![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW204_GAGTGG_L003_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 7171554 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
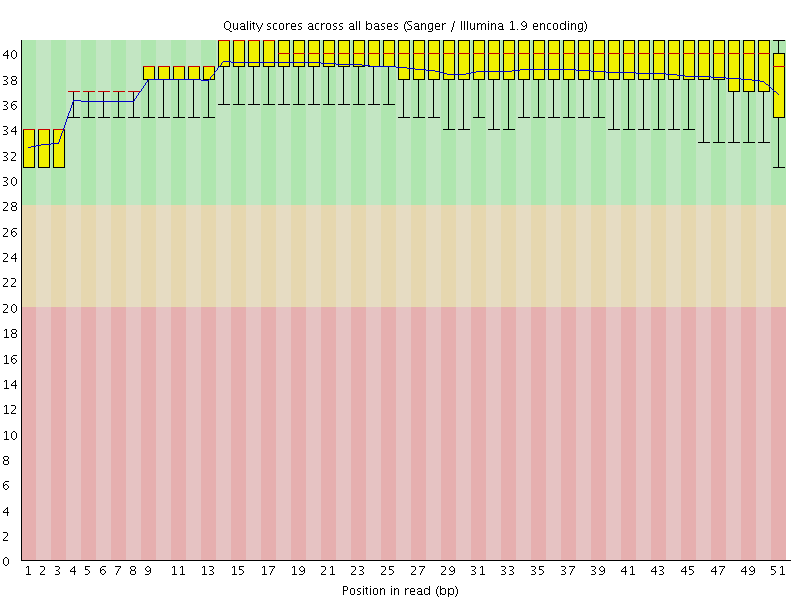
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
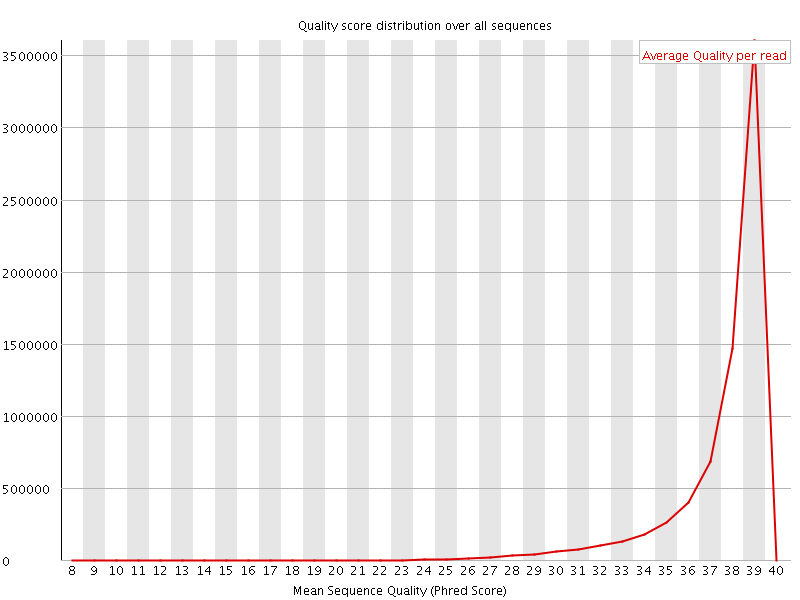
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
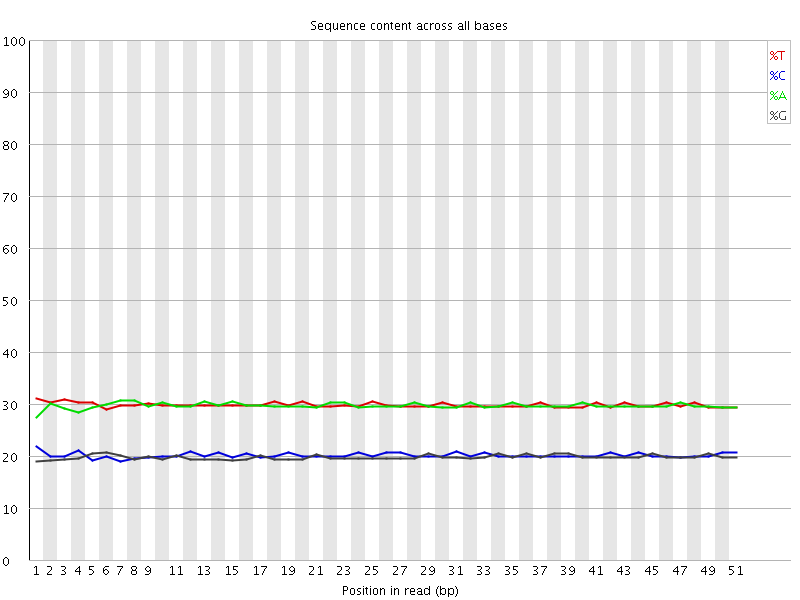
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
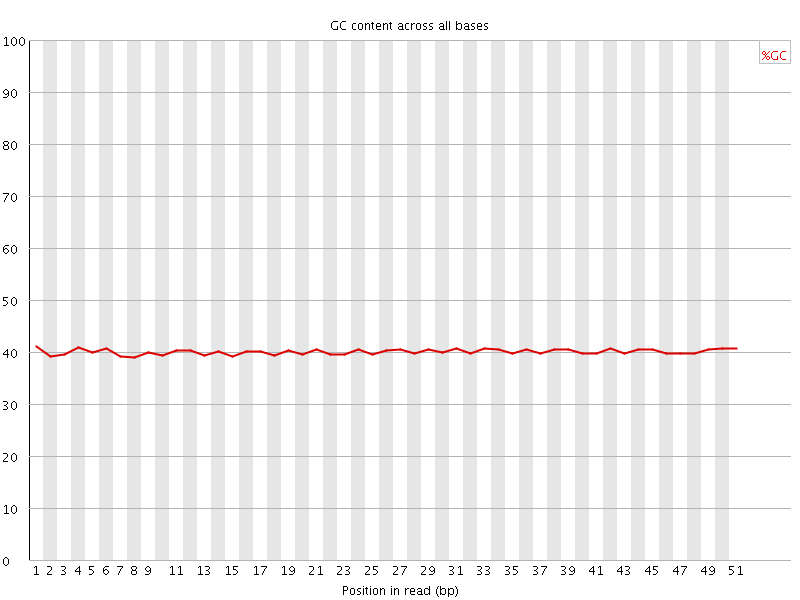
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
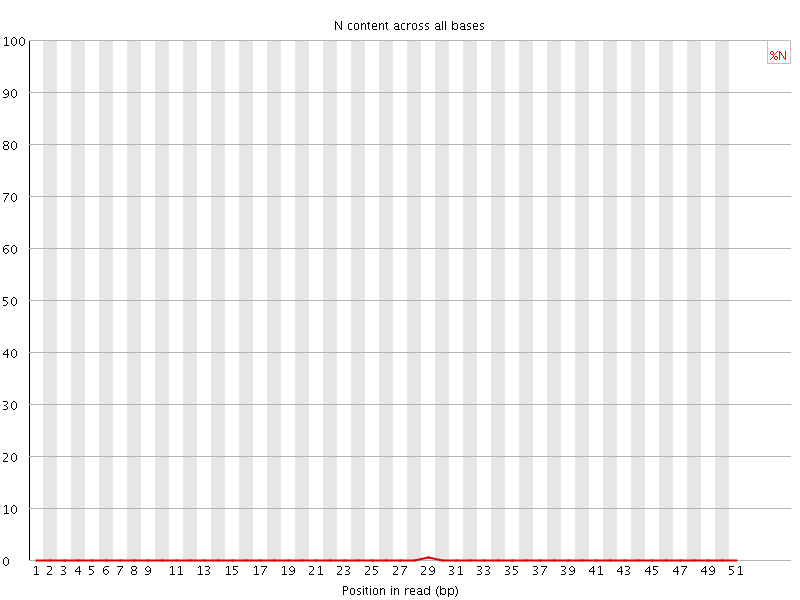
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
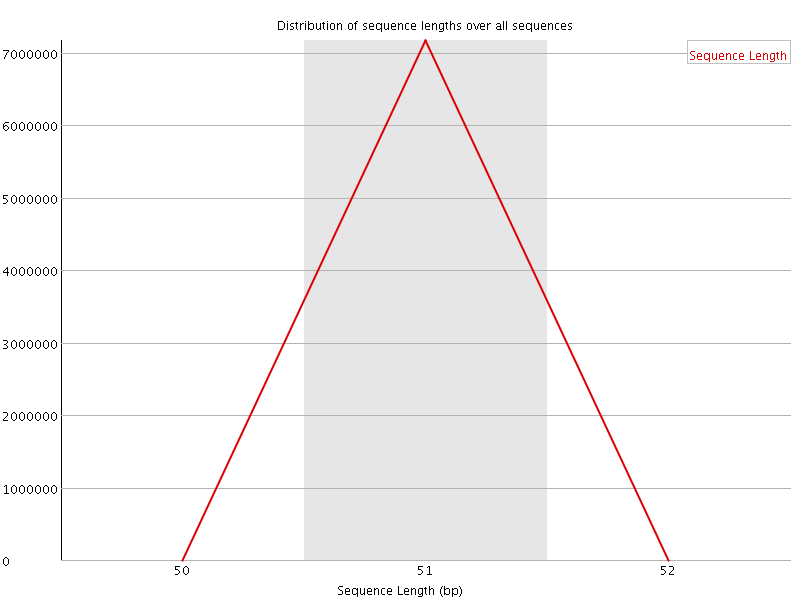
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
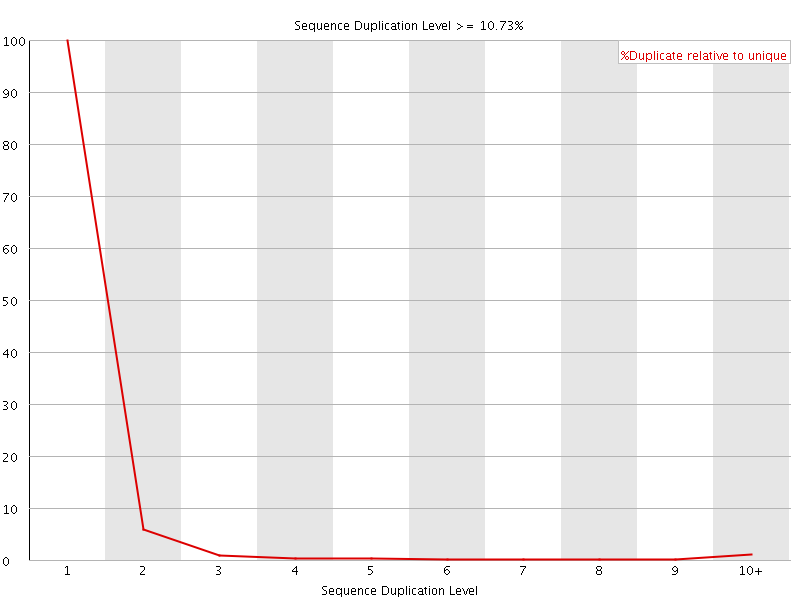
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGAGTGGATCTCGTATGCC | 52828 | 0.7366325345943152 | TruSeq Adapter, Index 7 (97% over 36bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
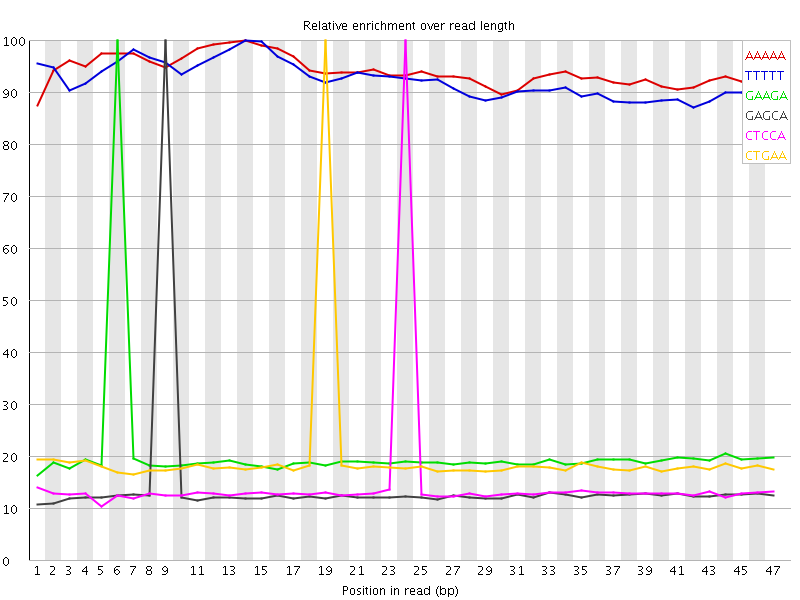
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 2838875 | 3.5727818 | 3.7938952 | 14 |
| TTTTT | 2882360 | 3.540719 | 3.8331857 | 14 |
| GAAGA | 711605 | 2.0107007 | 9.735145 | 6 |
| GAGCA | 450090 | 1.8683181 | 13.129313 | 9 |
| CTCCA | 467075 | 1.8546945 | 12.591306 | 24 |
| CTGAA | 663315 | 1.8287055 | 9.278696 | 19 |
| GGAAG | 427665 | 1.8106575 | 13.483749 | 5 |
| TCCAG | 429440 | 1.7392795 | 12.673235 | 25 |
| CGGAA | 416105 | 1.7272468 | 13.31934 | 4 |
| GTGGA | 367535 | 1.5485595 | 12.360244 | 36 |
| TCTCG | 382955 | 1.5435164 | 12.11295 | 41 |
| TCGGA | 350475 | 1.4477884 | 12.918471 | 3 |
| TCTGA | 518965 | 1.4238315 | 8.84415 | 18 |
| CACAC | 354050 | 1.4127123 | 12.301166 | 12 |
| AGAGC | 334815 | 1.389813 | 12.742441 | 8 |
| AGCAC | 332570 | 1.3534849 | 12.469252 | 10 |
| GCACA | 332070 | 1.3514498 | 12.425301 | 11 |
| AAGAG | 474780 | 1.3415315 | 9.07844 | 7 |
| AGTGG | 311545 | 1.3126532 | 12.182248 | 35 |
| ATCGG | 313255 | 1.2940352 | 12.637236 | 2 |
| TGAAC | 456385 | 1.2582164 | 8.716836 | 20 |
| GATCG | 301480 | 1.2453934 | 12.493417 | 1 |
| GGATC | 298055 | 1.231245 | 11.875479 | 38 |
| CACGA | 301720 | 1.2279322 | 11.908948 | 31 |
| CCAGT | 303030 | 1.227305 | 12.091086 | 26 |
| ACGAG | 293585 | 1.2186676 | 12.04449 | 32 |
| CTCGT | 301135 | 1.2137375 | 11.75605 | 42 |
| CGTCT | 293110 | 1.1813923 | 12.133355 | 16 |
| GAGTG | 277750 | 1.1702623 | 12.098107 | 34 |
| TGGAT | 414145 | 1.1589218 | 8.351778 | 37 |
| CACGT | 284105 | 1.1506567 | 12.135171 | 14 |
| GTCAC | 282020 | 1.1422123 | 11.832567 | 29 |
| ATCTC | 423755 | 1.1398672 | 8.108361 | 40 |
| AACTC | 421290 | 1.1387383 | 8.482833 | 22 |
| GTCTG | 274795 | 1.1296749 | 12.362573 | 17 |
| ATGCC | 277320 | 1.1231768 | 11.595638 | 47 |
| GAACT | 402545 | 1.1097839 | 8.566016 | 21 |
| ACTCC | 275185 | 1.092724 | 11.914914 | 23 |
| ACGTC | 268595 | 1.0878395 | 12.08101 | 15 |
| TCACG | 268520 | 1.0875357 | 11.721568 | 30 |
| ACACG | 263865 | 1.073871 | 12.170203 | 13 |
| CAGTC | 254915 | 1.032434 | 11.856503 | 27 |
| CGAGT | 249650 | 1.0312873 | 11.751466 | 33 |
| GATCT | 368445 | 1.0108651 | 8.11469 | 39 |
| AGTCA | 340460 | 0.93862057 | 8.234989 | 28 |
| TCGTA | 290815 | 0.7978795 | 7.92621 | 43 |
| TATGC | 263805 | 0.72377497 | 7.799909 | 46 |
| GTATG | 247025 | 0.6912619 | 7.941355 | 45 |
| CGTAT | 249510 | 0.68455523 | 7.824402 | 44 |