![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW209_TGACCA_L003_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 6596382 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
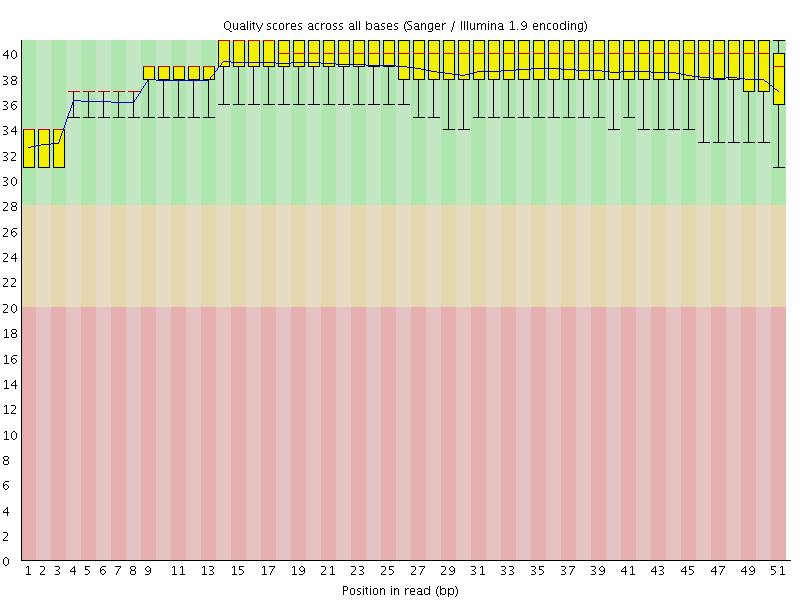
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
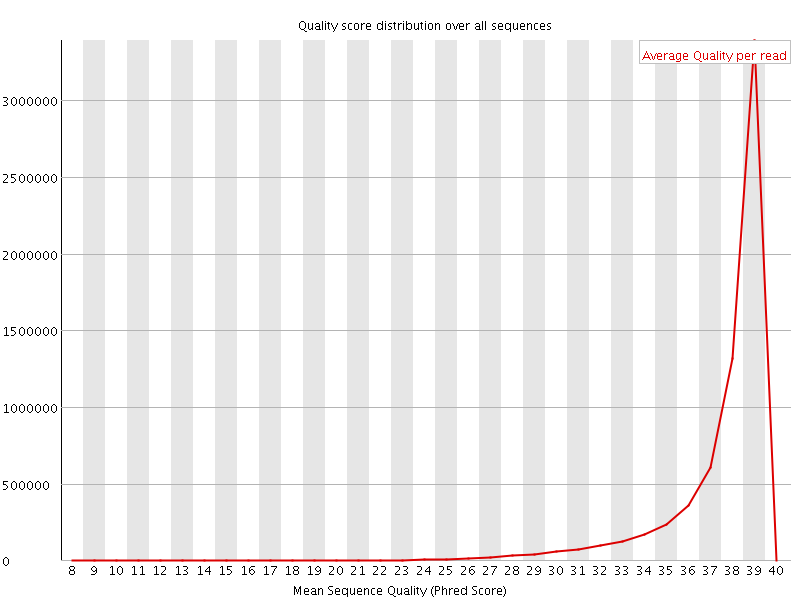
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
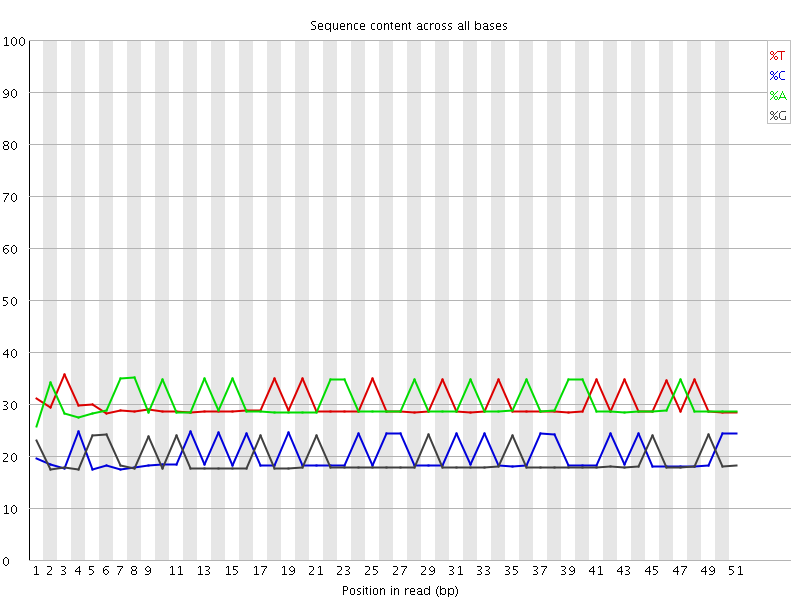
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
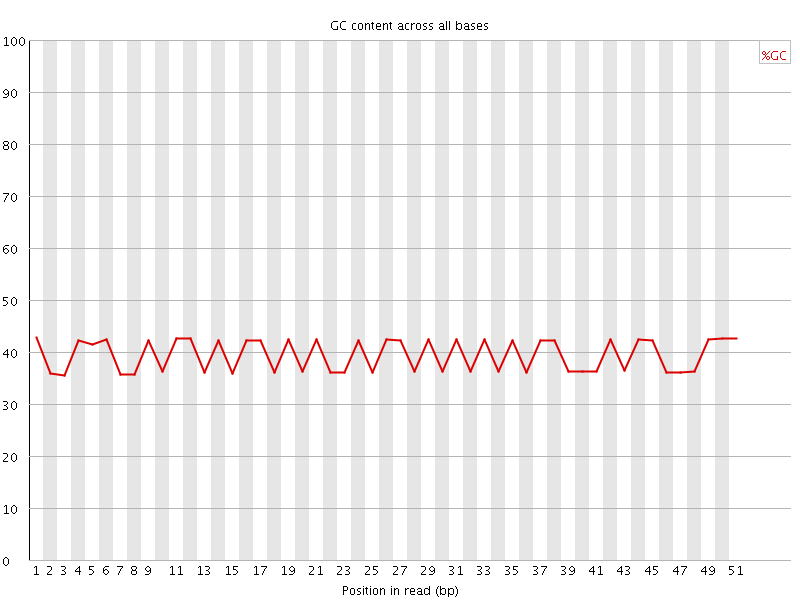
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
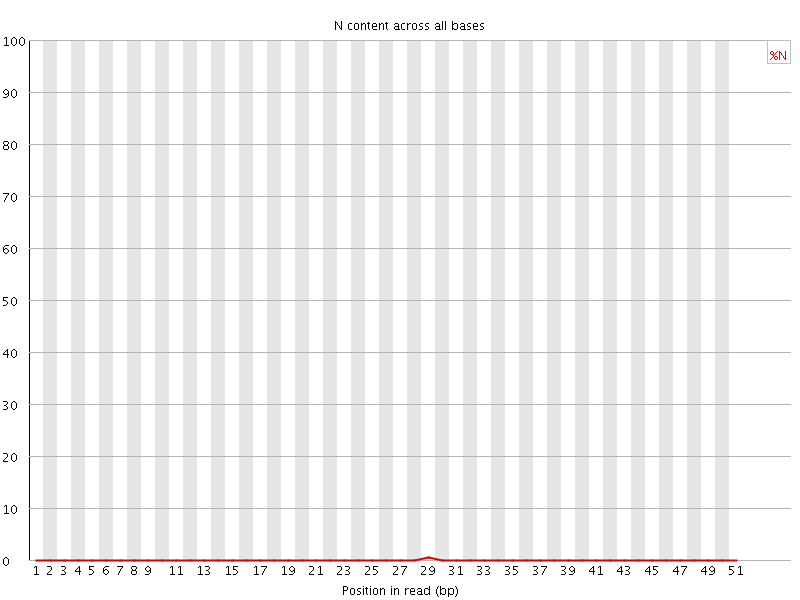
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
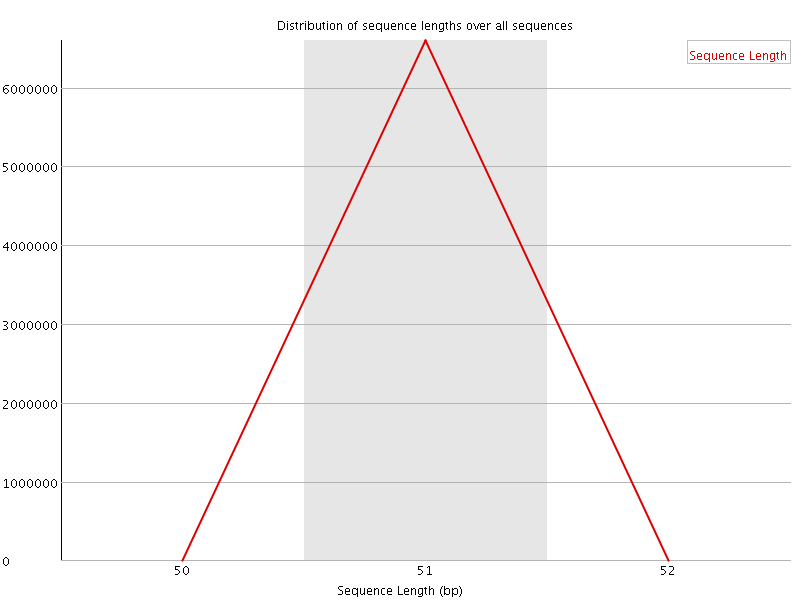
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
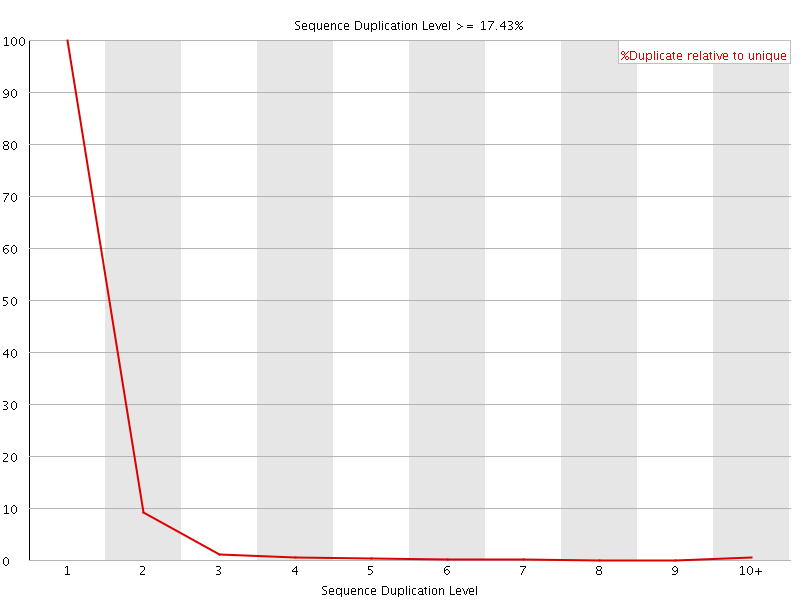
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACTGACCAATCTCGTATGCC | 359269 | 5.446455344763235 | TruSeq Adapter, Index 4 (100% over 51bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
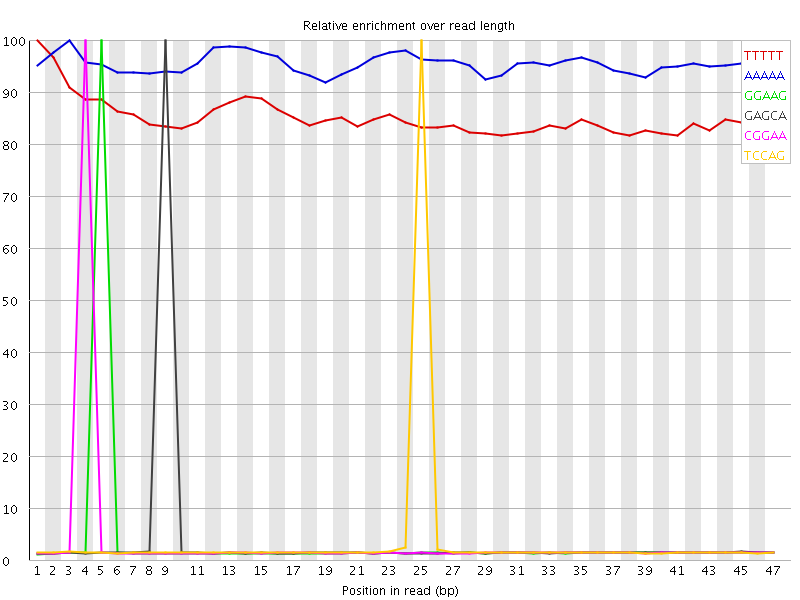
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 2913350 | 3.817884 | 4.4828944 | 1 |
| AAAAA | 2862065 | 3.6011858 | 3.7689846 | 3 |
| GGAAG | 718565 | 3.5128407 | 96.87403 | 5 |
| GAGCA | 726320 | 3.374974 | 91.71834 | 9 |
| CGGAA | 721895 | 3.3544126 | 91.986694 | 4 |
| TCCAG | 718665 | 3.2000139 | 85.77631 | 25 |
| CTCCA | 735245 | 3.1117704 | 81.305405 | 24 |
| TCGGA | 649155 | 3.041051 | 92.46857 | 3 |
| TCTCG | 658105 | 2.9542913 | 85.68689 | 41 |
| AGAGC | 632935 | 2.941044 | 91.38452 | 8 |
| GAAGA | 941705 | 2.9284093 | 62.201115 | 6 |
| ATCGG | 620640 | 2.9074686 | 92.14413 | 2 |
| CACTG | 642740 | 2.8619409 | 85.40792 | 31 |
| GCACA | 646505 | 2.8553834 | 86.52255 | 11 |
| CTGAA | 946660 | 2.82094 | 58.800365 | 19 |
| GTCTG | 595490 | 2.8124354 | 91.61408 | 17 |
| AGCAC | 636370 | 2.810621 | 86.544334 | 10 |
| CCAGT | 621575 | 2.7676992 | 85.07643 | 26 |
| CACAC | 651410 | 2.7346208 | 81.75737 | 12 |
| GATCG | 582860 | 2.7304833 | 91.6179 | 1 |
| GTCAC | 603900 | 2.6889975 | 85.48583 | 29 |
| CTGAC | 601415 | 2.6779325 | 84.72142 | 33 |
| CTCGT | 594390 | 2.6682694 | 84.87485 | 42 |
| CGTCT | 593895 | 2.6660469 | 86.78175 | 16 |
| CACGT | 596755 | 2.657183 | 86.42223 | 14 |
| ATGCC | 590800 | 2.630667 | 84.26386 | 47 |
| GACCA | 590940 | 2.6099725 | 82.88874 | 35 |
| TGACC | 576980 | 2.5691304 | 83.783905 | 34 |
| ACGTC | 574530 | 2.558221 | 86.04721 | 15 |
| ACACG | 577480 | 2.5505247 | 85.77392 | 13 |
| CAGTC | 571190 | 2.543349 | 84.826996 | 27 |
| ACTCC | 592005 | 2.505537 | 81.17766 | 23 |
| TCTGA | 817630 | 2.4563456 | 58.91027 | 18 |
| AAGAG | 752425 | 2.3398077 | 61.660877 | 7 |
| TGAAC | 767220 | 2.2862294 | 58.20269 | 20 |
| ACCAA | 788120 | 2.2141626 | 53.072483 | 36 |
| ACTGA | 729475 | 2.1737533 | 57.43794 | 32 |
| CCAAT | 764120 | 2.1642702 | 53.468758 | 37 |
| GAACT | 711660 | 2.1206663 | 57.968903 | 21 |
| TCACT | 736150 | 2.102079 | 54.987225 | 30 |
| AACTC | 728185 | 2.0624893 | 54.976334 | 22 |
| ATCTC | 709575 | 2.0261939 | 54.505707 | 40 |
| CAATC | 707790 | 2.004723 | 53.368027 | 38 |
| AGTCA | 661960 | 1.9725661 | 57.18139 | 28 |
| GTATG | 579765 | 1.83246 | 59.696526 | 45 |
| TATGC | 591585 | 1.7772552 | 56.664455 | 46 |
| TCGTA | 589065 | 1.7696846 | 56.837173 | 43 |
| CGTAT | 566735 | 1.7026001 | 56.75026 | 44 |
| AATCT | 839900 | 1.6050359 | 36.556973 | 39 |