![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW201_GTGGCC_L003_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 10825090 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
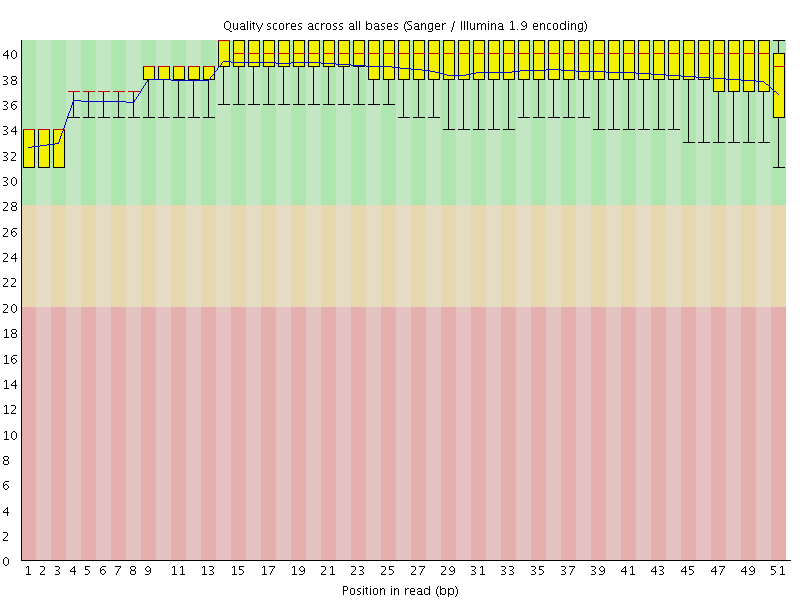
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
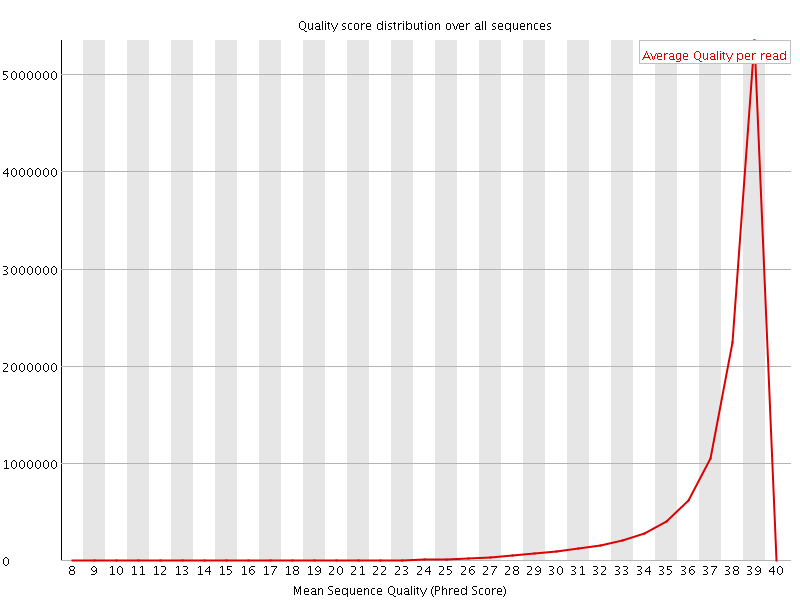
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
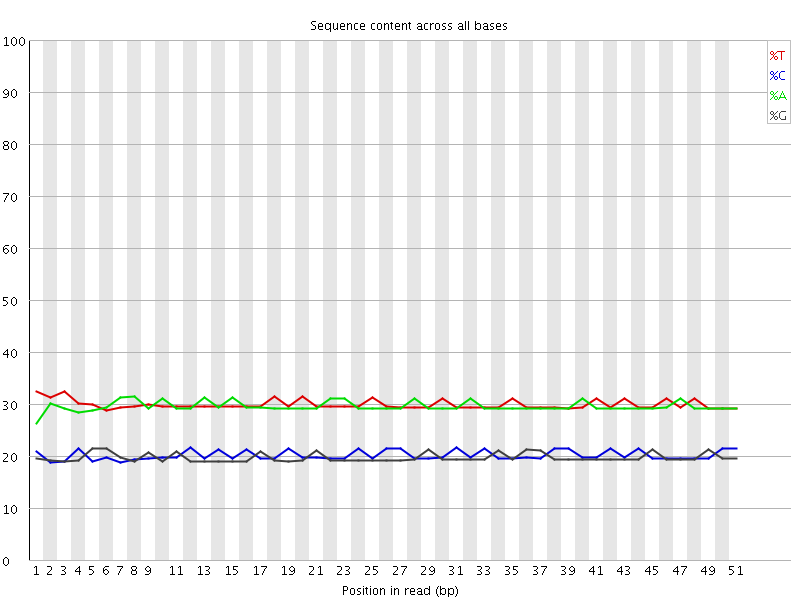
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
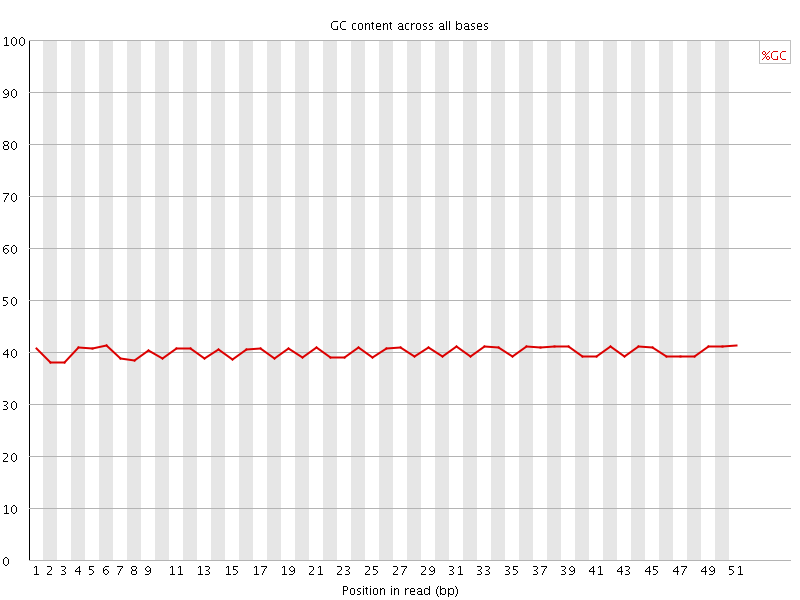
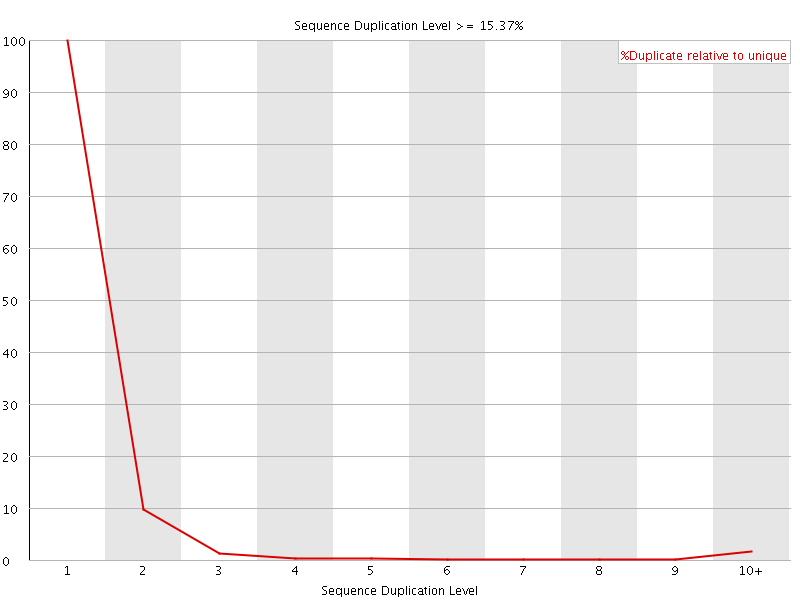
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
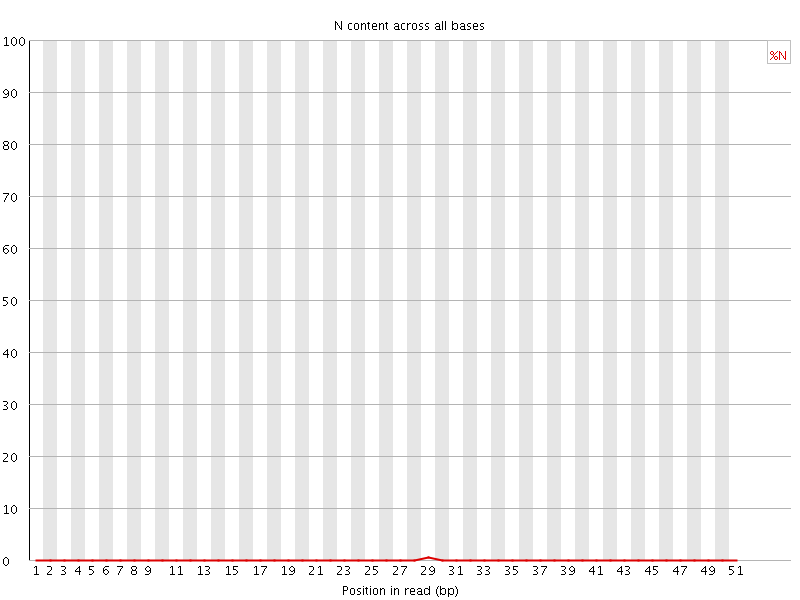
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
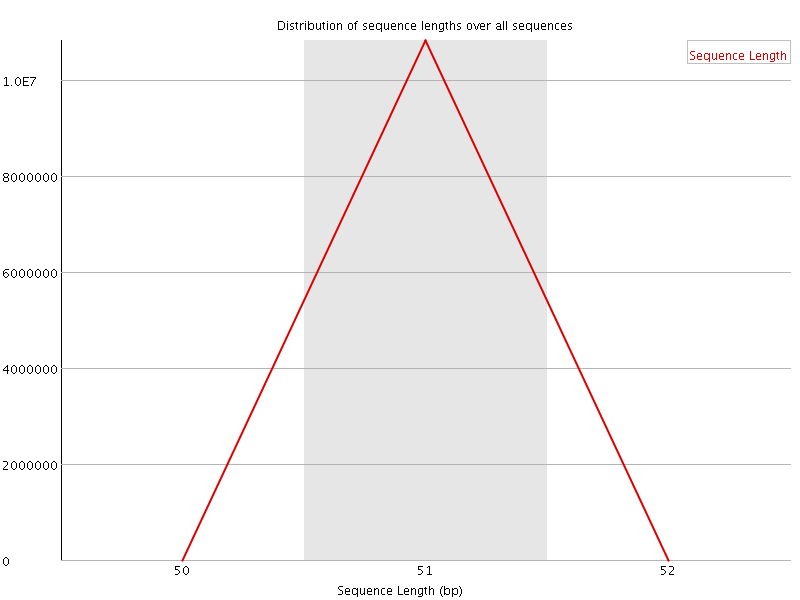
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGTGGCCATCTCGTATGCC | 183604 | 1.6960967530062105 | TruSeq Adapter, Index 1 (97% over 35bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
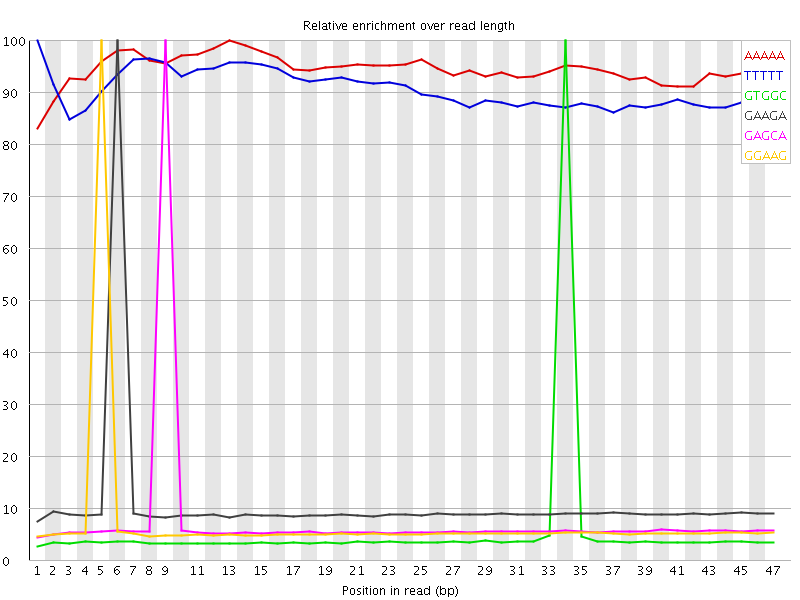
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 4303580 | 3.6414428 | 3.8551555 | 13 |
| TTTTT | 4444315 | 3.5536819 | 3.9218087 | 1 |
| GTGGC | 545840 | 2.2452128 | 39.594772 | 34 |
| GAAGA | 1164805 | 2.2123554 | 20.412155 | 6 |
| GAGCA | 787165 | 2.1857116 | 28.751776 | 9 |
| GGAAG | 746885 | 2.1253693 | 29.465801 | 5 |
| CTCCA | 811470 | 2.121185 | 26.58706 | 24 |
| TCCAG | 763835 | 2.0462523 | 27.169739 | 25 |
| CGGAA | 730095 | 2.0272462 | 28.771986 | 4 |
| CGTGG | 491840 | 2.023094 | 39.42574 | 33 |
| CTGAA | 1101665 | 2.0187554 | 19.3331 | 19 |
| CACGT | 748840 | 2.0060818 | 26.848217 | 14 |
| GGCCA | 473600 | 1.922491 | 38.619556 | 36 |
| TGGCC | 475020 | 1.90656 | 38.092567 | 35 |
| TCTCG | 692955 | 1.8354844 | 26.178923 | 41 |
| TCGGA | 637555 | 1.750374 | 28.23686 | 3 |
| CCATC | 665000 | 1.7383118 | 25.403042 | 38 |
| AGAGC | 620865 | 1.7239484 | 28.266777 | 8 |
| CACAC | 639520 | 1.6907296 | 26.890474 | 12 |
| AGCAC | 617415 | 1.6728258 | 27.559547 | 10 |
| GCACA | 610710 | 1.6546593 | 27.510569 | 11 |
| TCTGA | 891605 | 1.6154467 | 18.753553 | 18 |
| ATCGG | 581615 | 1.5967935 | 27.95068 | 2 |
| AAGAG | 822440 | 1.5620894 | 19.701384 | 7 |
| GATCG | 566900 | 1.5563942 | 27.749554 | 1 |
| CCAGT | 561365 | 1.5038514 | 26.526367 | 26 |
| CTCGT | 566710 | 1.5010893 | 25.69923 | 42 |
| CGTCT | 558215 | 1.478588 | 26.610931 | 16 |
| ACGTG | 531750 | 1.4598918 | 26.473843 | 32 |
| TGAAC | 795640 | 1.4579773 | 18.815023 | 20 |
| GTCTG | 530310 | 1.4395572 | 27.127737 | 17 |
| GTCAC | 534590 | 1.4321233 | 26.402872 | 29 |
| GCCAT | 534310 | 1.4313734 | 25.62264 | 37 |
| ATGCC | 532390 | 1.4262297 | 25.839851 | 47 |
| ACACG | 516435 | 1.3992304 | 27.161997 | 13 |
| ACGTC | 517325 | 1.3858718 | 26.836355 | 15 |
| ACTCC | 530040 | 1.3855262 | 26.006794 | 23 |
| TCACG | 516015 | 1.3823625 | 26.103456 | 30 |
| CATCT | 759675 | 1.343057 | 17.404318 | 39 |
| AACTC | 746585 | 1.3349341 | 18.233004 | 22 |
| CAGTC | 496695 | 1.3306057 | 26.335762 | 27 |
| GAACT | 719200 | 1.3179041 | 18.634735 | 21 |
| ATCTC | 744640 | 1.3164761 | 17.500216 | 40 |
| AGTCA | 619630 | 1.1354462 | 18.178583 | 28 |
| TCGTA | 571455 | 1.0353857 | 17.502941 | 43 |
| TATGC | 511755 | 0.92721885 | 17.34376 | 46 |
| CGTAT | 504965 | 0.91491646 | 17.374865 | 44 |
| GTATG | 487970 | 0.9060806 | 17.784706 | 45 |