![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW203_CGTACG_L003_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 7572129 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
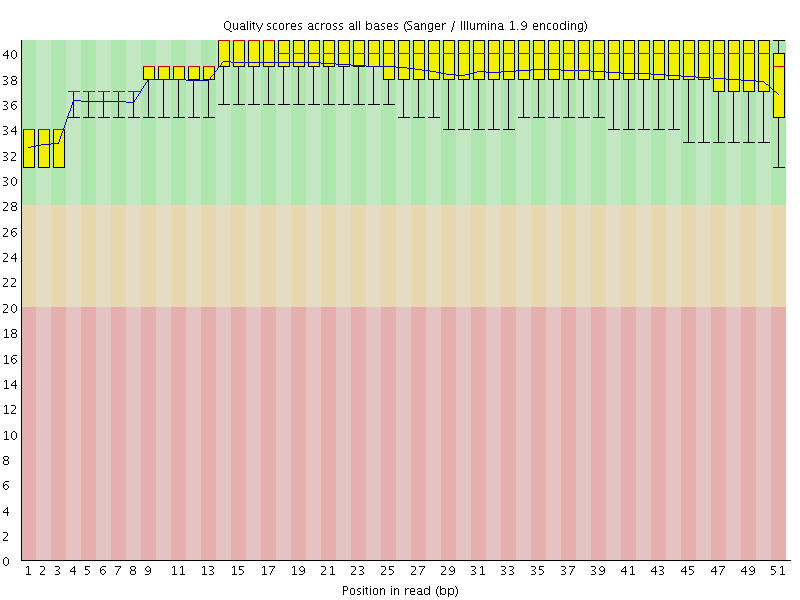
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
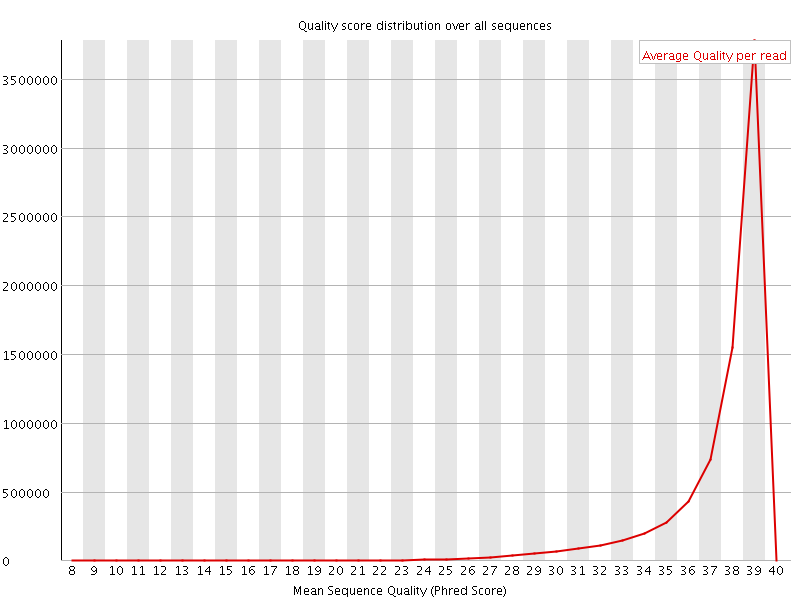
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
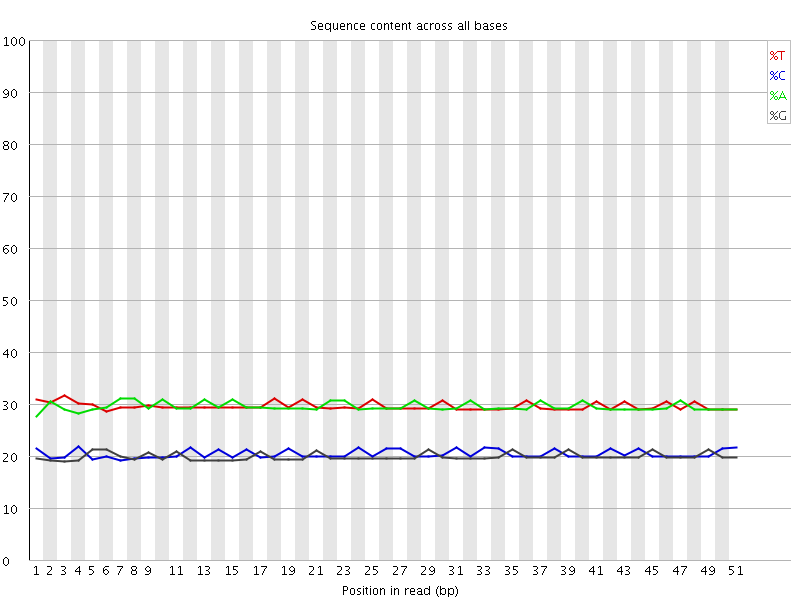
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
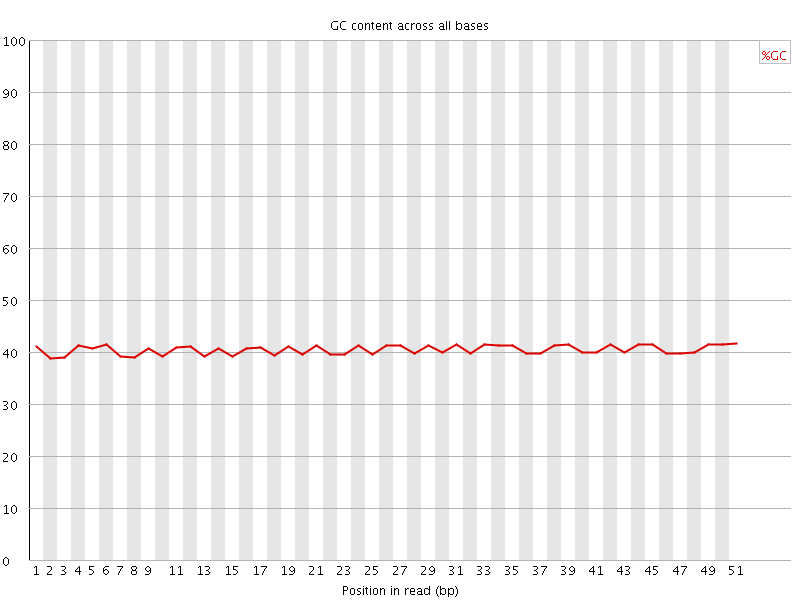
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
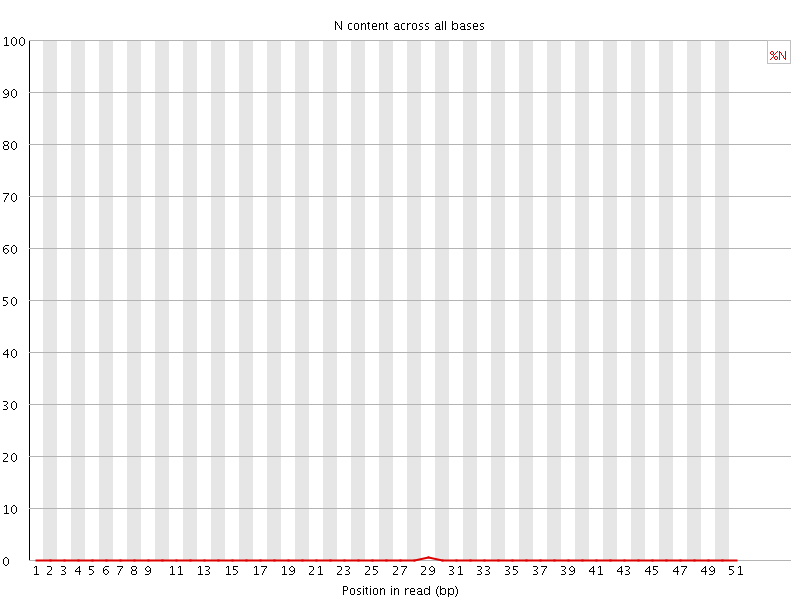
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
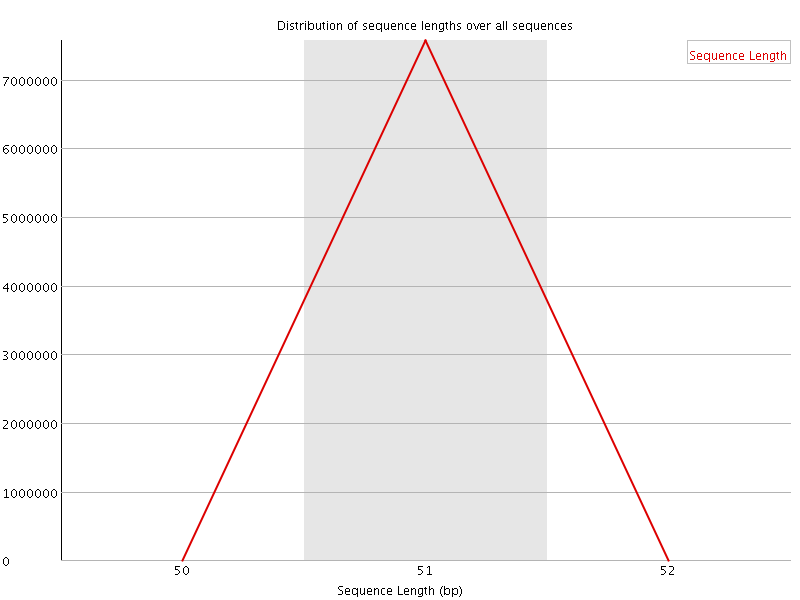
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
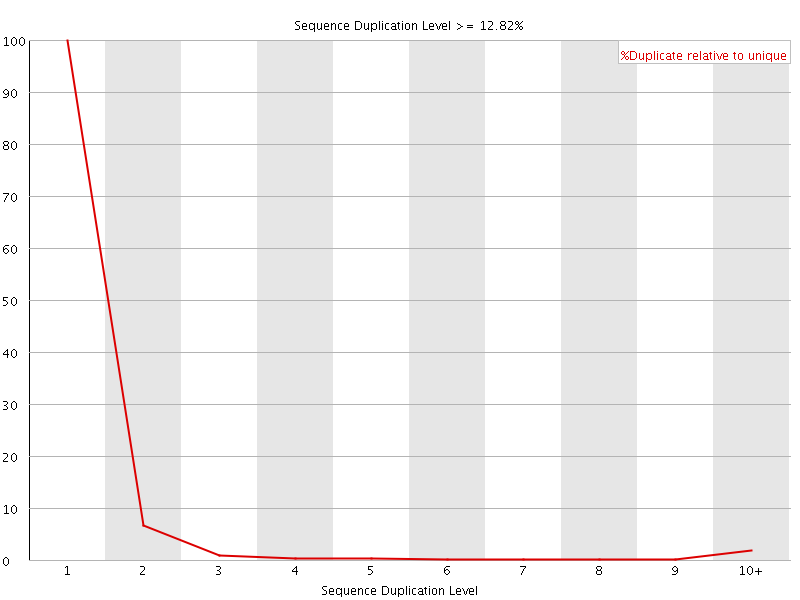
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCGTACGATCTCGTATGCC | 109232 | 1.4425533426596404 | TruSeq Adapter, Index 12 (97% over 36bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
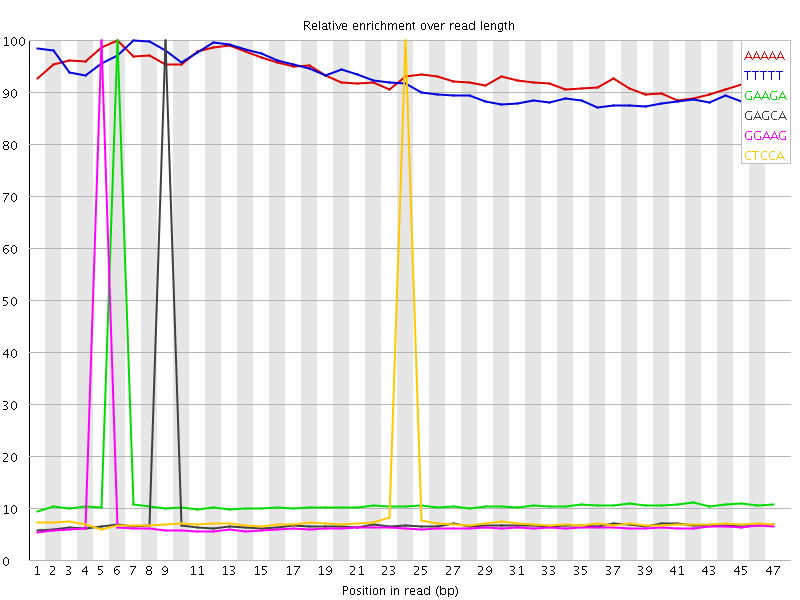
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 2848615 | 3.5035067 | 3.7469194 | 6 |
| TTTTT | 2874275 | 3.4716158 | 3.7576442 | 7 |
| GAAGA | 802765 | 2.160066 | 17.486029 | 6 |
| GAGCA | 535420 | 2.074947 | 23.90576 | 9 |
| GGAAG | 511445 | 2.0355518 | 24.749317 | 5 |
| CTCCA | 551585 | 2.019343 | 22.242867 | 24 |
| CTGAA | 743910 | 1.9420257 | 16.575417 | 19 |
| TCCAG | 512785 | 1.9279854 | 22.756977 | 25 |
| CGGAA | 491190 | 1.9035398 | 24.04587 | 4 |
| CACCG | 345440 | 1.8773624 | 31.387686 | 31 |
| TCTCG | 460285 | 1.7243367 | 21.907295 | 41 |
| TCACC | 452840 | 1.6578393 | 21.565971 | 30 |
| TCGGA | 427010 | 1.6488353 | 23.76493 | 3 |
| AGCAC | 422275 | 1.5934447 | 22.900759 | 10 |
| AGAGC | 408750 | 1.5840548 | 23.547302 | 8 |
| CACAC | 426270 | 1.56623 | 22.309021 | 12 |
| TCTGA | 598765 | 1.5574632 | 16.17041 | 18 |
| GCACA | 406810 | 1.5350878 | 22.797997 | 11 |
| ATCGG | 395060 | 1.5254653 | 23.533772 | 2 |
| GATCG | 393685 | 1.5201558 | 23.419312 | 1 |
| AAGAG | 552970 | 1.4879218 | 16.73494 | 7 |
| CGATC | 385100 | 1.4479114 | 21.463522 | 38 |
| CCAGT | 379625 | 1.4273262 | 22.149923 | 26 |
| TGAAC | 542970 | 1.4174587 | 16.007782 | 20 |
| CTCGT | 375210 | 1.4056258 | 21.510443 | 42 |
| CGTCT | 367465 | 1.3766111 | 22.354752 | 16 |
| GTCAC | 360535 | 1.3555511 | 22.002382 | 29 |
| ATGCC | 359060 | 1.3500054 | 21.524124 | 47 |
| CACGT | 357340 | 1.3435383 | 22.42163 | 14 |
| GTCTG | 348055 | 1.339103 | 22.923763 | 17 |
| ACTCC | 353030 | 1.2924366 | 21.69525 | 23 |
| ACGTC | 343495 | 1.2914835 | 22.342281 | 15 |
| ATCTC | 507430 | 1.2851882 | 14.847871 | 40 |
| AACTC | 504375 | 1.2820864 | 15.46483 | 22 |
| ACACG | 336495 | 1.2697558 | 22.523375 | 13 |
| GAACT | 482960 | 1.2607987 | 15.833691 | 21 |
| CAGTC | 328275 | 1.2342589 | 21.91439 | 27 |
| ACCGT | 314315 | 1.1817715 | 21.540665 | 32 |
| ACGAT | 451480 | 1.1786182 | 15.168547 | 37 |
| GATCT | 450905 | 1.1728607 | 15.069438 | 39 |
| AGTCA | 423235 | 1.1048826 | 15.415156 | 28 |
| CCGTA | 279980 | 1.0526778 | 21.332863 | 33 |
| TCGTA | 369355 | 0.96073896 | 14.941746 | 43 |
| TACGA | 360760 | 0.94178754 | 14.944161 | 36 |
| GTACG | 243400 | 0.9398528 | 21.851473 | 35 |
| CGTAC | 247600 | 0.9309344 | 21.192932 | 34 |
| TATGC | 341965 | 0.8894941 | 14.825828 | 46 |
| GTATG | 324815 | 0.8676974 | 15.2254305 | 45 |
| CGTAT | 328475 | 0.8544049 | 14.784767 | 44 |