![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | LW200_GTGAAA_L003_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 9633780 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
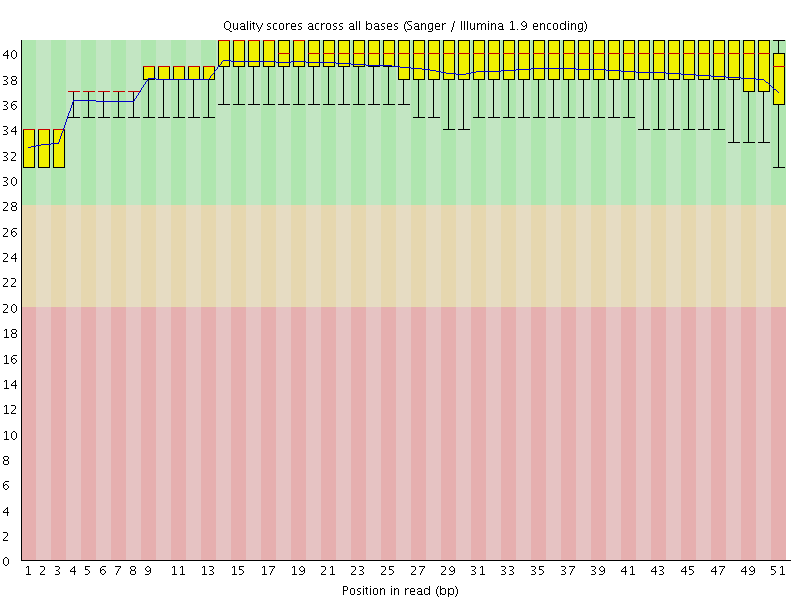
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
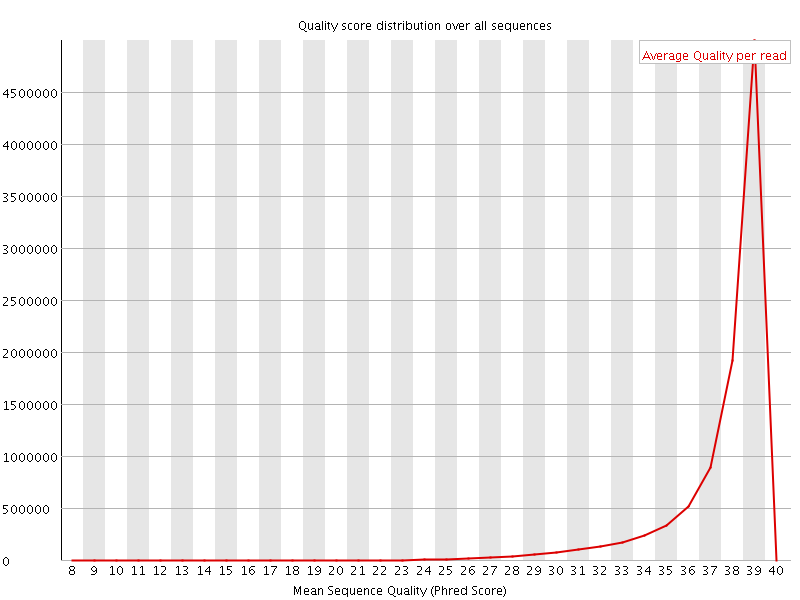
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
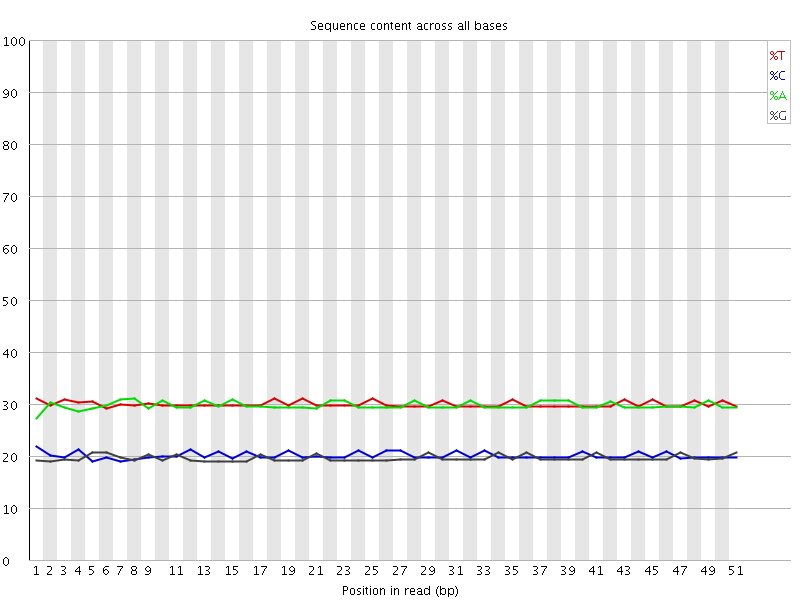
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
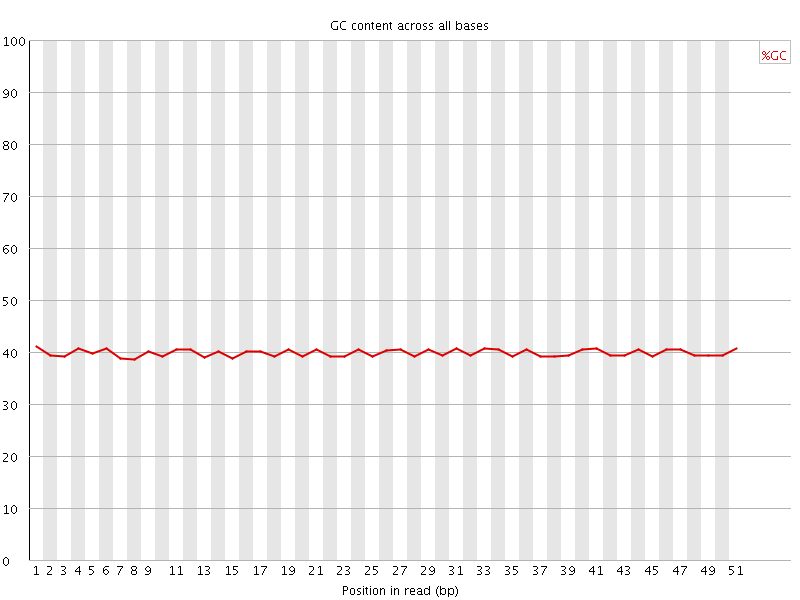
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
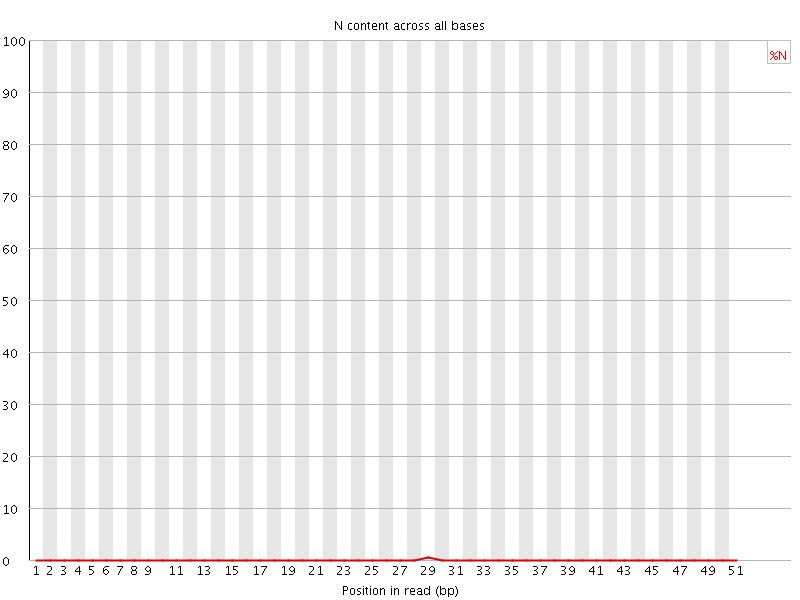
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
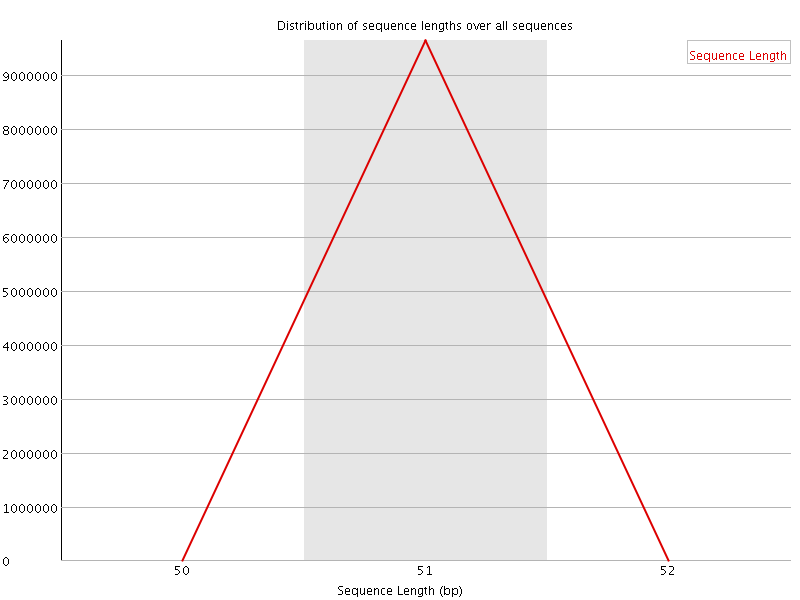
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
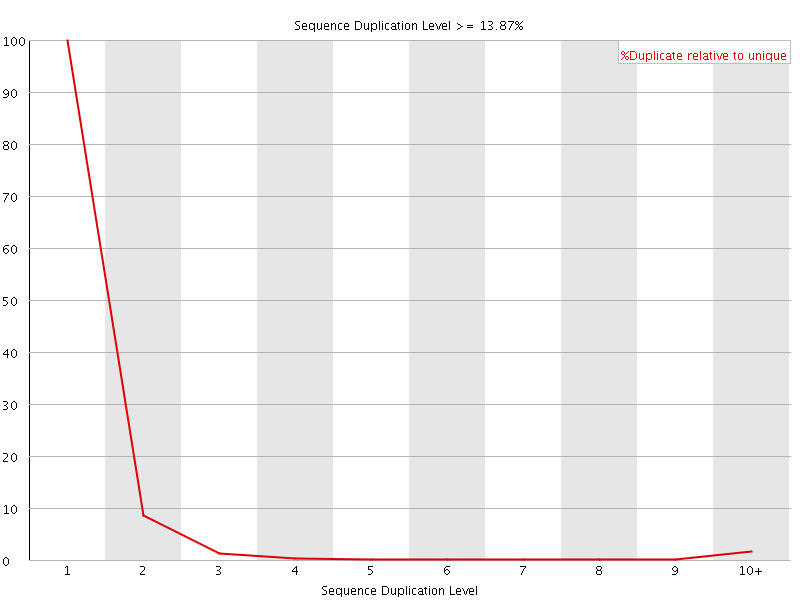
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGTGAAACGATCTCGTATG | 106515 | 1.1056407765176286 | TruSeq Adapter, Index 1 (97% over 35bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
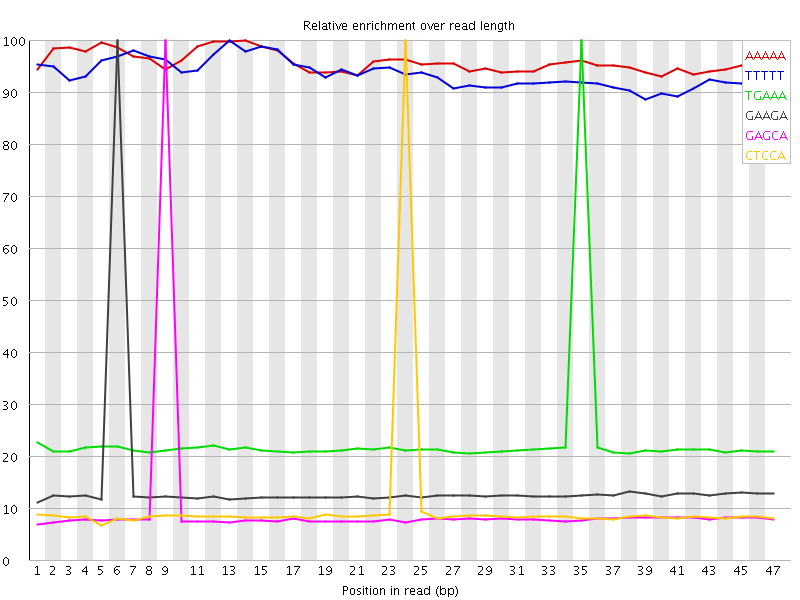
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 3835890 | 3.5584726 | 3.7110236 | 14 |
| TTTTT | 3985770 | 3.5583336 | 3.8029275 | 13 |
| TGAAA | 1674790 | 2.3335145 | 10.128292 | 35 |
| GAAGA | 993785 | 2.1118379 | 14.749843 | 6 |
| GAGCA | 647270 | 2.0289617 | 20.582945 | 9 |
| CTCCA | 668130 | 1.9741794 | 18.952 | 24 |
| GGAAG | 610585 | 1.9638107 | 21.189184 | 5 |
| CTGAA | 926830 | 1.9048928 | 13.89027 | 19 |
| CGGAA | 606320 | 1.9005979 | 20.594719 | 4 |
| TCCAG | 616835 | 1.8700777 | 19.352114 | 25 |
| CACGT | 562430 | 1.7051364 | 19.082994 | 14 |
| TCTCG | 564485 | 1.6982846 | 18.450594 | 43 |
| GAAAC | 792330 | 1.6410022 | 13.06056 | 36 |
| CACAC | 548115 | 1.6320367 | 19.129173 | 12 |
| TCGGA | 516410 | 1.6063879 | 20.188265 | 3 |
| AGAGC | 495885 | 1.5544236 | 20.117125 | 8 |
| GCACA | 502775 | 1.5360205 | 19.640959 | 11 |
| TCTGA | 740890 | 1.5110943 | 13.419174 | 18 |
| AGCAC | 492560 | 1.5048126 | 19.585014 | 10 |
| GTGAA | 712790 | 1.503132 | 13.337605 | 34 |
| AAGAG | 688390 | 1.4628599 | 14.066878 | 7 |
| ATCGG | 463960 | 1.4432325 | 19.890316 | 2 |
| AAACG | 667975 | 1.3834493 | 12.808134 | 37 |
| GATCG | 439375 | 1.3667564 | 19.660181 | 1 |
| CTCGT | 454210 | 1.3665161 | 18.081757 | 44 |
| CCAGT | 447345 | 1.3562297 | 18.753609 | 26 |
| TGAAC | 657200 | 1.3507284 | 13.328516 | 20 |
| CGTCT | 447000 | 1.3448244 | 18.841164 | 16 |
| ACGTG | 423620 | 1.3177476 | 18.86655 | 32 |
| GTCAC | 427285 | 1.2954131 | 18.671888 | 29 |
| GTCTG | 415945 | 1.2839824 | 19.307196 | 17 |
| CGATC | 422810 | 1.2818463 | 17.706495 | 40 |
| ACACG | 407360 | 1.2445194 | 19.180506 | 13 |
| ACGTC | 409240 | 1.2407056 | 18.921518 | 15 |
| CGTGA | 398560 | 1.2397939 | 18.778065 | 33 |
| ACTCC | 419205 | 1.23866 | 18.318695 | 23 |
| TCACG | 406765 | 1.2332021 | 18.566828 | 30 |
| AACGA | 594140 | 1.2305292 | 12.549183 | 38 |
| AACTC | 611925 | 1.2257549 | 12.867359 | 22 |
| ATCTC | 614355 | 1.2212151 | 12.236258 | 42 |
| GAACT | 581830 | 1.1958221 | 13.136937 | 21 |
| CAGTC | 386975 | 1.1732041 | 18.490555 | 27 |
| ACGAT | 532175 | 1.0937673 | 12.308937 | 39 |
| GATCT | 526145 | 1.0731076 | 12.330287 | 41 |
| AGTCA | 503285 | 1.0343903 | 12.846619 | 28 |
| TCGTA | 440560 | 0.89855134 | 12.329308 | 45 |
| CGTAT | 389190 | 0.79377884 | 12.224575 | 46 |
| GTATG | 378655 | 0.7924038 | 12.508594 | 47 |