
| GGAAG | 566590 | 3.9750345 | 122.42379 | 5 |
| GAGCA | 573310 | 3.8419755 | 116.19314 | 9 |
| TTTTT | 1542815 | 3.723958 | 4.367069 | 1 |
| CGGAA | 555440 | 3.7222216 | 116.69973 | 4 |
| TCCAG | 565395 | 3.6903594 | 110.64424 | 25 |
| CTCCA | 580870 | 3.6215014 | 105.59225 | 24 |
| TCGGA | 513760 | 3.5106208 | 118.796005 | 3 |
| CAGTG | 510285 | 3.486876 | 114.53764 | 35 |
| TCTCG | 521590 | 3.4713998 | 112.65553 | 41 |
| AGAGC | 513485 | 3.4410646 | 115.86747 | 8 |
| ATCGG | 499485 | 3.4130774 | 118.5618 | 2 |
| GTCTG | 482975 | 3.36517 | 119.31884 | 17 |
| AAAAA | 1535575 | 3.3625498 | 3.6263413 | 13 |
| AGCAC | 520265 | 3.330295 | 110.611664 | 10 |
| GAAGA | 699710 | 3.3299065 | 83.47181 | 6 |
| GCACA | 513840 | 3.2891676 | 110.43133 | 11 |
| CCAGT | 503595 | 3.2869883 | 110.06875 | 26 |
| GATCG | 479640 | 3.2774727 | 118.07816 | 1 |
| CACAG | 507650 | 3.2495441 | 107.82862 | 33 |
| CTCGT | 486260 | 3.2362635 | 112.20351 | 42 |
| CGTCT | 484540 | 3.2248166 | 113.691765 | 16 |
| CTGAA | 686245 | 3.1808617 | 80.09047 | 19 |
| ATGCC | 487135 | 3.1795528 | 110.21083 | 47 |
| CACGT | 485170 | 3.1667273 | 112.068474 | 14 |
| GTCAC | 482505 | 3.1493328 | 110.241554 | 29 |
| ACGTC | 472355 | 3.0830832 | 111.67033 | 15 |
| CAGTC | 468335 | 3.0568445 | 109.69074 | 27 |
| ACACG | 474925 | 3.0400665 | 109.966965 | 13 |
| ACTCC | 484890 | 3.0231032 | 105.6613 | 23 |
| TCTGA | 611035 | 2.8879542 | 81.51807 | 18 |
| AAGAG | 584220 | 2.7802913 | 82.87184 | 7 |
| TGAAC | 580265 | 2.6896265 | 79.54179 | 20 |
| AGTGA | 551625 | 2.6768038 | 81.62916 | 36 |
| GTGAT | 533200 | 2.638283 | 83.14979 | 37 |
| TGATC | 555305 | 2.624556 | 79.28057 | 38 |
| ACAGT | 553950 | 2.5676517 | 77.78335 | 34 |
| TCACA | 578965 | 2.5633678 | 74.824524 | 30 |
| GAACT | 548605 | 2.5428774 | 79.228264 | 21 |
| ATCTC | 552140 | 2.4926798 | 76.27419 | 40 |
| AACTC | 560840 | 2.4831192 | 75.58869 | 22 |
| GATCT | 525105 | 2.4818208 | 79.57039 | 39 |
| ACACA | 567490 | 2.4640992 | 73.39687 | 32 |
| AGTCA | 515850 | 2.391052 | 78.18044 | 28 |
| GTATG | 471730 | 2.3341284 | 83.49014 | 45 |
| TCGTA | 493210 | 2.3310742 | 79.874954 | 43 |
| TATGC | 483440 | 2.284898 | 79.72691 | 46 |
| CGTAT | 473485 | 2.2378476 | 79.75286 | 44 |