![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW207_CGATGT_L003_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 4768014 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
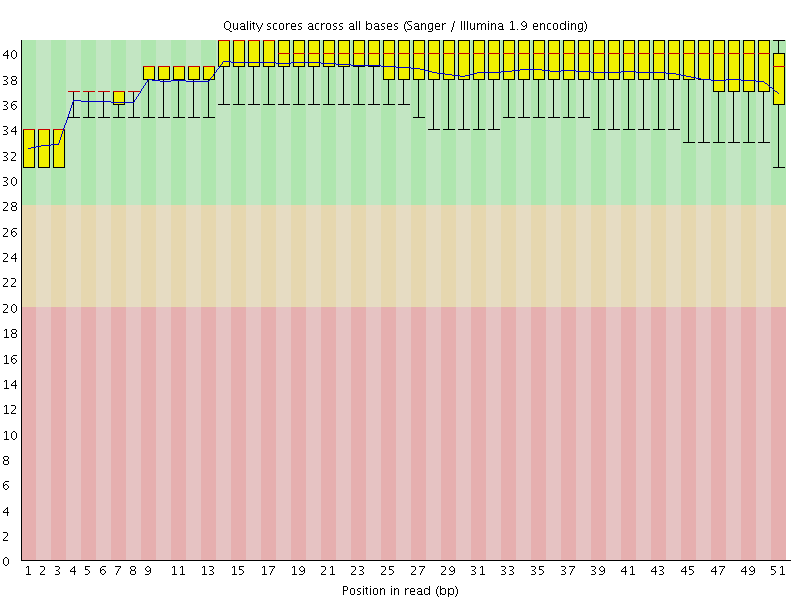
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
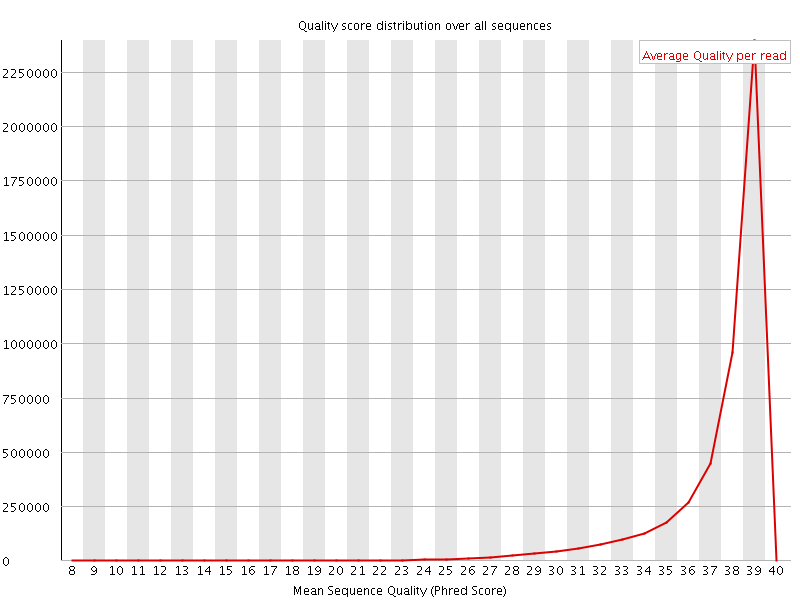
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
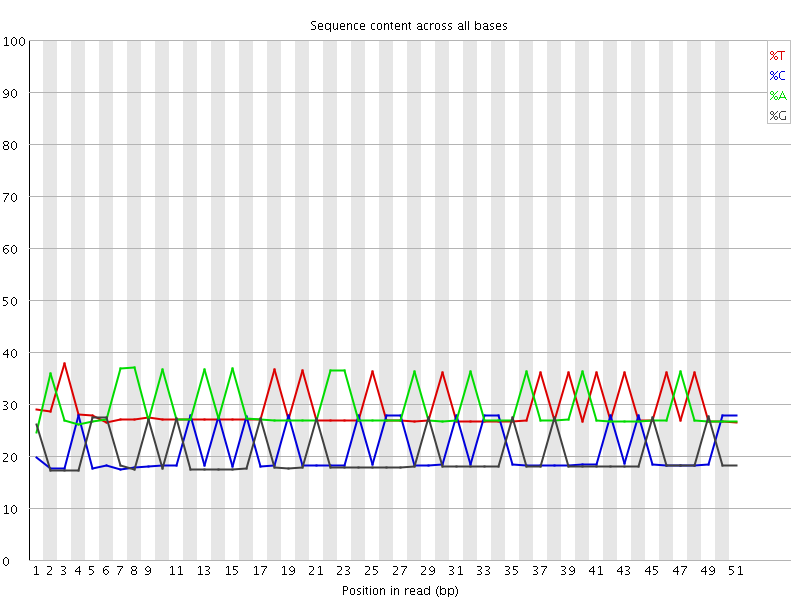
![[WARN]](Icons/warning.png) Per base GC content
Per base GC content
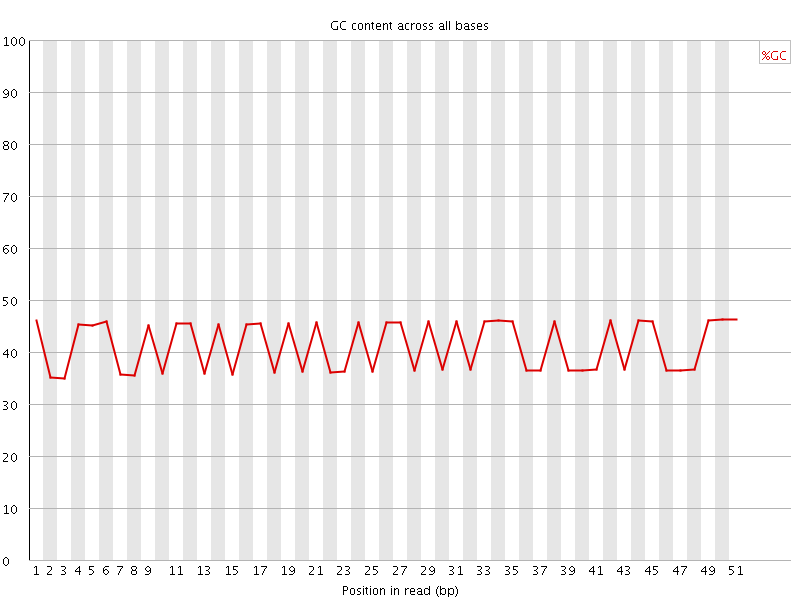
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
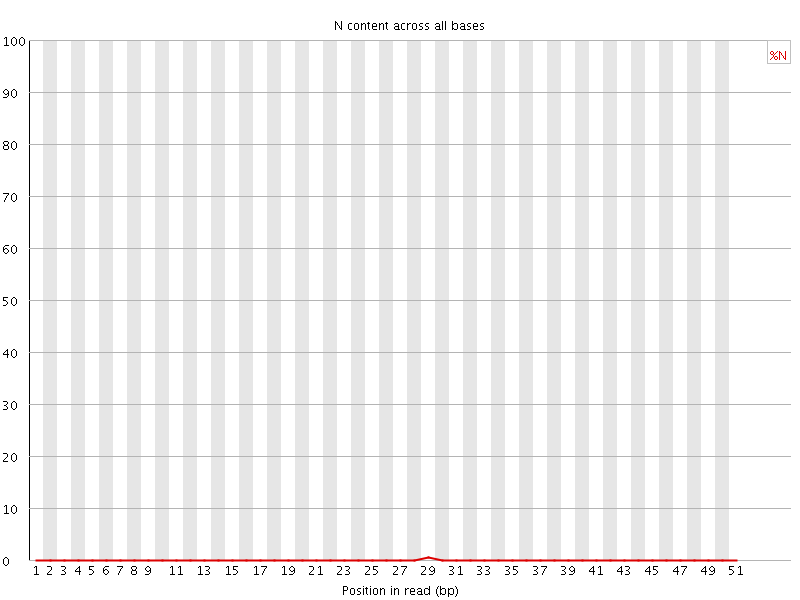
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
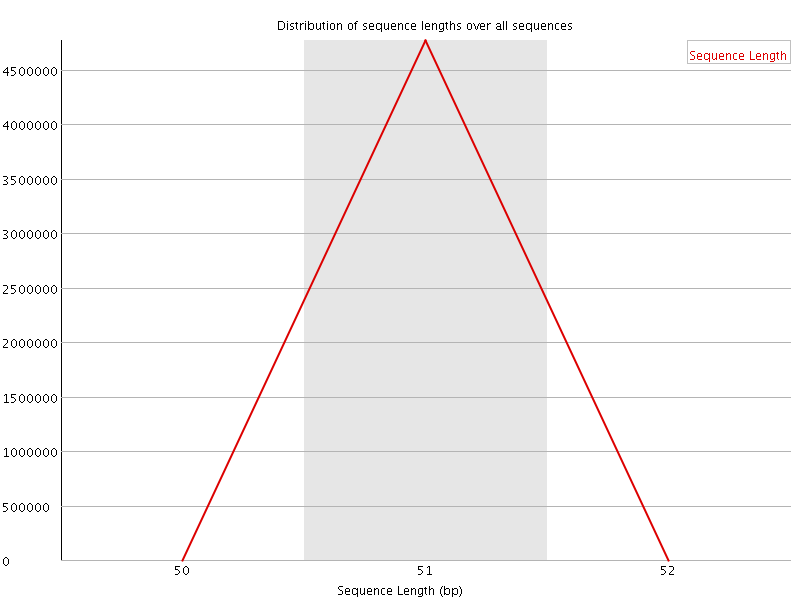
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
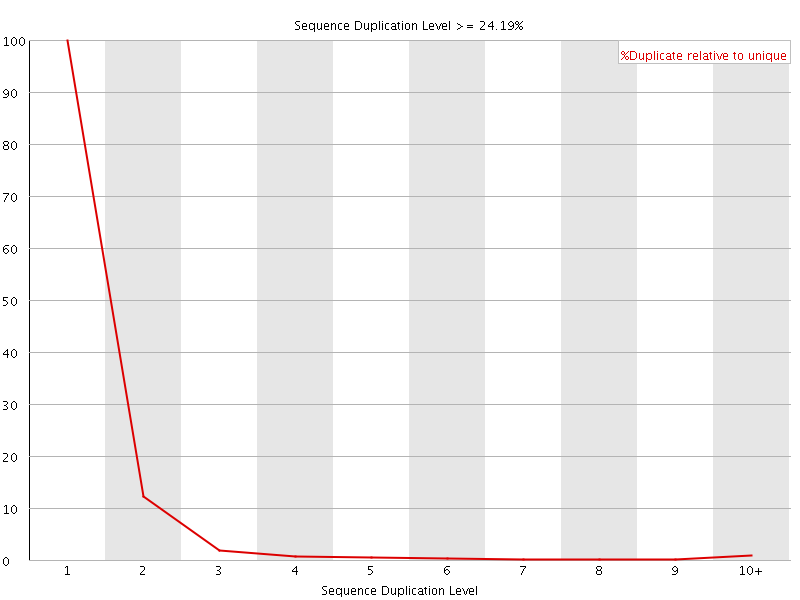
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCGATGTATCTCGTATGCC | 411945 | 8.639760705400613 | TruSeq Adapter, Index 2 (100% over 51bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
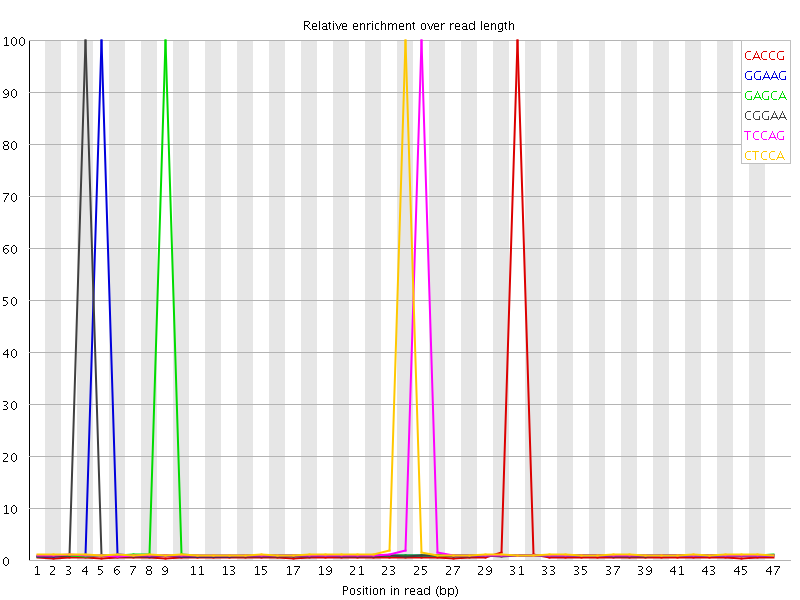
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CACCG | 585935 | 4.6220145 | 167.11206 | 31 |
| GGAAG | 684000 | 4.2699184 | 136.28952 | 5 |
| GAGCA | 690630 | 4.1133647 | 129.42252 | 9 |
| CGGAA | 667880 | 3.9778662 | 129.91469 | 4 |
| TCCAG | 686995 | 3.9376812 | 122.46284 | 25 |
| CTCCA | 701770 | 3.8376915 | 116.73252 | 24 |
| TCGGA | 624395 | 3.7510953 | 130.88385 | 3 |
| CGATG | 621890 | 3.7360463 | 127.06402 | 34 |
| AGAGC | 622155 | 3.70553 | 129.04918 | 8 |
| TCTCG | 633070 | 3.660038 | 122.8513 | 41 |
| ATCGG | 607485 | 3.6495073 | 130.60178 | 2 |
| GTCTG | 593395 | 3.5957499 | 130.205 | 17 |
| TCACC | 655660 | 3.5855348 | 116.35373 | 30 |
| AGCAC | 630380 | 3.5821395 | 123.0295 | 10 |
| GAAGA | 832690 | 3.5726733 | 94.06063 | 6 |
| GCACA | 623470 | 3.542873 | 122.89342 | 11 |
| GATCG | 586460 | 3.5231981 | 130.14026 | 1 |
| CCAGT | 614450 | 3.521871 | 121.8464 | 26 |
| TTTTT | 1651655 | 3.4950778 | 4.4049225 | 1 |
| CCGAT | 600890 | 3.4441488 | 121.259964 | 33 |
| CGTCT | 593940 | 3.4338114 | 124.06934 | 16 |
| CTCGT | 589970 | 3.4108596 | 122.05903 | 42 |
| CACAC | 628480 | 3.4073741 | 116.95964 | 12 |
| ATGCC | 593260 | 3.400416 | 120.82965 | 47 |
| CACGT | 592725 | 3.3973494 | 123.2801 | 14 |
| GTCAC | 591465 | 3.3901274 | 121.98324 | 29 |
| CTGAA | 819215 | 3.382541 | 89.617455 | 19 |
| AAAAA | 1659850 | 3.3641224 | 3.685223 | 13 |
| ACCGA | 590760 | 3.3569984 | 120.18774 | 32 |
| ACGTC | 579995 | 3.3243842 | 123.085365 | 15 |
| CAGTC | 573675 | 3.2881596 | 121.46029 | 27 |
| ACACG | 578585 | 3.287814 | 122.279755 | 13 |
| ACTCC | 591870 | 3.2366936 | 116.68693 | 23 |
| TCTGA | 735570 | 3.0634873 | 90.056625 | 18 |
| AAGAG | 700970 | 3.007526 | 93.42101 | 7 |
| TGAAC | 701540 | 2.8966606 | 88.964745 | 20 |
| GATGT | 647285 | 2.8255262 | 92.32848 | 35 |
| GAACT | 664135 | 2.7422152 | 88.813324 | 21 |
| AACTC | 676605 | 2.6654384 | 84.67586 | 22 |
| ATCTC | 670380 | 2.663799 | 84.496544 | 40 |
| AGTCA | 629480 | 2.5991247 | 87.799416 | 28 |
| GTATG | 576650 | 2.5171902 | 91.96637 | 45 |
| TCGTA | 601480 | 2.505032 | 87.87027 | 43 |
| GTATC | 590715 | 2.4601982 | 87.74413 | 38 |
| TATGC | 587410 | 2.4464338 | 87.600975 | 46 |
| CGTAT | 578960 | 2.411241 | 87.75867 | 44 |
| TGTAT | 659495 | 1.9957619 | 64.22498 | 37 |
| ATGTA | 654155 | 1.9625962 | 63.462585 | 36 |
| TATCT | 639350 | 1.8459682 | 61.02652 | 39 |